

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0214.10
(74 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
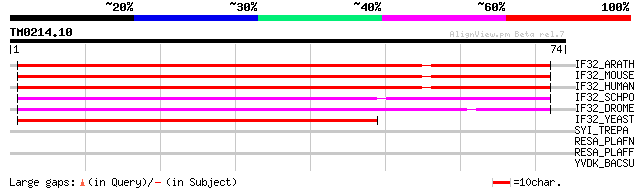
Score E
Sequences producing significant alignments: (bits) Value
IF32_ARATH (Q38884) Eukaryotic translation initiation factor 3 s... 74 9e-14
IF32_MOUSE (Q9QZD9) Eukaryotic translation initiation factor 3 s... 55 3e-08
IF32_HUMAN (Q13347) Eukaryotic translation initiation factor 3 s... 55 3e-08
IF32_SCHPO (P79083) Eukaryotic translation initiation factor 3 3... 55 4e-08
IF32_DROME (O02195) Eukaryotic translation initiation factor 3 s... 54 6e-08
IF32_YEAST (P40217) Eukaryotic translation initiation factor 3 3... 49 3e-06
SYI_TREPA (O83466) Isoleucyl-tRNA synthetase (EC 6.1.1.5) (Isole... 27 7.3
RESA_PLAFN (P13831) Ring-infected erythrocyte surface antigen (F... 27 7.3
RESA_PLAFF (P13830) Ring-infected erythrocyte surface antigen pr... 27 7.3
YVDK_BACSU (O06993) Hypothetical glycosyl hydrolase yvdK (EC 3.2... 27 9.6
>IF32_ARATH (Q38884) Eukaryotic translation initiation factor 3
subunit 2 (eIF-3 beta) (eIF3 p36) (eIF3i) (TGF-beta
receptor interacting protein 1) (TRIP-1)
Length = 328
Score = 73.6 bits (179), Expect = 9e-14
Identities = 40/71 (56%), Positives = 48/71 (67%), Gaps = 1/71 (1%)
Query: 2 VLRGGQVASAVSTTDHHAGNFEAKFYDMILQEEFRDVKGHSGLINVMAFIMEERPALSDP 61
VL GGQ ASAV+TTDH AG FEAKFYD ILQEE VKGH G IN +AF + + + S
Sbjct: 250 VLGGGQDASAVTTTDHRAGKFEAKFYDKILQEEIGGVKGHFGPINALAFNPDGK-SFSSG 308
Query: 62 VSDGHVLVQFY 72
DG+V + +
Sbjct: 309 GEDGYVRLHHF 319
>IF32_MOUSE (Q9QZD9) Eukaryotic translation initiation factor 3
subunit 2 (eIF-3 beta) (eIF3 p36) (eIF3i) (TGF-beta
receptor interacting protein 1) (TRIP-1)
Length = 325
Score = 55.5 bits (132), Expect = 3e-08
Identities = 30/71 (42%), Positives = 43/71 (60%), Gaps = 1/71 (1%)
Query: 2 VLRGGQVASAVSTTDHHAGNFEAKFYDMILQEEFRDVKGHSGLINVMAFIMEERPALSDP 61
VL GGQ A V+TT G FEA+F+ + +EEF VKGH G IN +AF + + + S
Sbjct: 245 VLGGGQEAMDVTTTSTRIGKFEARFFHLAFEEEFGRVKGHFGPINSVAFHPDGK-SYSSG 303
Query: 62 VSDGHVLVQFY 72
DG+V + ++
Sbjct: 304 GEDGYVRIHYF 314
>IF32_HUMAN (Q13347) Eukaryotic translation initiation factor 3
subunit 2 (eIF-3 beta) (eIF3 p36) (eIF3i) (TGF-beta
receptor interacting protein 1) (TRIP-1)
Length = 325
Score = 55.5 bits (132), Expect = 3e-08
Identities = 30/71 (42%), Positives = 43/71 (60%), Gaps = 1/71 (1%)
Query: 2 VLRGGQVASAVSTTDHHAGNFEAKFYDMILQEEFRDVKGHSGLINVMAFIMEERPALSDP 61
VL GGQ A V+TT G FEA+F+ + +EEF VKGH G IN +AF + + + S
Sbjct: 245 VLGGGQEAMDVTTTSTRIGKFEARFFHLAFEEEFGRVKGHFGPINSVAFHPDGK-SYSSG 303
Query: 62 VSDGHVLVQFY 72
DG+V + ++
Sbjct: 304 GEDGYVRIHYF 314
>IF32_SCHPO (P79083) Eukaryotic translation initiation factor 3 39
kDa subunit (eIF3 p39) (Translation initiation factor
eIF3, p39 subunit) (Suppressor of uncontrolled mitosis
1)
Length = 328
Score = 54.7 bits (130), Expect = 4e-08
Identities = 31/71 (43%), Positives = 41/71 (57%), Gaps = 1/71 (1%)
Query: 2 VLRGGQVASAVSTTDHHAGNFEAKFYDMILQEEFRDVKGHSGLINVMAFIMEERPALSDP 61
+L GGQ A V+TT G FEA+FY IL+EE VKGH G IN +A + + +
Sbjct: 248 ILGGGQEARDVTTTAARQGKFEARFYHAILEEELGRVKGHFGPINTIA-VHPKGTGYASG 306
Query: 62 VSDGHVLVQFY 72
DG+V V F+
Sbjct: 307 GEDGYVRVHFF 317
>IF32_DROME (O02195) Eukaryotic translation initiation factor 3
subunit 2 (eIF-3 beta) (eIF3i) (TRIP-1 homolog)
Length = 326
Score = 54.3 bits (129), Expect = 6e-08
Identities = 31/71 (43%), Positives = 43/71 (59%), Gaps = 1/71 (1%)
Query: 2 VLRGGQVASAVSTTDHHAGNFEAKFYDMILQEEFRDVKGHSGLINVMAFIMEERPALSDP 61
VL GGQ A V+TT AG F+++F+ +I +EEF +KGH G IN +AF + + S
Sbjct: 247 VLGGGQDAMEVTTTSTKAGKFDSRFFHLIYEEEFARLKGHFGPINSLAFHPDGKSYASGG 306
Query: 62 VSDGHVLVQFY 72
DG V VQ +
Sbjct: 307 -EDGFVRVQTF 316
>IF32_YEAST (P40217) Eukaryotic translation initiation factor 3 39
kDa subunit (eIF3 p39) (Translation initiation factor
eIF3, p39 subunit)
Length = 347
Score = 48.5 bits (114), Expect = 3e-06
Identities = 23/48 (47%), Positives = 31/48 (63%)
Query: 2 VLRGGQVASAVSTTDHHAGNFEAKFYDMILQEEFRDVKGHSGLINVMA 49
+L GGQ A V+TT + G FEA+FY I +EE V+GH G +N +A
Sbjct: 253 ILGGGQEAKDVTTTSANEGKFEARFYHKIFEEEIGRVQGHFGPLNTVA 300
>SYI_TREPA (O83466) Isoleucyl-tRNA synthetase (EC 6.1.1.5)
(Isoleucine--tRNA ligase) (IleRS)
Length = 1091
Score = 27.3 bits (59), Expect = 7.3
Identities = 15/51 (29%), Positives = 24/51 (46%)
Query: 18 HAGNFEAKFYDMILQEEFRDVKGHSGLINVMAFIMEERPALSDPVSDGHVL 68
HA +FE F + E +G + ++A + ERPA + + G VL
Sbjct: 569 HATDFERYFPAHFISEGLDQTRGWFYTLTILAVALFERPAFENCIVTGLVL 619
>RESA_PLAFN (P13831) Ring-infected erythrocyte surface antigen
(Fragment)
Length = 760
Score = 27.3 bits (59), Expect = 7.3
Identities = 12/46 (26%), Positives = 26/46 (56%)
Query: 26 FYDMILQEEFRDVKGHSGLINVMAFIMEERPALSDPVSDGHVLVQF 71
FY + E+F+D G ++ ++ F E+R +++D + L++F
Sbjct: 388 FYLLSSLEKFKDFTGTPQIVTLLRFFFEKRLSMNDLENKSEHLLKF 433
>RESA_PLAFF (P13830) Ring-infected erythrocyte surface antigen
precursor
Length = 1073
Score = 27.3 bits (59), Expect = 7.3
Identities = 12/46 (26%), Positives = 26/46 (56%)
Query: 26 FYDMILQEEFRDVKGHSGLINVMAFIMEERPALSDPVSDGHVLVQF 71
FY + E+F+D G ++ ++ F E+R +++D + L++F
Sbjct: 602 FYLLSSLEKFKDFTGTPQIVTLLRFFFEKRLSMNDLENKSEHLLKF 647
>YVDK_BACSU (O06993) Hypothetical glycosyl hydrolase yvdK (EC
3.2.1.-)
Length = 757
Score = 26.9 bits (58), Expect = 9.6
Identities = 13/43 (30%), Positives = 20/43 (46%)
Query: 1 EVLRGGQVASAVSTTDHHAGNFEAKFYDMILQEEFRDVKGHSG 43
E L G V + V D + N ++ + + QEE + HSG
Sbjct: 158 ECLTGDAVITLVPYLDGNVANEDSNYQEQFWQEEAKGADSHSG 200
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.137 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,415,529
Number of Sequences: 164201
Number of extensions: 258568
Number of successful extensions: 511
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 501
Number of HSP's gapped (non-prelim): 10
length of query: 74
length of database: 59,974,054
effective HSP length: 50
effective length of query: 24
effective length of database: 51,764,004
effective search space: 1242336096
effective search space used: 1242336096
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0214.10


