

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0209.4
(342 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
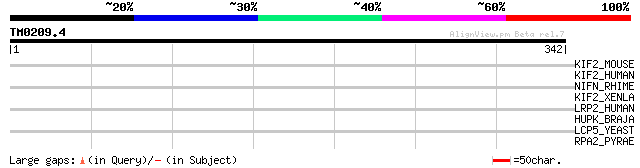
KIF2_MOUSE (P28740) Kinesin-like protein KIF2 35 0.36
KIF2_HUMAN (O00139) Kinesin-like protein KIF2 (Kinesin-2) (HK2) 35 0.36
NIFN_RHIME (P12781) Nitrogenase iron-molybdenum cofactor biosynt... 33 1.0
KIF2_XENLA (Q91637) Kinesin-like protein KIF2 (Kinesin-related p... 33 1.0
LRP2_HUMAN (P98164) Low-density lipoprotein receptor-related pro... 32 1.8
HUPK_BRAJA (P48342) Hydrogenase expression/formation protein hupK 31 3.9
LCP5_YEAST (P40079) U3 small nucleolar ribonucleoprotein protein... 30 6.7
RPA2_PYRAE (Q8ZYQ7) DNA-directed RNA polymerase subunit A'' (EC ... 30 8.8
>KIF2_MOUSE (P28740) Kinesin-like protein KIF2
Length = 716
Score = 34.7 bits (78), Expect = 0.36
Identities = 19/50 (38%), Positives = 31/50 (62%), Gaps = 5/50 (10%)
Query: 153 DRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDV--SART 200
D +RKTKL +E +Q +QVVG+++R++ CV+ +D+ S RT
Sbjct: 348 DLLNRKTKLRVLEDGKQQVQVVGLQEREVK---CVEDVLKLIDIGNSCRT 394
>KIF2_HUMAN (O00139) Kinesin-like protein KIF2 (Kinesin-2) (HK2)
Length = 679
Score = 34.7 bits (78), Expect = 0.36
Identities = 19/50 (38%), Positives = 31/50 (62%), Gaps = 5/50 (10%)
Query: 153 DRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDV--SART 200
D +RKTKL +E +Q +QVVG+++R++ CV+ +D+ S RT
Sbjct: 349 DLLNRKTKLRVLEDGKQQVQVVGLQEREVK---CVEDVLKLIDIGNSCRT 395
>NIFN_RHIME (P12781) Nitrogenase iron-molybdenum cofactor
biosynthesis protein nifN
Length = 441
Score = 33.1 bits (74), Expect = 1.0
Identities = 21/70 (30%), Positives = 33/70 (47%), Gaps = 6/70 (8%)
Query: 204 STLGILAKVPSLGGRIRPLGVGGALPIWRSGLASSSFATAIRAIMASSVCGVGGVEPLEP 263
+T+ L LGG + LGVGGA+P++ +SFA + PL+
Sbjct: 11 ATINPLKSSQPLGGALAFLGVGGAIPLFHGSQGCTSFALVLLVRHFKEAI------PLQT 64
Query: 264 VAVDLVSTIL 273
A+D V+ +L
Sbjct: 65 TAMDDVAIVL 74
>KIF2_XENLA (Q91637) Kinesin-like protein KIF2 (Kinesin-related
protein XKIF2)
Length = 682
Score = 33.1 bits (74), Expect = 1.0
Identities = 18/50 (36%), Positives = 31/50 (62%), Gaps = 5/50 (10%)
Query: 153 DRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDV--SART 200
D +RKTKL +E +Q +QVVG+++R++ CV+ +++ S RT
Sbjct: 351 DLLNRKTKLRVLEDGKQQVQVVGLQEREVK---CVEDVLKLIEIGNSCRT 397
>LRP2_HUMAN (P98164) Low-density lipoprotein receptor-related protein
2 precursor (Megalin) (Glycoprotein 330) (gp330)
Length = 4655
Score = 32.3 bits (72), Expect = 1.8
Identities = 21/68 (30%), Positives = 31/68 (44%), Gaps = 8/68 (11%)
Query: 28 PQKKLIVGDLPCPKMRNDNQSCDTTPTLTQSESRTINSLFTSMTSLLTVRVTDCSRHTDR 87
PQ + GD+ C ++NQ+C T T +++E FT L ++ C RH D
Sbjct: 2967 PQSWVCDGDVDCTDGYDENQNC-TRRTCSENE-------FTCGYGLCIPKIFRCDRHNDC 3018
Query: 88 GGLIDSPG 95
G D G
Sbjct: 3019 GDYSDERG 3026
>HUPK_BRAJA (P48342) Hydrogenase expression/formation protein hupK
Length = 362
Score = 31.2 bits (69), Expect = 3.9
Identities = 27/68 (39%), Positives = 35/68 (50%), Gaps = 8/68 (11%)
Query: 241 ATAIRAIMASSVCGVGGVEPLEPVAVDLVSTILLQCHTDRG--GLIDSHV---GHALIAS 295
A A+RA+M +S +GG P E V L + Q T G G+ + H G AL AS
Sbjct: 112 AAAMRAVMQASAL-LGG--PSEAVPQALRRDAVAQIRTAVGTLGISEDHALASGSALAAS 168
Query: 296 LPCCDGRL 303
+ CDGRL
Sbjct: 169 VEGCDGRL 176
>LCP5_YEAST (P40079) U3 small nucleolar ribonucleoprotein protein
LCP5
Length = 357
Score = 30.4 bits (67), Expect = 6.7
Identities = 13/35 (37%), Positives = 22/35 (62%)
Query: 135 GRDIELTSDKTKWQDAMADRFSRKTKLAWIESQRQ 169
G D + S K K +D+ + R ++KT+ AW +QR+
Sbjct: 322 GEDFGIFSSKRKLEDSTSRRGAKKTRSAWDRAQRR 356
>RPA2_PYRAE (Q8ZYQ7) DNA-directed RNA polymerase subunit A'' (EC
2.7.7.6)
Length = 379
Score = 30.0 bits (66), Expect = 8.8
Identities = 14/47 (29%), Positives = 27/47 (56%)
Query: 208 ILAKVPSLGGRIRPLGVGGALPIWRSGLASSSFATAIRAIMASSVCG 254
++A + GR+RP+G G + S LA ++F ++ ++ +SV G
Sbjct: 302 LVADAMTWSGRLRPIGRHGVVGSKESPLARAAFEVTVKTLIEASVRG 348
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,422,237
Number of Sequences: 164201
Number of extensions: 1629137
Number of successful extensions: 3313
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 3303
Number of HSP's gapped (non-prelim): 11
length of query: 342
length of database: 59,974,054
effective HSP length: 111
effective length of query: 231
effective length of database: 41,747,743
effective search space: 9643728633
effective search space used: 9643728633
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0209.4


