

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0214.6
(398 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
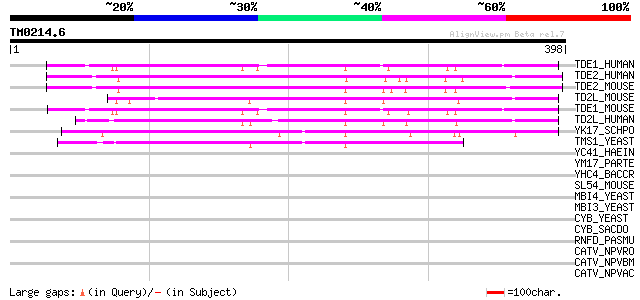
Sequences producing significant alignments: (bits) Value
TDE1_HUMAN (Q13530) Tumor differentially expressed protein 1 (Tr... 146 1e-34
TDE2_HUMAN (Q9NRX5) Tumor differentially expressed protein 2 (Tu... 140 4e-33
TDE2_MOUSE (Q9QZI8) Tumor differentially expressed protein 2 (Tu... 136 1e-31
TD2L_MOUSE (Q8K0E7) Tumor differentially expressed 2-like 133 9e-31
TDE1_MOUSE (Q9QZI9) Tumor differentially expressed protein 1 (Me... 130 8e-30
TD2L_HUMAN (Q96SA4) Tumor differentially expressed 2-like (FKSG8... 129 1e-29
YK17_SCHPO (Q9HDY3) Membrane protein PB1A10.07c 118 2e-26
TMS1_YEAST (Q12116) Membrane protein TMS1 95 3e-19
YC41_HAEIN (P44133) Hypothetical protein HI1241 35 0.26
YM17_PARTE (P15618) Hypothetical 26.3 kDa protein (ORF17) 35 0.34
YHC4_BACCR (Q814U5) Hypothetical UPF0059 protein BC5324 35 0.44
SL54_MOUSE (Q9ET37) Low affinity sodium-glucose cotransporter (S... 34 0.57
MBI4_YEAST (P03879) Intron-encoded RNA maturase bI4 precursor (R... 33 1.3
MBI3_YEAST (Q9ZZW7) Cytochrome b mRNA maturase bI3 33 1.3
CYB_YEAST (P00163) Cytochrome b 33 1.3
CYB_SACDO (Q35819) Cytochrome b 33 1.3
RNFD_PASMU (Q9CNP3) Electron transport complex protein rnfD 33 1.7
CATV_NPVRO (Q8B9D5) Viral cathepsin (EC 3.4.22.50) (V-cath) (Cys... 32 2.9
CATV_NPVBM (P41721) Viral cathepsin (EC 3.4.22.50) (V-cath) (Cys... 32 2.9
CATV_NPVAC (P25783) Viral cathepsin (EC 3.4.22.50) (V-cath) (Cys... 32 2.9
>TDE1_HUMAN (Q13530) Tumor differentially expressed protein 1
(Transmembrane protein SBBI99)
Length = 473
Score = 146 bits (368), Expect = 1e-34
Identities = 108/449 (24%), Positives = 203/449 (45%), Gaps = 91/449 (20%)
Query: 27 NASNPGMARYVYALMFLVANLLAWAARDYGRGALTEMERLKG-CNGG-----------KD 74
N+ N + R +YA + L++ ++++ + + T ++++ G C GG KD
Sbjct: 31 NSKNSTVTRLIYAFILLLSTVVSYIMQR--KEMETYLKKIPGFCEGGFKIHEADINADKD 88
Query: 75 C---LGAEGVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIP 131
C +G + V R+S IF+F+ L + ++R H+G+W KI +G+ V
Sbjct: 89 CDVLVGYKAVYRISFAMAIFFFVFSLLMFKVKTSKDLRAAVHNGFWFFKIAALIGIMVGS 148
Query: 132 FLLPSE-FIQIYGEVAHFGAGVFLLIQLISIISFI-----TWLNDCCESEK---YAAKCQ 182
F +P F ++ V GA +F+LIQL+ ++ F +W+N E YAA
Sbjct: 149 FYIPGGYFSSVWFVVGMIGAALFILIQLVLLVDFAHSWNESWVNRMEEGNPRLWYAA--- 205
Query: 183 IHVMLFATTAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPKV--- 239
++ F + Y++ ++ + L++ +Y C N FFI+ L+L + + +S+HPK+
Sbjct: 206 --LLSFTSAFYILSIICVGLLYTYYTKPDGCTENKFFISINLILCVVASIISIHPKIQEH 263
Query: 240 --NAGILTPGLMGLYVVFLCWNAIRSEPDGGSC--------------------------- 270
+G+L L+ LY ++L W+A+ +EPD SC
Sbjct: 264 QPRSGLLQSSLITLYTMYLTWSAMSNEPD-RSCNPNLMSFITRITAPTLAPGNSTAVVPT 322
Query: 271 -IRKSDSSTKTDWLSIISFIIAILAIVIATFSTGIDSKCFQFR----------------- 312
S S + D + I + +L ++ ++ T +S+ +
Sbjct: 323 PTPPSKSGSLLDSDNFIGLFVFVLCLLYSSIRTSTNSQVDKLTLSGSDSVILGDTTTSGA 382
Query: 313 --KDDDIP------AEDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWT 364
++D P ++ V Y Y FH + ++Y M L W S + + ++ W
Sbjct: 383 SDEEDGQPRRAVDNEKEGVQYSYSLFHLMLCLASLYIMMTLTSWYSPDA-KFQSMTSKWP 441
Query: 365 STWVRIVNEWLAVCVYLWMLVAPIIWKSR 393
+ WV+I + W+ + +Y+W LVAP++ SR
Sbjct: 442 AVWVKISSSWVCLLLYVWTLVAPLVLTSR 470
>TDE2_HUMAN (Q9NRX5) Tumor differentially expressed protein 2 (Tumor
differentially expressed 1 protein like) (UNQ396/PRO732)
Length = 453
Score = 140 bits (354), Expect = 4e-33
Identities = 109/427 (25%), Positives = 192/427 (44%), Gaps = 61/427 (14%)
Query: 27 NASNPGMARYVYALMFLVANLLAWAARDYGRGALTEMERLKG-CNGGKDCL------GAE 79
+ +N + R +YAL LV +A G ++ ++ G C K + G +
Sbjct: 31 SGNNSTVTRLIYALFLLVGVCVACVMLIPGMEE--QLNKIPGFCENEKGVVPCNILVGYK 88
Query: 80 GVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPS-EF 138
V R+ G +FY ++ L + ++ R H+G+W K + + + F +P F
Sbjct: 89 AVYRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFAAAIAIIIGAFFIPEGTF 148
Query: 139 IQIYGEVAHFGAGVFLLIQLISIISFI-TWLNDCCES-EKYAAKCQIHVMLFATTA-YVV 195
++ V GA F+LIQL+ +I F +W E E+ ++C +L AT Y++
Sbjct: 149 TTVWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNYLL 208
Query: 196 CLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPKVN-----AGILTPGLMG 250
LV I+L F++Y SC N FI+ ++L + +S+ PK+ +G+L ++
Sbjct: 209 SLVAIVLFFVYYTHPASCSENKAFISVNMLLCVGASVMSILPKIQESQPRSGLLQSSVIT 268
Query: 251 LYVVFLCWNAIRSEPDGG------SCIRKSDSST------KTDWL---SIISFIIAILAI 295
+Y ++L W+A+ +EP+ S I + +ST W II I+ +L +
Sbjct: 269 VYTMYLTWSAMTNEPETNCNPSLLSIIGYNTTSTVPKEGQSVQWWHAQGIIGLILFLLCV 328
Query: 296 VIATFSTGIDSKCFQF---------------RKDDDIPAEDDV-----------PYGYGF 329
++ T +S+ + R D + DDV Y Y F
Sbjct: 329 FYSSIRTSNNSQVNKLTLTSDESTLIEDGGARSDGSLEDGDDVHRAVDNERDGVTYSYSF 388
Query: 330 FHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPII 389
FHF+ ++Y M L W + R+ WT+ WV+I + W+ + +Y+W LVAP++
Sbjct: 389 FHFMLFLASLYIMMTLTNWYRYEPSREMKSQ--WTAVWVKISSSWIGIVLYVWTLVAPLV 446
Query: 390 WKSRQTD 396
+R D
Sbjct: 447 LTNRDFD 453
>TDE2_MOUSE (Q9QZI8) Tumor differentially expressed protein 2 (Tumor
differentially expressed 1 protein like) (Membrane
protein TMS-2) (Axotomy induced glyco/Golgi protein 2)
Length = 453
Score = 136 bits (342), Expect = 1e-31
Identities = 106/427 (24%), Positives = 191/427 (43%), Gaps = 61/427 (14%)
Query: 27 NASNPGMARYVYALMFLVANLLAWAARDYGRGALTEMERLKG-CNGGKDCL------GAE 79
+ +N + R +YAL LV +A G ++ ++ G C K + G +
Sbjct: 31 SGNNSTVTRLIYALFLLVGVCVACVMLIPGMEE--QLNKIPGFCENEKGVVPCNILVGYK 88
Query: 80 GVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPS-EF 138
V R+ G +FY ++ L + ++ R H+G+W K V + + F +P F
Sbjct: 89 AVYRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFATAVAIIIGAFFIPEGTF 148
Query: 139 IQIYGEVAHFGAGVFLLIQLISIISFI-TWLNDCCES-EKYAAKCQIHVMLFATTA-YVV 195
++ V GA F+LIQL+ +I F +W E E+ ++C +L AT Y++
Sbjct: 149 TTVWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNYLL 208
Query: 196 CLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPKV-----NAGILTPGLMG 250
LV ++L F++Y SC N FI+ ++L + +S+ PK+ +G+L ++
Sbjct: 209 SLVAVVLFFVYYTHPASCAENKAFISVNMLLCIGASVMSILPKIQESQPRSGLLQSSVIT 268
Query: 251 LYVVFLCWNAIRSEPD-----------GGSCIR--KSDSSTKTDW--LSIISFIIAILAI 295
+Y ++L W+A+ +EP+ G + R D + W II ++ +L +
Sbjct: 269 VYTMYLTWSAMTNEPETNCNPSLLSIIGFNTTRPIPKDGQSVQWWHPQGIIGLVLFLLCV 328
Query: 296 VIATFSTGIDSKCFQF---------------RKDDDIP-----------AEDDVPYGYGF 329
++ T +S+ + R D + D V Y Y F
Sbjct: 329 FYSSIRTSNNSQVNKLTLTSDESTLIEDGNGRSDGSLDDGDGIHRAVDNERDGVTYSYSF 388
Query: 330 FHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPII 389
FHF+ ++Y M L W + R+ + WT+ WV+I + W+ + +Y+W LVAP++
Sbjct: 389 FHFMLFLASLYIMMTLTNWYRYEPSRE--MKSQWTAVWVKISSSWIGLVLYVWTLVAPLV 446
Query: 390 WKSRQTD 396
+R D
Sbjct: 447 LTNRDFD 453
>TD2L_MOUSE (Q8K0E7) Tumor differentially expressed 2-like
Length = 450
Score = 133 bits (334), Expect = 9e-31
Identities = 101/368 (27%), Positives = 167/368 (44%), Gaps = 49/368 (13%)
Query: 71 GGKDC---LGAEGVLRV--SLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWV 125
G DC LG V R+ + F F+F++ + R+S+ + R +G+W K ++ V
Sbjct: 84 GPLDCGSLLGFRAVYRMCFATAAFFFFFMLLMICVRSSR--DPRAAIQNGFWFFKFLILV 141
Query: 126 GMTVIPFLLPS-EFIQIYGEVAHFGAGVFLLIQLISIISFITWLND--CCESEKYAAKCQ 182
G+TV F +P F +I+ G+ +F+LIQLI + F N C++E+ +
Sbjct: 142 GITVGAFYIPDGSFPKIWFYFGVVGSFLFILIQLILFVDFAHSWNQRWLCKAEECDSPAW 201
Query: 183 IHVMLFATTA-YVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPKV-- 239
+ F T Y++ + + LMF++Y +C FI+ L ++ +++ PKV
Sbjct: 202 YAGLFFFTFLFYLLSIAAVALMFVYYTESGACHEGKVFISLNLTFCVCVSIIAVLPKVQD 261
Query: 240 ---NAGILTPGLMGLYVVFLCWNAIRSEPD-----------GGSCIRKSDSSTKT-DWLS 284
N+G+L ++ LY +F+ W+A+ + PD G + D ST D S
Sbjct: 262 AQPNSGLLQASVITLYTMFVTWSALSNVPDQKCNPHLPTKNGTGQVDLEDYSTVWWDAPS 321
Query: 285 IISFIIAILAIVIATFSTGIDSKCFQFRKDDDIPAE-------------------DDVPY 325
I+ +I IL + + + + ++ PAE D V Y
Sbjct: 322 IVGLVIFILCTFFISLRSSDHRQVNSLMQTEECPAEMVQQQQVAVSDGRAYDNEQDGVTY 381
Query: 326 GYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLV 385
Y FFHF +++ M L W S RK WTS WV+I W + +YLW LV
Sbjct: 382 SYSFFHFCLVLASLHVMMTLTNWYSPGETRKMIST--WTSVWVKICASWAGLFLYLWTLV 439
Query: 386 APIIWKSR 393
AP++ +R
Sbjct: 440 APLLLPNR 447
>TDE1_MOUSE (Q9QZI9) Tumor differentially expressed protein 1
(Membrane protein TMS-1) (Axotomy induced glycoprotein
1) (Axotomy induced glyco/Golgi protein 1) (AIGP-1)
Length = 472
Score = 130 bits (326), Expect = 8e-30
Identities = 101/447 (22%), Positives = 193/447 (42%), Gaps = 90/447 (20%)
Query: 28 ASNPGMARYVYALMFLVANLLAWAARDYGRGALTEMERLKG-CNGG-----------KDC 75
+ N + R +YA + + +++ G T+++++ G C GG KDC
Sbjct: 32 SKNSTVTRLIYAFILFLGTIVSCIMMT--EGIQTQLKKIPGFCEGGFQIKMVDTKAEKDC 89
Query: 76 ---LGAEGVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPF 132
+G + V R++ IF+F FL + + R H+G+W KI +G+ + F
Sbjct: 90 DVLVGFKAVYRINFAVAIFFFAFFLLMLKVKTSKDPRAAVHNGFWFFKIAAIIGIMIGSF 149
Query: 133 LLP-SEFIQIYGEVAHFGAGVFLLIQLISIISFI-----TWLNDCCESEK---YAAKCQI 183
+P F +++ GA F++IQL+ ++ W+N E YAA
Sbjct: 150 YIPGGSFTEVWFVAGMLGASFFIIIQLVLLVDMAHSWNELWVNRMEEGNPRLWYAA---- 205
Query: 184 HVMLFATTAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPKV---- 239
++ F + Y++ +V L++++Y C N FI+ L+ ++ VS+ PKV
Sbjct: 206 -LLSFTSLFYILSIVFAALLYVFYTKPDDCTENKVFISLNLIFCVAVSIVSILPKVQEHQ 264
Query: 240 -NAGILTPGLMGLYVVFLCWNAIRSEPDGGSC---------------IRKSDSSTKTDWL 283
+G+L ++ LY ++L W+A+ +EP+ SC + ++S+T
Sbjct: 265 PRSGLLQSSIITLYTLYLTWSAMTNEPE-RSCNPSLMSIITHLTSPTVSPANSTTLAPAY 323
Query: 284 S-------------IISFIIAILAIVIATFSTGIDSKCFQFR------------------ 312
+ I II + ++ ++F T +S+ +
Sbjct: 324 APPSQSGHFMNLDDIWGLIIFVFCLIYSSFRTSSNSQVNKLTLSGSDSVILGDTTNGAND 383
Query: 313 KDDDIP------AEDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTST 366
++D P ++ V Y Y FFH + ++Y M + W S + + + W +
Sbjct: 384 EEDGQPRRAVDNEKEGVQYSYSFFHLMLCCASLYIMMTITSWYSPDA-KFQKVSSKWLAV 442
Query: 367 WVRIVNEWLAVCVYLWMLVAPIIWKSR 393
W ++ + WL + +YLW LVAP++ R
Sbjct: 443 WFKMGSSWLCLLLYLWTLVAPLVLTGR 469
>TD2L_HUMAN (Q96SA4) Tumor differentially expressed 2-like (FKSG84
protein) (UNQ263/PRO300)
Length = 456
Score = 129 bits (325), Expect = 1e-29
Identities = 99/396 (25%), Positives = 163/396 (41%), Gaps = 60/396 (15%)
Query: 48 LAWAARDYGRGALTEMERLKGCNGGKDCLGAEGVLRVSLGCFIFYFIMFLTTARTSKLNE 107
L W + G G T ++ C LG V R+ F+F L S +
Sbjct: 68 LPWVCEE-GAGIPTVLQGHIDCGS---LLGYRAVYRMCFATAAFFFFFTLLMLCVSSSRD 123
Query: 108 VRDTWHSGWWSVKIVLWVGMTVIPFLLPS-EFIQIYGEVAHFGAGVFLLIQLISIISFI- 165
R +G+W K ++ VG+TV F +P F I+ G+ +F+LIQL+ +I F
Sbjct: 124 PRAAIQNGFWFFKFLILVGLTVGAFYIPDGSFTNIWFYFGVVGSFLFILIQLVLLIDFAH 183
Query: 166 ----TWLNDC--CESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYAPKPSCLLNIFF 219
WL C+S + A +LF Y++ + + LMF++Y C F
Sbjct: 184 SWNQRWLGKAEECDSRAWYAGLFFFTLLF----YLLSIAAVALMFMYYTEPSGCHEGKVF 239
Query: 220 IAWTLVLLQLMTSVSLHPKV-----NAGILTPGLMGLYVVFLCWNAIRSEPD-------- 266
I+ L ++ ++ PKV N+G+L ++ LY +F+ W+A+ S P+
Sbjct: 240 ISLNLTFCVCVSIAAVLPKVQDAQPNSGLLQASVITLYTMFVTWSALSSIPEQKCNPHLP 299
Query: 267 ---GGSCIRKSDSSTKTDWL---SIISFIIAILAIVIATFSTGIDSKCFQFRKDDDIPA- 319
G + +T W SI+ II +L + + + + + ++ P
Sbjct: 300 TQLGNETVVAGPEGYETQWWDAPSIVGLIIFLLCTLFISLRSSDHRQVNSLMQTEECPPM 359
Query: 320 ----------------------EDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKW 357
+D V Y Y FFHF +++ M L W RK
Sbjct: 360 LDATQQQQQQVAACEGRAFDNEQDGVTYSYSFFHFCLVLASLHVMMTLTNWYKPGETRKM 419
Query: 358 TIDVGWTSTWVRIVNEWLAVCVYLWMLVAPIIWKSR 393
WT+ WV+I W + +YLW LVAP++ ++R
Sbjct: 420 IST--WTAVWVKICASWAGLLLYLWTLVAPLLLRNR 453
>YK17_SCHPO (Q9HDY3) Membrane protein PB1A10.07c
Length = 441
Score = 118 bits (296), Expect = 2e-26
Identities = 93/393 (23%), Positives = 169/393 (42%), Gaps = 38/393 (9%)
Query: 38 YALMFLVANLLAWAA-RDYGRGALTEMER---LKGCNGGKDCLGAEGVLRVSLGCFIFYF 93
YA+++ V +LL+W + L+++ C C V R+S +F+
Sbjct: 47 YAVLYFVNSLLSWCMLSSWFNSKLSKLSAGYLQFDCQNDGKCYSVIAVHRLSFTLVMFHL 106
Query: 94 IMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGE-VAHFGAGV 152
+ + + + V +G W KIVLW + + F +P++F+ +G ++ G+ +
Sbjct: 107 FLAFILSLCNTRSRVAIKIQNGLWPFKIVLWFVLGIFSFFIPTKFLSFWGNIISVMGSAL 166
Query: 153 FLLIQLISIISFI-TWLNDCCESEKYAAKCQIHVMLFATTA--YVVCLVGIILMFIWYAP 209
F++ L+ ++ F TW C + + L +T YVV LV IL ++++
Sbjct: 167 FIVYGLMLLVDFAHTWAERCVDRVLTSDSSSSKFYLIGSTVGMYVVGLVLTILTYVFFCA 226
Query: 210 KPSCLLNIFFIAWTLVLLQLMTSVSLHPKV-----NAGILTPGLMGLYVVFLCWNAIRSE 264
SC N L+L ++ +S+HP + +G+ ++ Y +L +A+ +
Sbjct: 227 S-SCSFNQAINTINLLLCIAVSCLSVHPTIQEYNPRSGLAQSSMVMCYTCYLILSALANR 285
Query: 265 PDGGSCIRKSDSSTKTDWLSII------SFIIAILAIVIATFSTGIDSKCFQFRKDDDI- 317
PD G C +S++ T S + F I A+ A+ DS + + D+
Sbjct: 286 PDEGQCNPWGNSASGTREFSKVIGAAFTFFTILYSAVRAASSRESDDSYSYLYADSHDMG 345
Query: 318 ---PAED--------DVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDV----- 361
P ED Y + +FH VF A Y A LL WN+ DV
Sbjct: 346 VSTPLEDGSSEEDKHQSDYNFIWFHIVFVLAAFYTASLLTNWNTTSVYENQKNDVFVRIG 405
Query: 362 -GWTSTWVRIVNEWLAVCVYLWMLVAPIIWKSR 393
+ + WV+I+ W+ +Y+W +AP+ + R
Sbjct: 406 FSYAAVWVKIITSWVCHGLYVWSCLAPVFFPYR 438
>TMS1_YEAST (Q12116) Membrane protein TMS1
Length = 473
Score = 95.1 bits (235), Expect = 3e-19
Identities = 70/302 (23%), Positives = 141/302 (46%), Gaps = 16/302 (5%)
Query: 35 RYVYALMFLVANLLAWAARDYGRGALTEMERLKGCNGGKDCLGAEGVLRVSLGCFIFYFI 94
R +YA+ L+ +L++W + + L K C G +C G V R++ + I
Sbjct: 43 RLLYAVWLLLNSLISWVSYSANKSILWPG---KTCTGTGEC-GFFTVHRLNFALGCLHLI 98
Query: 95 MFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAG-VF 153
+ L N+VR + WWS+K +L++ + V+ F++P++F + + +G +F
Sbjct: 99 LALVLTGVKSTNDVRAALQNSWWSLKFILYLCLIVLSFVIPNDFYIFFSKWVSVPSGAIF 158
Query: 154 LLIQLISIISFI-TWLNDC---CESE-KYAAKCQIHVMLFATTAYVVCLVGIILMFIWYA 208
+L+ LI ++ F W C ESE + ++ Q ++L T+ Y ++ ++M++ +
Sbjct: 159 ILVGLILLVDFAHEWAETCISHVESEDEDSSFWQRFLVLGTTSMYTASIIMTVVMYVMFC 218
Query: 209 PKPSCLLNIFFIAWTLVLLQLMTSVSLHPKV-----NAGILTPGLMGLYVVFLCWNAIRS 263
+ C +N + L+L + +S++PK+ +G+ ++ +Y +L +A+ S
Sbjct: 219 HQ-QCNMNQTAVTVNLILTVITLVLSVNPKIQEANPKSGLAQSSMVSVYCTYLTMSAMSS 277
Query: 264 EPDGGSCIRKSDSSTKTDWLSIISFIIAILAIVIATFSTGIDSKCFQFRKDDDIPAEDDV 323
EPD C SS + I+ + +AI T +S + I +D+
Sbjct: 278 EPDDKMCNPLVRSSGTRKFSIILGSLFTFIAIAYTTTRAAANSAFQGTNTNGAIYLGNDI 337
Query: 324 PY 325
Y
Sbjct: 338 EY 339
Score = 41.2 bits (95), Expect = 0.005
Identities = 24/71 (33%), Positives = 36/71 (49%), Gaps = 3/71 (4%)
Query: 325 YGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTS--TWVRIVNEWLAVCVYLW 382
Y Y FH +F + A+LL + + + I VG T +WV+IV+ W+ +Y W
Sbjct: 397 YNYTLFHVIFFLATQWIAILLTINVTQDDVGDF-IPVGRTYFYSWVKIVSAWICYALYGW 455
Query: 383 MLVAPIIWKSR 393
+VAP I R
Sbjct: 456 TVVAPAIMPDR 466
>YC41_HAEIN (P44133) Hypothetical protein HI1241
Length = 266
Score = 35.4 bits (80), Expect = 0.26
Identities = 65/289 (22%), Positives = 114/289 (38%), Gaps = 63/289 (21%)
Query: 119 VKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAGVFLLIQLISIISFITWLNDCCESEK-- 176
+ I + +G+ + F L SE + H G GV L+ QLIS++ F + + C ++
Sbjct: 16 LSISMVLGIDLFIFSLQSE----KQTMPHLGVGV-LVAQLISLLVF--YRGEICPGQRGR 68
Query: 177 -------YAAKCQIHVMLFA-----TTAYVVCLVGI-ILMFIWYAPKPSCLLNIFFIAWT 223
+A + +++ T V+ + G+ ++ FIW PK + N F +
Sbjct: 69 LIKVNMTFAIYWAVWLLISLLQNNHTLTNVMSVCGLSVVYFIWKQPKTEKIRNSFLLMAA 128
Query: 224 LV-------LLQLMTSVSLHPKVNAGILTPGLMGLYVVFLCWNAIRSEPDGGSCIRKSDS 276
L+ L + T + P L G+ + L R+ G +
Sbjct: 129 LIAGLGCLSYLMIFTELPASDFAEYNPFAPILSGVILANLVLVIARNRLQGFIAL----- 183
Query: 277 STKTDWLSIISFIIAILAIVIATFSTGIDSKCFQFRKDDDIPAEDDVPY-GYGFFHFVFA 335
II + LA+ + G++S + +E Y Y HFV A
Sbjct: 184 ---LPLAMIILLALNALAMFLFLLLNGMESA---------VNSESVFAYIIYFVCHFVIA 231
Query: 336 TGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWML 384
A+L++ HS +KWT+ ST + ++AVC+ LWM+
Sbjct: 232 ------AILIL-----HSFQKWTL-----STNSLFILLFIAVCLPLWMV 264
>YM17_PARTE (P15618) Hypothetical 26.3 kDa protein (ORF17)
Length = 221
Score = 35.0 bits (79), Expect = 0.34
Identities = 20/83 (24%), Positives = 40/83 (48%), Gaps = 15/83 (18%)
Query: 158 LISIISFITWLNDCCESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYAPKPSCLLNI 217
L+S +F +W+ C ++ C + CL + + ++++P S L+ +
Sbjct: 13 LVSTSAFCSWV---CSRPSQSSLCSL------------CLSCLFFILLFWSPSTSYLILV 57
Query: 218 FFIAWTLVLLQLMTSVSLHPKVN 240
FF+ +T V+LQ + SL +N
Sbjct: 58 FFLLYTCVVLQTQLAYSLSAFMN 80
>YHC4_BACCR (Q814U5) Hypothetical UPF0059 protein BC5324
Length = 182
Score = 34.7 bits (78), Expect = 0.44
Identities = 39/149 (26%), Positives = 67/149 (44%), Gaps = 20/149 (13%)
Query: 122 VLWVGMTVIPFLLPSEFIQI---------YGEVAHFGAGVFLLIQLISIISFITWLNDCC 172
+L++GMT+ F + FI + YG++AHF AG LLI L I + T L +
Sbjct: 37 ILYIGMTIGIFHIIMPFIGMVLGRFLSEKYGDIAHF-AGAILLIGLGFYIVYSTILQN-- 93
Query: 173 ESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTS 232
E A I + +FA + + + I+ A +L F++ L + L+
Sbjct: 94 -EETRTAPIGISLFVFAFGVSIDSFSVGLSLGIYGAQTIITILLFGFVSMLLAWIGLLIG 152
Query: 233 ------VSLHPKVNAGILTPGLMGLYVVF 255
+ + ++ GI+ G GLY++F
Sbjct: 153 RHAKDMLGTYGEIVGGIILVG-FGLYILF 180
>SL54_MOUSE (Q9ET37) Low affinity sodium-glucose cotransporter
(Sodium/glucose cotransporter 3) (Na(+)/glucose
cotransporter 3)
Length = 656
Score = 34.3 bits (77), Expect = 0.57
Identities = 29/138 (21%), Positives = 62/138 (44%), Gaps = 5/138 (3%)
Query: 90 IFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHF- 148
+F ++ +T+ + EV ++ I ++G + L + F + E F
Sbjct: 428 LFVSVLIVTSILWVPIVEVSQGGQLVHYTEAISSYLGPPIAAVFLVAVFCKRANEQGAFW 487
Query: 149 GAGVFLLIQLISIISFITWLNDCCESEKYAAK--CQIHVMLFATTAYVVCLVGIILMFIW 206
G V L++ LI +I+ ++ C + K C +H + FA + VC+ ++++ +
Sbjct: 488 GLMVGLVMGLIRMIAEFSYGTGSCLAPSSCPKIICGVHYLYFAIILFFVCI--LVILGVS 545
Query: 207 YAPKPSCLLNIFFIAWTL 224
Y KP +++ + W L
Sbjct: 546 YLTKPIPDVHLHRLCWAL 563
>MBI4_YEAST (P03879) Intron-encoded RNA maturase bI4 precursor (RNA
maturase SCbI4) [Contains: Truncated, nonfunctional
cytochrome b; RNA maturase bI4 (EC 3.1.-.-)]
Length = 638
Score = 33.1 bits (74), Expect = 1.3
Identities = 21/83 (25%), Positives = 42/83 (50%), Gaps = 14/83 (16%)
Query: 87 GCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLW-VGMTVIPFLLPSEFIQ---IY 142
G F+ +MF+ A+ ++ + S ++ LW VG+ + + + F+ +Y
Sbjct: 85 GASFFFMVMFMHMAK--------GLYYGSYRSPRVTLWNVGVIIFILTIATAFLGYCCVY 136
Query: 143 GEVAHFGAGVFLLIQLISIISFI 165
G+++H+GA V + L S I F+
Sbjct: 137 GQMSHWGATV--ITNLFSAIPFV 157
>MBI3_YEAST (Q9ZZW7) Cytochrome b mRNA maturase bI3
Length = 517
Score = 33.1 bits (74), Expect = 1.3
Identities = 21/83 (25%), Positives = 42/83 (50%), Gaps = 14/83 (16%)
Query: 87 GCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLW-VGMTVIPFLLPSEFIQ---IY 142
G F+ +MF+ A+ ++ + S ++ LW VG+ + + + F+ +Y
Sbjct: 85 GASFFFMVMFMHMAK--------GLYYGSYRSPRVTLWNVGVIIFILTIATAFLGYCCVY 136
Query: 143 GEVAHFGAGVFLLIQLISIISFI 165
G+++H+GA V + L S I F+
Sbjct: 137 GQMSHWGATV--ITNLFSAIPFV 157
>CYB_YEAST (P00163) Cytochrome b
Length = 385
Score = 33.1 bits (74), Expect = 1.3
Identities = 21/83 (25%), Positives = 42/83 (50%), Gaps = 14/83 (16%)
Query: 87 GCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLW-VGMTVIPFLLPSEFIQ---IY 142
G F+ +MF+ A+ ++ + S ++ LW VG+ + + + F+ +Y
Sbjct: 85 GASFFFMVMFMHMAK--------GLYYGSYRSPRVTLWNVGVIIFILTIATAFLGYCCVY 136
Query: 143 GEVAHFGAGVFLLIQLISIISFI 165
G+++H+GA V + L S I F+
Sbjct: 137 GQMSHWGATV--ITNLFSAIPFV 157
>CYB_SACDO (Q35819) Cytochrome b
Length = 385
Score = 33.1 bits (74), Expect = 1.3
Identities = 21/83 (25%), Positives = 42/83 (50%), Gaps = 14/83 (16%)
Query: 87 GCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLW-VGMTVIPFLLPSEFIQ---IY 142
G F+ +MF+ A+ ++ + S ++ LW VG+ + + + F+ +Y
Sbjct: 85 GASFFFMVMFMHMAK--------GLYYGSYRSPRVTLWNVGVIIFILTIATAFLGYCCVY 136
Query: 143 GEVAHFGAGVFLLIQLISIISFI 165
G+++H+GA V + L S I F+
Sbjct: 137 GQMSHWGATV--ITNLFSAIPFV 157
>RNFD_PASMU (Q9CNP3) Electron transport complex protein rnfD
Length = 349
Score = 32.7 bits (73), Expect = 1.7
Identities = 44/150 (29%), Positives = 65/150 (43%), Gaps = 35/150 (23%)
Query: 113 HSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAGVFLLIQLISIISFITWLNDCC 172
HSG +++I+LWV + +LP+ +Q+Y +FG GV LIQ ISF L
Sbjct: 11 HSGKLTMRIMLWVILA----MLPAMGVQLY----YFGFGV--LIQAALAISFALCL---- 56
Query: 173 ESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIW--YAPKPSCLLNIFFIAWTLVLLQLM 230
E + LF + + V L +IL YAP + L+L+
Sbjct: 57 --EGIVTVLRQKPTLFYLSDFSVILTALILAMAIPPYAP------------YWLILIGTF 102
Query: 231 TSVSLHPKVNAG----ILTPGLMGLYVVFL 256
+V L V G + P ++G YVV L
Sbjct: 103 CAVILGKHVYGGLGQNLFNPAMVG-YVVLL 131
>CATV_NPVRO (Q8B9D5) Viral cathepsin (EC 3.4.22.50) (V-cath)
(Cysteine proteinase) (CP)
Length = 323
Score = 32.0 bits (71), Expect = 2.9
Identities = 16/51 (31%), Positives = 26/51 (50%), Gaps = 1/51 (1%)
Query: 317 IPAEDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTW 367
I A D V Y G + F +G + A+LL+G+ +++ WT W + W
Sbjct: 246 IDAADIVNYKQGIIKYCFNSGLNH-AVLLVGYGVENNIPYWTFKNTWGTDW 295
>CATV_NPVBM (P41721) Viral cathepsin (EC 3.4.22.50) (V-cath)
(Cysteine proteinase) (CP)
Length = 323
Score = 32.0 bits (71), Expect = 2.9
Identities = 16/51 (31%), Positives = 26/51 (50%), Gaps = 1/51 (1%)
Query: 317 IPAEDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTW 367
I A D V Y G + F +G + A+LL+G+ +++ WT W + W
Sbjct: 246 IDAADIVNYKQGIIKYCFDSGLNH-AVLLVGYGVENNIPYWTFKNTWGTDW 295
>CATV_NPVAC (P25783) Viral cathepsin (EC 3.4.22.50) (V-cath)
(Cysteine proteinase) (CP)
Length = 323
Score = 32.0 bits (71), Expect = 2.9
Identities = 16/51 (31%), Positives = 26/51 (50%), Gaps = 1/51 (1%)
Query: 317 IPAEDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTW 367
I A D V Y G + F +G + A+LL+G+ +++ WT W + W
Sbjct: 246 IDAADIVNYKQGIIKYCFNSGLNH-AVLLVGYGVENNIPYWTFKNTWGTDW 295
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.328 0.140 0.472
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,522,064
Number of Sequences: 164201
Number of extensions: 2008009
Number of successful extensions: 5396
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 5345
Number of HSP's gapped (non-prelim): 37
length of query: 398
length of database: 59,974,054
effective HSP length: 112
effective length of query: 286
effective length of database: 41,583,542
effective search space: 11892893012
effective search space used: 11892893012
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0214.6


