

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0209.3
(1547 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
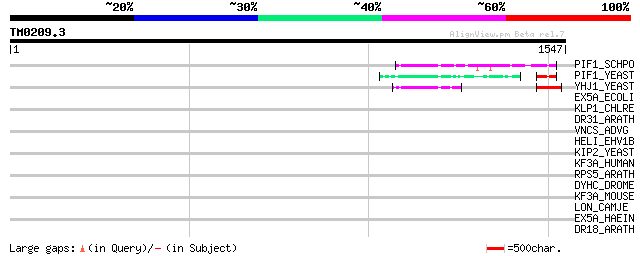
Score E
Sequences producing significant alignments: (bits) Value
PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1, m... 73 5e-12
PIF1_YEAST (P07271) DNA repair and recombination protein PIF1, m... 54 3e-06
YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergeni... 51 2e-05
EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 38 0.25
KLP1_CHLRE (P46870) Kinesin-like protein KLP1 36 0.72
DR31_ARATH (Q9FLB4) Putative disease resistance protein At5g05400 36 0.72
VNCS_ADVG (P24030) Noncapsid protein NS-1 (Nonstructural protein... 35 1.2
HELI_EHV1B (P28934) Probable helicase 35 1.6
KIP2_YEAST (P28743) Kinesin-like protein KIP2 35 2.1
KF3A_HUMAN (Q9Y496) Kinesin-like protein KIF3A (Microtubule plus... 34 3.6
RPS5_ARATH (O64973) Disease resistance protein RPS5 (Resistance ... 33 4.7
DYHC_DROME (P37276) Dynein heavy chain, cytosolic (DYHC) 33 4.7
KF3A_MOUSE (P28741) Kinesin-like protein KIF3A (Microtubule plus... 33 6.1
LON_CAMJE (O69300) ATP-dependent protease La (EC 3.4.21.53) 33 8.0
EX5A_HAEIN (P45158) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 33 8.0
DR18_ARATH (O64789) Putative disease resistance protein At1g61310 33 8.0
>PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1,
mitochondrial precursor
Length = 805
Score = 73.2 bits (178), Expect = 5e-12
Identities = 106/482 (21%), Positives = 202/482 (40%), Gaps = 68/482 (14%)
Query: 1075 KKVLDVVMSNKGGFLFLYGFGGTGKTFVWNTLSAAVRSK----GLIVLNVASSGIASLLL 1130
K++LD+V+ + +F G GTGK+ + + ++SK V AS+G+A+ +
Sbjct: 315 KRILDMVVEQQHS-IFFTGSAGTGKSVLLRKIIEVLKSKYRKQSDRVAVTASTGLAACNI 373
Query: 1131 PGGRTAHSRFSILISINEVS--TCNLRQGSPKAELLQKASLIIWDETPMMNKHCFEALDR 1188
GG T HS + ++ V +++ + ++I DE M++ + L+
Sbjct: 374 -GGVTLHSFAGVGLARESVDLLVSKIKKNKKCVNRWLRTRVLIIDEVSMVDAELMDKLEE 432
Query: 1189 SLNDVMKTRANYGNDRPFRGKVVVLGGDFRQILPVISKGSRADMVGSTVTSSYLWKYCK- 1247
+ K + +PF G +VL GDF Q+ PV G + + T WK
Sbjct: 433 VARVIRK------DSKPFGGIQLVLTGDFFQLPPVPENGKESKFCFESQT----WKSALD 482
Query: 1248 -VMKLTVNMRLQSASS*TSATEIREFAQWILKVGDETVDTIDEDETTIEIPSDLL----- 1301
+ LT R + E+R + K+ DE+V TIE LL
Sbjct: 483 FTIGLTHVFRQKDEEFVKMLNELR-----LGKLSDESVRKFKVLNRTIEYEDGLLPTELF 537
Query: 1302 -----IGQGPDPLLELVN---FAYPDLVANLESDSYFQERAI---LAPT---------LE 1341
+ + D ++ +N + + + D F++R + +AP +
Sbjct: 538 PTRYEVERSNDMRMQQINQNPVTFTAIDSGTVRDKEFRDRLLQGCMAPATLVLKVNAQVM 597
Query: 1342 SVVHVNNYILSKLPGVEREYLSYDTPCRSDEDSEVHAEWFTSEFLNDVQCSGIPNHRLIL 1401
+ ++++ +++ G ++ +T +D+E+ D+ +P+ +
Sbjct: 598 LIKNIDDQLVNGSLGKVIGFIDDETYQMEKKDAEMQGRNAFEYDSLDISPFDLPD---VK 654
Query: 1402 KERVPIMLLRNIDQAGGLCNGTRLRVTHLTQYIIVATVLSGIRLGKTEYIPRITLTPSDS 1461
+++ ++ +R R ++ + + IV + R T
Sbjct: 655 QKKYKLIAMRKASSTAIKWPLVRFKLPNGGERTIV--------------VQRETWNIELP 700
Query: 1462 GLPFKFSRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLK 1521
+ SR Q P+ L +A++I+K+QGQ+L V + L R VF GQ YVALSR +++GL+
Sbjct: 701 NGEVQASRSQIPLILAYAISIHKAQGQTLDRVKVDLGR-VFEKGQAYVALSRATTQEGLQ 759
Query: 1522 VL 1523
VL
Sbjct: 760 VL 761
>PIF1_YEAST (P07271) DNA repair and recombination protein PIF1,
mitochondrial precursor
Length = 857
Score = 53.9 bits (128), Expect = 3e-06
Identities = 31/56 (55%), Positives = 41/56 (72%), Gaps = 1/56 (1%)
Query: 1468 SRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKVL 1523
SR Q P+ L ++++I+KSQGQ+L V + L R VF GQ YVALSR SR+GL+VL
Sbjct: 690 SRVQLPLMLAWSLSIHKSQGQTLPKVKVDLRR-VFEKGQAYVALSRAVSREGLQVL 744
Score = 47.4 bits (111), Expect = 3e-04
Identities = 94/400 (23%), Positives = 157/400 (38%), Gaps = 80/400 (20%)
Query: 1031 PTFAEIMNFENKFVADELNYNKEEMSKIHEELVRSLTGEQKAAYKKVLDVVMSNKGGFLF 1090
P+F E+ N + D+ N N K+ ++ L+ EQ++ K ++ G +F
Sbjct: 209 PSFTELQNDQ-----DDSNLNPHNGVKV--KIPICLSKEQESIIK------LAENGHNIF 255
Query: 1091 LYGFGGTGKTFVWNTLSAAVRSKGLI----VLNVASSGIASLLLPGGRTAHSRFSILISI 1146
G GTGK+ + + + KG+ V AS+G+A+ + GG T HS F+ ++
Sbjct: 256 YTGSAGTGKSILLREMIKVL--KGIYGRENVAVTASTGLAACNI-GGITIHS-FAGILGK 311
Query: 1147 NEVSTCNLRQGSPKAELLQK---ASLIIWDETPMMNKHCFEALDRSLNDVMKTRANYGND 1203
+ + G + L++ ++ DE M++ + LD + K N
Sbjct: 312 GDADKLYKKVGRRSRKHLRRWENIGALVVDEISMLDAELLDKLDFIARKIRK------NH 365
Query: 1204 RPFRGKVVVLGGDFRQILPVISKGSRADMVGSTVTSSYLWKYCKVMKLTVNMRLQSASS* 1263
+PF G ++ GDF Q+ PV +R S WK + +K+T+ ++
Sbjct: 366 QPFGGIQLIFCGDFFQLPPVSKDPNRPT---KFAFESKAWK--EGVKMTIMLQ------- 413
Query: 1264 TSATEIREFAQWILKVGDETVDTIDEDETTIEIPSDLLIGQGPDPLLELVNFAYPDLVAN 1323
R ++I + + ID DET E + L
Sbjct: 414 -KVFRQRGDVKFIEMLNRMRLGNID-DETERE---------------------FKKLSRP 450
Query: 1324 LESDSYFQERAILAPTLESVVHVNNYILSKLPGVEREYLSYDTPCRSDEDSEVHAEWFTS 1383
L D A L T V NN LSKLPG + + D DE+ + E
Sbjct: 451 LPDDEIIP--AELYSTRMEVERANNSRLSKLPGQVHIFNAIDGGALEDEELK---ERLLQ 505
Query: 1384 EFLNDVQCSGIPNHRLILKERVPIMLLRNIDQAGGLCNGT 1423
FL + L LK +M+++N+D L NG+
Sbjct: 506 NFLAPKE--------LHLKVGAQVMMVKNLDAT--LVNGS 535
>YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergenic
region
Length = 723
Score = 51.2 bits (121), Expect = 2e-05
Identities = 58/200 (29%), Positives = 93/200 (46%), Gaps = 22/200 (11%)
Query: 1066 LTGEQKAAYKKVLDVVMSNKGGFLFLYGFGGTGKTFVWNTLSAAVRS---KGLIVLNVAS 1122
LT EQ+ +V+++++ + +F G GTGK+ + T+ + S K I + AS
Sbjct: 232 LTMEQE----RVVNLIVKKRTN-VFYTGSAGTGKSVILQTIIRQLSSLYGKESIAIT-AS 285
Query: 1123 SGIASLLLPGGRTAHSRFSILISINEVSTCNLRQGSPKAELL--QKASLIIWDETPMMNK 1180
+G+A++ + GG T H I I + + S K L + ++I DE M++
Sbjct: 286 TGLAAVTI-GGSTLHKWSGIGIGNKTIDQLVKKIQSQKDLLAAWRYTKVLIIDEISMVDG 344
Query: 1181 HCFEALDRSLNDVMKTRANYGNDRPFRGKVVVLGGDFRQILPVISKGSRADMVGSTVTSS 1240
+ + L++ + K ND PF G +VL GDF Q+ PV K V S
Sbjct: 345 NLLDKLEQIARRIRK------NDDPFGGIQLVLTGDFFQLPPVAKKDEH--NVVKFCFES 396
Query: 1241 YLWKYC--KVMKLTVNMRLQ 1258
+WK C K + LT R Q
Sbjct: 397 EMWKRCIQKTILLTKVFRQQ 416
Score = 48.5 bits (114), Expect = 1e-04
Identities = 28/68 (41%), Positives = 45/68 (66%), Gaps = 2/68 (2%)
Query: 1469 RRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKVLIVVEQ 1528
R Q P+ LC+A++I+K+QGQ++ + + L R +F GQ+YVALSR + L+VL +
Sbjct: 646 RTQIPLMLCWALSIHKAQGQTIQRLKVDL-RRIFEAGQVYVALSRAVTMDTLQVL-NFDP 703
Query: 1529 GVVSTSTR 1536
G + T+ R
Sbjct: 704 GKIRTNER 711
>EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC 3.1.11.5)
(Exodeoxyribonuclease V 67 kDa polypeptide)
Length = 608
Score = 37.7 bits (86), Expect = 0.25
Identities = 37/133 (27%), Positives = 53/133 (39%), Gaps = 22/133 (16%)
Query: 1406 PIMLLRNIDQAGGLCNGTRLRVTHLTQYIIVATVLSGIRLGKTEYIPRITLTPSDSGLPF 1465
P+M+ RN D A GL NG GI L + + R+ D +
Sbjct: 481 PVMIARN-DSALGLFNGD-----------------IGIALDRGQGT-RVWFAMPDGNIKS 521
Query: 1466 KFSRRQFPVTLCFAMTINKSQGQSLSHVGLYLP---RPVFTHGQLYVALSRVKSRKGLKV 1522
R +AMT++KSQG H L LP PV T +Y A++R + R L
Sbjct: 522 VQPSRLPEHETTWAMTVHKSQGSEFDHAALILPSQRTPVVTRELVYTAVTRARRRLSLYA 581
Query: 1523 LIVVEQGVVSTST 1535
+ ++T T
Sbjct: 582 DERILSAAIATRT 594
>KLP1_CHLRE (P46870) Kinesin-like protein KLP1
Length = 776
Score = 36.2 bits (82), Expect = 0.72
Identities = 25/84 (29%), Positives = 43/84 (50%), Gaps = 6/84 (7%)
Query: 1037 MNFENKFVADELNYNKEEMSKIHEELVRSLTGEQKAAYK----KVLDVVMSNKGGFLFLY 1092
+N A +N +E+ S + ++ +++ Q+AAY +V+D +M+ G +F Y
Sbjct: 33 VNVPKDLSAGPVNNQQEQFSFKFDGVLENVS--QEAAYTTLAHEVVDSLMAGYHGTIFAY 90
Query: 1093 GFGGTGKTFVWNTLSAAVRSKGLI 1116
G G GKTF + A +GLI
Sbjct: 91 GQTGAGKTFTMSGGGTAYAHRGLI 114
>DR31_ARATH (Q9FLB4) Putative disease resistance protein At5g05400
Length = 874
Score = 36.2 bits (82), Expect = 0.72
Identities = 35/126 (27%), Positives = 54/126 (42%), Gaps = 12/126 (9%)
Query: 996 LQIDDDELMNLCLIEIEKELQCNARSLK--DFPSLSYPTFAEIMNFENKFVADELNYNKE 1053
L D+E+ NLC + C+ R D+ ++ N +K V DE+ K
Sbjct: 89 LSQSDEEIDNLCCGQY-----CSKRCKYSYDYSKSVINKLQDVENLLSKGVFDEVA-QKG 142
Query: 1054 EMSKIHEELVRSLTGEQKAAYKKVLDVVMSNKGGFLFLYGFGGTGKTFVWNTLSAAVRSK 1113
+ K+ E L Q+A + + +M G L +YG GG GKT TL + + +K
Sbjct: 143 PIPKVEERLFHQEIVGQEAIVESTWNSMMEVGVGLLGIYGMGGVGKT----TLLSQINNK 198
Query: 1114 GLIVLN 1119
V N
Sbjct: 199 FRTVSN 204
>VNCS_ADVG (P24030) Noncapsid protein NS-1 (Nonstructural protein NS1)
(NCVP1)
Length = 590
Score = 35.4 bits (80), Expect = 1.2
Identities = 21/72 (29%), Positives = 40/72 (55%), Gaps = 4/72 (5%)
Query: 1048 LNYNKEEMSKIHEELVRSLTGEQKAAYKKVLDVVMSNKGG---FLFLYGFGGTGKTFVWN 1104
+ +N+EE K ++ + G + KKVL +++ +GG ++ YG GGTGKT + +
Sbjct: 385 IKFNEEEDDKPLLATIKDM-GLNEQYLKKVLCTILTKQGGKRGCIWFYGPGGTGKTLLAS 443
Query: 1105 TLSAAVRSKGLI 1116
+ A + G++
Sbjct: 444 LICKATVNYGMV 455
>HELI_EHV1B (P28934) Probable helicase
Length = 881
Score = 35.0 bits (79), Expect = 1.6
Identities = 19/44 (43%), Positives = 26/44 (58%)
Query: 1479 AMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKV 1522
AMTI +SQG SL V + PR +YVA+SR S + L++
Sbjct: 806 AMTIARSQGLSLDKVAICFPRNNLRINSVYVAMSRTVSSRFLRM 849
>KIP2_YEAST (P28743) Kinesin-like protein KIP2
Length = 706
Score = 34.7 bits (78), Expect = 2.1
Identities = 22/75 (29%), Positives = 39/75 (51%), Gaps = 5/75 (6%)
Query: 1075 KKVLDVVMSNKGGFLFLYGFGGTGKTFVWNTLSAAVRSKGLIVLNVASSGIASLLLPGGR 1134
K ++D ++ +F YG G+GKTF T+S + GLI L+V S + + ++
Sbjct: 184 KPMIDKLLMGFNATIFAYGMTGSGKTF---TMSGNEQELGLIPLSV--SYLFTNIMEQSM 238
Query: 1135 TAHSRFSILISINEV 1149
+F ++IS E+
Sbjct: 239 NGDKKFDVIISYLEI 253
>KF3A_HUMAN (Q9Y496) Kinesin-like protein KIF3A (Microtubule plus
end-directed kinesin motor 3A)
Length = 702
Score = 33.9 bits (76), Expect = 3.6
Identities = 15/42 (35%), Positives = 23/42 (54%)
Query: 1075 KKVLDVVMSNKGGFLFLYGFGGTGKTFVWNTLSAAVRSKGLI 1116
+ ++D V+ G +F YG GTGKTF + A +G+I
Sbjct: 82 RPIIDSVLEGYNGTIFAYGQTGTGKTFTMEGVRAIPELRGII 123
>RPS5_ARATH (O64973) Disease resistance protein RPS5 (Resistance to
Pseudomonas syringae protein 5) (pNd3/pNd10)
Length = 889
Score = 33.5 bits (75), Expect = 4.7
Identities = 32/114 (28%), Positives = 52/114 (45%), Gaps = 22/114 (19%)
Query: 996 LQIDDDELMNLCLIEI-EKELQCNAR-------SLKDFPSLSYPTFAEIMNFENKFV-AD 1046
L+ ++ EL LCL K+L+ + R LK+ SLS F ++++ F D
Sbjct: 90 LRSNEVELQRLCLCGFCSKDLKLSYRYGKRVIMMLKEVESLSSQGFFDVVSEATPFADVD 149
Query: 1047 ELNYNKEEMSKIHEELVRSLTGEQKAAYKKVLDVVMSNKGGFLFLYGFGGTGKT 1100
E+ + ++ G Q+ +K + +M + G L LYG GG GKT
Sbjct: 150 EIPFQP------------TIVG-QEIMLEKAWNRLMEDGSGILGLYGMGGVGKT 190
>DYHC_DROME (P37276) Dynein heavy chain, cytosolic (DYHC)
Length = 4639
Score = 33.5 bits (75), Expect = 4.7
Identities = 32/102 (31%), Positives = 53/102 (51%), Gaps = 12/102 (11%)
Query: 1047 ELNYNKEEMSKIHEE--LVRSLTGEQKAAY-KKVLDVV-MSNKGGFLFLYGFGGTGKTFV 1102
E+ KEE+ K+ +E LV EQ AA+ +KVL + +SN L + G G+GK+
Sbjct: 2160 EMKGLKEEIRKVCQEDYLVCGEGDEQGAAWMEKVLQLYQISNLNHGLMMVGPSGSGKSTA 2219
Query: 1103 WNTLSAAV-RSKGLIVLNVASSGIASLLLPGGRTAHSRFSIL 1143
W TL A+ R +G+ G+A ++ P + + + +L
Sbjct: 2220 WKTLLKALERFEGV-------EGVAHVIDPKAISKEALYGVL 2254
>KF3A_MOUSE (P28741) Kinesin-like protein KIF3A (Microtubule plus
end-directed kinesin motor 3A)
Length = 701
Score = 33.1 bits (74), Expect = 6.1
Identities = 15/42 (35%), Positives = 23/42 (54%)
Query: 1075 KKVLDVVMSNKGGFLFLYGFGGTGKTFVWNTLSAAVRSKGLI 1116
+ ++D V+ G +F YG GTGKTF + A +G+I
Sbjct: 82 RPIIDSVLEGYNGTIFAYGQTGTGKTFTMEGVRAVPGLRGVI 123
>LON_CAMJE (O69300) ATP-dependent protease La (EC 3.4.21.53)
Length = 791
Score = 32.7 bits (73), Expect = 8.0
Identities = 37/146 (25%), Positives = 67/146 (45%), Gaps = 32/146 (21%)
Query: 1042 KFVADELNYNKEEMSKIHEEL-----VRSLTGEQKAAYKKVLDVVMSNKGGFLFLYGFGG 1096
K V+ +LN++ ++K E + VR L ++K A K V+ L LYG G
Sbjct: 315 KEVSKQLNHDHYALNKPKERIEEYFAVRELLEKRKIAEKDGAKVI-------LCLYGPPG 367
Query: 1097 TGKTFVWNTLSAAVRSKGLIVLNVASSGIASL------------LLPG----GRTAHSRF 1140
GKT + N++S A++ + ++ +A G+ + +PG G +
Sbjct: 368 VGKTSLANSVSKALKRE---LIRIALGGLEDVNELRGHRRTYIGAMPGRITQGLIEAKQI 424
Query: 1141 SILISINEVSTCNLR-QGSPKAELLQ 1165
+ +I ++E+ N +G P A LL+
Sbjct: 425 NPVIVLDEIDKLNRSFRGDPSAVLLE 450
>EX5A_HAEIN (P45158) Exodeoxyribonuclease V alpha chain (EC 3.1.11.5)
Length = 640
Score = 32.7 bits (73), Expect = 8.0
Identities = 18/68 (26%), Positives = 34/68 (49%), Gaps = 7/68 (10%)
Query: 1478 FAMTINKSQGQSLSHVGLYLP---RPVFTHGQLYVALSRVKSRKGLKVLIVVEQGVVSTS 1534
F MTI+KSQG H + LP PV + ++ ++R K ++ + ++ + T+
Sbjct: 566 FMMTIHKSQGSEFKHTVMVLPTEVNPVLSRELVFTGVTRAKK----ELTVFADEKIWKTA 621
Query: 1535 TRNVVYKE 1542
R V ++
Sbjct: 622 IRQTVKRQ 629
>DR18_ARATH (O64789) Putative disease resistance protein At1g61310
Length = 925
Score = 32.7 bits (73), Expect = 8.0
Identities = 24/115 (20%), Positives = 55/115 (46%), Gaps = 5/115 (4%)
Query: 996 LQIDDDELMNLCLIEIEKELQCNARSLKDFPSLSYPTFAEIMNFENKFVADELNYNKEEM 1055
+ I+ +L+++ +E++K C + S Y ++ E K + E N+++
Sbjct: 81 IDIECKDLLSVSPVELQKLCLCGLCTKYVCSSYKYGKKVFLLLEEVKILKSEGNFDEVSQ 140
Query: 1056 ----SKIHEELVRSLTGEQKAAYKKVLDVVMSNKGGFLFLYGFGGTGKTFVWNTL 1106
S++ E + G+++ +K + +M + G + L+G GG GKT ++ +
Sbjct: 141 PPPRSEVEERPTQPTIGQEEML-EKAWNRLMEDGVGIMGLHGMGGVGKTTLFKKI 194
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.331 0.145 0.455
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 178,412,300
Number of Sequences: 164201
Number of extensions: 7638616
Number of successful extensions: 20048
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 20031
Number of HSP's gapped (non-prelim): 25
length of query: 1547
length of database: 59,974,054
effective HSP length: 123
effective length of query: 1424
effective length of database: 39,777,331
effective search space: 56642919344
effective search space used: 56642919344
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0209.3


