

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.1
(305 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
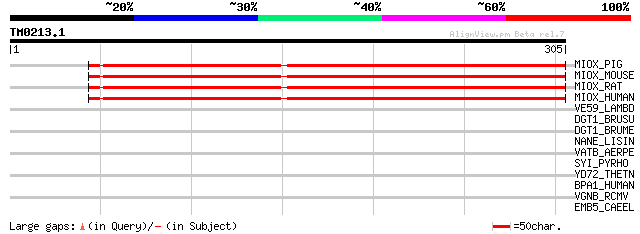
MIOX_PIG (Q8WN98) Inositol oxygenase (EC 1.13.99.1) (Myo-inosito... 270 3e-72
MIOX_MOUSE (Q9QXN5) Inositol oxygenase (EC 1.13.99.1) (Myo-inosi... 268 1e-71
MIOX_RAT (Q9QXN4) Inositol oxygenase (EC 1.13.99.1) (Myo-inosito... 266 5e-71
MIOX_HUMAN (Q9UGB7) Inositol oxygenase (EC 1.13.99.1) (Myo-inosi... 261 2e-69
VE59_LAMBD (P03754) EA59 gene protein 35 0.23
DGT1_BRUSU (P63934) Deoxyguanosinetriphosphate triphosphohydrola... 32 1.5
DGT1_BRUME (P63933) Deoxyguanosinetriphosphate triphosphohydrola... 32 1.5
NANE_LISIN (Q926V7) Putative N-acetylmannosamine-6-phosphate 2-e... 31 3.3
VATB_AERPE (Q9YF36) V-type ATP synthase beta chain (EC 3.6.3.14)... 30 5.7
SYI_PYRHO (O58792) Isoleucyl-tRNA synthetase (EC 6.1.1.5) (Isole... 30 5.7
YD72_THETN (Q8RA57) Hypothetical UPF0144 protein TTE1372 30 7.4
BPA1_HUMAN (Q03001) Bullous pemphigoid antigen 1 isoforms 1/2/3/... 30 7.4
VGNB_RCMV (P35930) Genome polyprotein B [Contains: Protease cofa... 30 9.7
EMB5_CAEEL (P34703) Abnormal embryogenesis protein 5 30 9.7
>MIOX_PIG (Q8WN98) Inositol oxygenase (EC 1.13.99.1) (Myo-inositol
oxygenase) (Aldehyde reductase-like 6)
Length = 282
Score = 270 bits (690), Expect = 3e-72
Identities = 131/263 (49%), Positives = 174/263 (65%), Gaps = 5/263 (1%)
Query: 44 TFRDYNAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVD 103
+FR+Y + + V Y+ H +Q+VDFV+K ++G + M++ E ++L+ +VD
Sbjct: 24 SFRNYTS-GPLLDRVFRTYKLMHTWQTVDFVRKKHAQFGGFSYKRMTVLEAVDMLDGLVD 82
Query: 104 ESDPDLDEPQIEHLLQTAEAIRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGD 163
ESDPD+D P H QTAE IRK +P++DW HL GL+HDLGKVL+L G PQWAVVGD
Sbjct: 83 ESDPDVDFPNSFHAFQTAEGIRKAHPDKDWFHLVGLLHDLGKVLVL---AGEPQWAVVGD 139
Query: 164 TFPVGCRFDESIVH-HKHFKENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVA 222
TFPVGCR S+V F++NPD +P Y+T GMY CGL N LMSWGHD+YMY +
Sbjct: 140 TFPVGCRPQASVVFCDSTFQDNPDLQDPVYSTELGMYQPHCGLENALMSWGHDEYMYQMM 199
Query: 223 KENKTTLPSAALFIIRYHSFYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRI 282
K NK +LP A +IIR+HSFY H G Y+ L NE+D L W+ FNK+DLY+K
Sbjct: 200 KFNKFSLPGEAFYIIRFHSFYPWHTGGDYRQLCNEQDLAMLPWVQEFNKFDLYTKGSDMP 259
Query: 283 DVEKVKPYYLSLIEKYFPAKLKW 305
DV++++PYY LI+KY P L W
Sbjct: 260 DVDELRPYYQGLIDKYCPGVLCW 282
>MIOX_MOUSE (Q9QXN5) Inositol oxygenase (EC 1.13.99.1) (Myo-inositol
oxygenase) (Aldehyde reductase-like 6) (Renal-specific
oxidoreductase)
Length = 285
Score = 268 bits (686), Expect = 1e-71
Identities = 129/263 (49%), Positives = 176/263 (66%), Gaps = 5/263 (1%)
Query: 44 TFRDYNAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVD 103
+FR+Y + + V Y+ H +Q+VDFV + R +YG + +M++ E +L+++VD
Sbjct: 27 SFRNYTS-GPLLDRVFTTYKLMHTHQTVDFVSRKRIQYGGFSYKKMTIMEAVGMLDDLVD 85
Query: 104 ESDPDLDEPQIEHLLQTAEAIRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGD 163
ESDPD+D P H QTAE IRK +P++DW HL GL+HDLGK++ L G PQWAVVGD
Sbjct: 86 ESDPDVDFPNSFHAFQTAEGIRKAHPDKDWFHLVGLLHDLGKIMAL---WGEPQWAVVGD 142
Query: 164 TFPVGCRFDESIVH-HKHFKENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVA 222
TFPVGCR S+V F++NPD +P Y+T GMY CGL NVLMSWGHD+Y+Y +
Sbjct: 143 TFPVGCRPQASVVFCDSTFQDNPDLQDPRYSTELGMYQPHCGLENVLMSWGHDEYLYQMM 202
Query: 223 KENKTTLPSAALFIIRYHSFYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRI 282
K NK +LPS A ++IR+HSFY H G Y+ L +++D + L W+ FNK+DLY+K
Sbjct: 203 KFNKFSLPSEAFYMIRFHSFYPWHTGGDYRQLCSQQDLDMLPWVQEFNKFDLYTKCPDLP 262
Query: 283 DVEKVKPYYLSLIEKYFPAKLKW 305
DVE ++PYY LI+KY P L W
Sbjct: 263 DVESLRPYYQGLIDKYCPGTLSW 285
>MIOX_RAT (Q9QXN4) Inositol oxygenase (EC 1.13.99.1) (Myo-inositol
oxygenase) (Aldehyde reductase-like 6) (Renal-specific
oxidoreductase) (Kidney-specific protein 32)
Length = 285
Score = 266 bits (680), Expect = 5e-71
Identities = 126/263 (47%), Positives = 177/263 (66%), Gaps = 5/263 (1%)
Query: 44 TFRDYNAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVD 103
+FR+Y + + V Y+ H +Q+VDFV + R ++G + +M++ E ++L+++VD
Sbjct: 27 SFRNYTS-GPLLDRVFTTYKLMHTHQTVDFVMRKRIQFGSFSYKKMTVMEAVDMLDDLVD 85
Query: 104 ESDPDLDEPQIEHLLQTAEAIRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGD 163
ESDPD+D P H QTAE IRK +P++DW HL GL+HDLGK+L L G PQWAVVGD
Sbjct: 86 ESDPDVDFPNSFHAFQTAEGIRKAHPDKDWFHLVGLLHDLGKILAL---WGEPQWAVVGD 142
Query: 164 TFPVGCRFDESIVH-HKHFKENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVA 222
TFPVGCR S+V F++NPD +P Y+T GMY CGL NVLMSWGHD+Y+Y +
Sbjct: 143 TFPVGCRPQASVVFCDSTFQDNPDLQDPRYSTELGMYQPHCGLENVLMSWGHDEYLYQMM 202
Query: 223 KENKTTLPSAALFIIRYHSFYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRI 282
K NK +LPS A +++R+HSFY H G Y+ L +++D + L W+ FNK+DLY+K
Sbjct: 203 KFNKFSLPSEAFYMVRFHSFYPWHTGGDYRQLCSQQDLDMLPWVQEFNKFDLYTKCPDLP 262
Query: 283 DVEKVKPYYLSLIEKYFPAKLKW 305
+V+ ++PYY LI+KY P L W
Sbjct: 263 EVKSLRPYYQGLIDKYCPGTLSW 285
>MIOX_HUMAN (Q9UGB7) Inositol oxygenase (EC 1.13.99.1) (Myo-inositol
oxygenase) (Aldehyde reductase-like 6) (Renal-specific
oxidoreductase) (Kidney-specific protein 32)
Length = 285
Score = 261 bits (667), Expect = 2e-69
Identities = 129/263 (49%), Positives = 173/263 (65%), Gaps = 5/263 (1%)
Query: 44 TFRDYNAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVD 103
+FR+Y + + V Y+ H +Q+VDFV+ ++G + +M++ E +LL+ +VD
Sbjct: 27 SFRNYTS-GPLLDRVFTTYKLMHTHQTVDFVRSKHAQFGGFSYKKMTVMEAVDLLDGLVD 85
Query: 104 ESDPDLDEPQIEHLLQTAEAIRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGD 163
ESDPD+D P H QTAE IRK +P++DW HL GL+HDLGKVL L G PQWAVVGD
Sbjct: 86 ESDPDVDFPNSFHAFQTAEGIRKAHPDKDWFHLVGLLHDLGKVLAL---FGEPQWAVVGD 142
Query: 164 TFPVGCRFDESIVH-HKHFKENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVA 222
TFPVGCR S+V F++NPD +P Y+T GMY CGL+ VLMSWGHD+YMY V
Sbjct: 143 TFPVGCRPQASVVFCDSTFQDNPDLQDPRYSTELGMYQPHCGLDRVLMSWGHDEYMYQVM 202
Query: 223 KENKTTLPSAALFIIRYHSFYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRI 282
K NK +LP A ++IR+HSFY H Y+ L +++D L W+ FNK+DLY+K
Sbjct: 203 KFNKFSLPPEAFYMIRFHSFYPWHTGRDYQQLCSQQDLAMLPWVREFNKFDLYTKCPDLP 262
Query: 283 DVEKVKPYYLSLIEKYFPAKLKW 305
DV+K++PYY LI+KY P L W
Sbjct: 263 DVDKLRPYYQGLIDKYCPGILSW 285
>VE59_LAMBD (P03754) EA59 gene protein
Length = 525
Score = 35.0 bits (79), Expect = 0.23
Identities = 32/119 (26%), Positives = 55/119 (45%), Gaps = 16/119 (13%)
Query: 182 KENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVAKENKTTLPSAALFIIRYHS 241
KE PD PA T+Y K +N L S G + + SA + +R
Sbjct: 256 KEQPD---PAKGTQYFYIGLKNAASNSLKSLG----------DLRLEFISAFIGCMRVDR 302
Query: 242 FYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKS--KVRIDVEKVKPYYLSLIEKY 298
L E A K L ++E+F N+E + + +KY+ ++ ++++D +K + I+KY
Sbjct: 303 KRQLWLE-AIKKLSSDENFSNMELISLISKYEELRRNEPQIQVDDDKFTKLFYDNIQKY 360
>DGT1_BRUSU (P63934) Deoxyguanosinetriphosphate
triphosphohydrolase-like protein
Length = 402
Score = 32.3 bits (72), Expect = 1.5
Identities = 18/58 (31%), Positives = 28/58 (48%), Gaps = 1/58 (1%)
Query: 39 NSFGHTFRDYNAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCE 96
+ FGHT + E E ++NF +H QS+ V K+ Y + + +S WE E
Sbjct: 105 HDFGHTPFGHTGEDALNERMKNFGGFDHNAQSLRIVTKLEHRYADFDGLNLS-WETLE 161
>DGT1_BRUME (P63933) Deoxyguanosinetriphosphate
triphosphohydrolase-like protein
Length = 402
Score = 32.3 bits (72), Expect = 1.5
Identities = 18/58 (31%), Positives = 28/58 (48%), Gaps = 1/58 (1%)
Query: 39 NSFGHTFRDYNAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCE 96
+ FGHT + E E ++NF +H QS+ V K+ Y + + +S WE E
Sbjct: 105 HDFGHTPFGHTGEDALNERMKNFGGFDHNAQSLRIVTKLEHRYADFDGLNLS-WETLE 161
>NANE_LISIN (Q926V7) Putative N-acetylmannosamine-6-phosphate
2-epimerase (EC 5.1.3.9) (ManNAc-6-P epimerase)
Length = 231
Score = 31.2 bits (69), Expect = 3.3
Identities = 27/102 (26%), Positives = 43/102 (41%), Gaps = 12/102 (11%)
Query: 43 HTFRDYNA-ESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEV 101
+T +D A +SE + Y+K++ V ++E + E CE E+
Sbjct: 49 NTAKDIRAIQSEVDVPIIGIYKKDYDDSDVFITPTLKE-----------VREICETGVEI 97
Query: 102 VDESDPDLDEPQIEHLLQTAEAIRKDYPNEDWLHLTGLIHDL 143
V P E L AIRK++PN ++ TG I D+
Sbjct: 98 VAMDATTRKRPHNEDLKDILSAIRKEFPNTLFMADTGSIEDV 139
>VATB_AERPE (Q9YF36) V-type ATP synthase beta chain (EC 3.6.3.14)
(V-type ATPase subunit B)
Length = 466
Score = 30.4 bits (67), Expect = 5.7
Identities = 21/77 (27%), Positives = 29/77 (37%)
Query: 13 SDVEGKTVPIDEKELVLDGGFVMPHTNSFGHTFRDYNAESERQEGVENFYRKNHIYQSVD 72
S ++G + ++L + G +PH R E +E F Y
Sbjct: 139 SAIDGMNTLVRGQKLPIFSGAGLPHNRLAAQIARQATVRGEEEEFAVVFSAIGIKYDDFL 198
Query: 73 FVKKMREEYGKLNRVEM 89
F KK EE G L RV M
Sbjct: 199 FFKKFFEETGALGRVAM 215
>SYI_PYRHO (O58792) Isoleucyl-tRNA synthetase (EC 6.1.1.5)
(Isoleucine--tRNA ligase) (IleRS)
Length = 1066
Score = 30.4 bits (67), Expect = 5.7
Identities = 13/37 (35%), Positives = 24/37 (64%), Gaps = 1/37 (2%)
Query: 45 FRDYNAESERQEGVENFYRKNHIYQSVDFVKKMREEY 81
FRDY + +E +E F+++N+IYQ + ++K +Y
Sbjct: 7 FRDYTP-GKLEEKIEEFWKENNIYQKIKELRKNGPKY 42
>YD72_THETN (Q8RA57) Hypothetical UPF0144 protein TTE1372
Length = 523
Score = 30.0 bits (66), Expect = 7.4
Identities = 27/106 (25%), Positives = 57/106 (53%), Gaps = 7/106 (6%)
Query: 49 NAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLN---EVVDES 105
+AE++++E + + H +S DF K++R+ G+L R+E + + E+L E +++
Sbjct: 60 DAENKKREALLEAKEEIHRLRS-DFEKEVRDRRGELQRLEKRLLQKEEILEKRAESLEQK 118
Query: 106 DPDLDEPQ--IEHLLQTAEAIRKDYPNEDWLHLTGLIHDLGKVLLL 149
+ L++ Q I+ L + + K E+ ++GL + K +LL
Sbjct: 119 EILLEQKQKEIQQLEEQITLLHKK-QLEELERISGLTQEEAKSILL 163
>BPA1_HUMAN (Q03001) Bullous pemphigoid antigen 1 isoforms 1/2/3/4/5/8
(230 kDa bullous pemphigoid antigen) (BPA)
(Hemidesmosomal plaque protein) (Dystonia musculorum
protein) (Dystonin) (Fragment)
Length = 3214
Score = 30.0 bits (66), Expect = 7.4
Identities = 22/77 (28%), Positives = 38/77 (48%), Gaps = 10/77 (12%)
Query: 37 HTNSFGHTFRDYNAESER-------QEGVENFYRKNH--IYQSVDFVKKMREEYGKLNRV 87
H NS H+FRD E ER + ++ + K+H + Q++ K+ ++ +LN
Sbjct: 2068 HLNSQIHSFRD-EKELERLQICQRKSDHLKEQFEKSHEQLLQNIKAEKENNDKIQRLNEE 2126
Query: 88 EMSMWECCELLNEVVDE 104
EC E+L + V+E
Sbjct: 2127 LEKSNECAEMLKQKVEE 2143
>VGNB_RCMV (P35930) Genome polyprotein B [Contains: Protease
cofactor; Membrane-binding protein; Viral genome-linked
protein (VPg); Protease (EC 3.4.22.-); RNA-directed RNA
polymerase (EC 2.7.7.48)]
Length = 1864
Score = 29.6 bits (65), Expect = 9.7
Identities = 10/27 (37%), Positives = 16/27 (59%)
Query: 239 YHSFYALHREGAYKHLMNEEDFENLEW 265
YH Y + R G+Y H+ +DF+ +W
Sbjct: 714 YHLLYVVPRNGSYLHVAANKDFQIQQW 740
>EMB5_CAEEL (P34703) Abnormal embryogenesis protein 5
Length = 1521
Score = 29.6 bits (65), Expect = 9.7
Identities = 25/103 (24%), Positives = 49/103 (47%), Gaps = 11/103 (10%)
Query: 33 FVMPHTNSFGHTFRDYNAESERQEGVENFYRKNHIYQSVDFVKK---MREEYGKLNRVEM 89
F+ +NSF TF + R+E ++N N++++ DF +K + E+ K+ +
Sbjct: 357 FIRVRSNSFEPTFIGFY----RKEDIDNLLTMNNLWRVYDFDEKWCHLSEKKNKIYDLMR 412
Query: 90 SMWECCELLNEVVDE----SDPDLDEPQIEHLLQTAEAIRKDY 128
M E EL +++ + SD DL + + L+ I ++
Sbjct: 413 RMREYQELSDDLTAKRRPISDADLMDTKYAETLEQLTDIHANF 455
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.139 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,094,177
Number of Sequences: 164201
Number of extensions: 1877453
Number of successful extensions: 4650
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 4633
Number of HSP's gapped (non-prelim): 14
length of query: 305
length of database: 59,974,054
effective HSP length: 110
effective length of query: 195
effective length of database: 41,911,944
effective search space: 8172829080
effective search space used: 8172829080
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0213.1


