

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
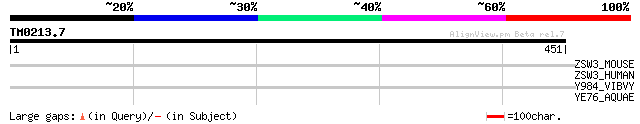
Query= TM0213.7
(451 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ZSW3_MOUSE (Q8CFL8) Zinc finger SWIM domain containing protein 3 37 0.14
ZSW3_HUMAN (Q96MP5) Zinc finger SWIM domain containing protein 3 35 0.51
Y984_VIBVY (Q7MMT3) Hypothetical UPF0319 protein VV0984 precursor 31 7.4
YE76_AQUAE (O67453) Hypothetical protein AQ_1476 30 9.7
>ZSW3_MOUSE (Q8CFL8) Zinc finger SWIM domain containing protein 3
Length = 695
Score = 36.6 bits (83), Expect = 0.14
Identities = 25/85 (29%), Positives = 41/85 (47%), Gaps = 14/85 (16%)
Query: 318 EVKHMVSQDQFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYM 377
E H VS+D +CSC+F + +PCRH +A + H+SQ+P R +
Sbjct: 536 ENSHQVSKDGC-------SCSCSFQQCYHLPCRHILALL-HTSQQPVGEAMVCCR---WQ 584
Query: 378 ATYGHVITP---INGQKLWPTTNDP 399
Y H++ P + + + P T+ P
Sbjct: 585 KKYQHLLDPNGELQDRGIIPNTDQP 609
>ZSW3_HUMAN (Q96MP5) Zinc finger SWIM domain containing protein 3
Length = 696
Score = 34.7 bits (78), Expect = 0.51
Identities = 18/46 (39%), Positives = 27/46 (58%), Gaps = 8/46 (17%)
Query: 318 EVKHMVSQDQFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRP 363
E H VS+D +CSC+F + +PCRH +A + H+SQ+P
Sbjct: 537 EDSHQVSKDGC-------SCSCSFQQWYHLPCRHILALL-HTSQQP 574
>Y984_VIBVY (Q7MMT3) Hypothetical UPF0319 protein VV0984 precursor
Length = 245
Score = 30.8 bits (68), Expect = 7.4
Identities = 12/45 (26%), Positives = 23/45 (50%)
Query: 283 EKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
EK + +P+ +E +EMV+YW EDK + + +++
Sbjct: 162 EKLSQSIAEQPIVEMEKGVEMVQYWYEKASNEDKKQFASLAFENR 206
>YE76_AQUAE (O67453) Hypothetical protein AQ_1476
Length = 180
Score = 30.4 bits (67), Expect = 9.7
Identities = 14/43 (32%), Positives = 26/43 (59%), Gaps = 2/43 (4%)
Query: 271 IMGRFATMNEKLEKYKG--EVMPKPLKRLEHEIEMVRYWVATR 311
+ G + E++EK K +++PK ++ LE E+E VR +A +
Sbjct: 14 VTGEIEKLRERIEKVKETLDLIPKEIEELERELERVRQEIAKK 56
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.341 0.149 0.525
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 51,848,273
Number of Sequences: 164201
Number of extensions: 2126823
Number of successful extensions: 6448
Number of sequences better than 10.0: 4
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 6445
Number of HSP's gapped (non-prelim): 4
length of query: 451
length of database: 59,974,054
effective HSP length: 114
effective length of query: 337
effective length of database: 41,255,140
effective search space: 13902982180
effective search space used: 13902982180
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0213.7


