

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0207.3
(227 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
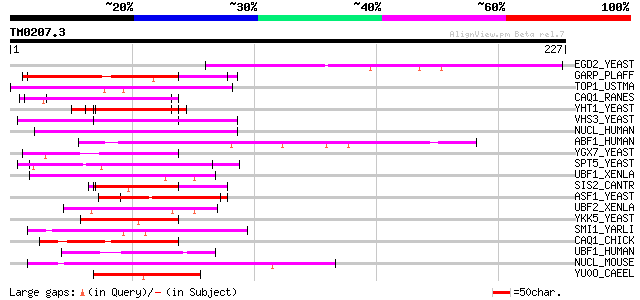
Score E
Sequences producing significant alignments: (bits) Value
EGD2_YEAST (P38879) EGD2 protein (GAL4 DNA-binding enhancer prot... 110 4e-24
GARP_PLAFF (P13816) Glutamic acid-rich protein precursor 58 2e-08
TOP1_USTMA (P41511) DNA topoisomerase I (EC 5.99.1.2) 54 2e-07
CAQ1_RANES (P31231) Calsequestrin, skeletal muscle isoform precu... 53 5e-07
YHT1_YEAST (P38835) Hypothetical 95.1 kDa protein in ACT5-YCK1 i... 52 9e-07
VHS3_YEAST (Q08438) Protein VHS3 (Viable in a HAL3 SIT4 backgrou... 52 2e-06
NUCL_HUMAN (P19338) Nucleolin (Protein C23) 51 2e-06
ABF1_HUMAN (Q15911) Alpha-fetoprotein enhancer binding protein (... 51 3e-06
YGX7_YEAST (P53076) Hypothetical 108.2 kDa protein in SAP4-OST5 ... 50 3e-06
SPT5_YEAST (P27692) Transcription initiation protein SPT5 50 4e-06
UBF1_XENLA (P25979) Nucleolar transcription factor 1 (Upstream b... 50 6e-06
SIS2_CANTR (Q12600) SIS2 protein (Halotolerance protein HAL3) 50 6e-06
ASF1_YEAST (P32447) Anti-silencing protein 1 50 6e-06
UBF2_XENLA (P25980) Nucleolar transcription factor 2 (Upstream b... 49 7e-06
YKK5_YEAST (P34250) Hypothetical 125.6 kDa protein in AAT1-GFA1 ... 49 1e-05
SMI1_YARLI (Q6CDX0) KNR4/SMI1 homolog 49 1e-05
CAQ1_CHICK (P19204) Calsequestrin, skeletal muscle isoform precu... 49 1e-05
UBF1_HUMAN (P17480) Nucleolar transcription factor 1 (Upstream b... 49 1e-05
NUCL_MOUSE (P09405) Nucleolin (Protein C23) 49 1e-05
YU0O_CAEEL (Q19753) Hypothetical protein F23B12.7 in chromosome V 48 2e-05
>EGD2_YEAST (P38879) EGD2 protein (GAL4 DNA-binding enhancer protein
2)
Length = 174
Score = 110 bits (274), Expect = 4e-24
Identities = 65/161 (40%), Positives = 96/161 (59%), Gaps = 16/161 (9%)
Query: 81 SRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISKPDVFKSPNSETYVIFGEAKIE 140
+++EKK+R+ + KLGLK + G+ RVT ++ N ++ I KP+VF+S YV+FGEAK++
Sbjct: 14 NKNEKKARELIGKLGLKQIPGIIRVTFRKKDNQIYAIEKPEVFRSAGGN-YVVFGEAKVD 72
Query: 141 DLSSQL---QTQAAQQFRMPDMGSVMAKQ------DQGTAADGG------QPEEEEEVDE 185
+ + +L Q QA MP V K D AA+G + +EE EVD
Sbjct: 73 NFTQKLAAAQQQAQASGIMPSNEDVATKSPEDIQADMQAAAEGSVNAAAEEDDEEGEVDA 132
Query: 186 TGVEPHDIDLVMTQAGVSRGKAVKALKTHNGDIVGAIMELT 226
+ DI+LV+ Q VS+ +A+KALK HNGD+V AIM L+
Sbjct: 133 GDLNKDDIELVVQQTNVSKNQAIKALKAHNGDLVNAIMSLS 173
>GARP_PLAFF (P13816) Glutamic acid-rich protein precursor
Length = 678
Score = 57.8 bits (138), Expect = 2e-08
Identities = 28/82 (34%), Positives = 47/82 (57%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDK 67
+ E E E+ EE + + + EDDA ED ++D+D+A++D+DE+DDDE+DD
Sbjct: 585 EEEEEEEEEEEEEEEEEEEEEDEDEEDEDDAEEDEDDAEEDEDDAEEDDDEEDDDEEDDD 644
Query: 68 EDGALGGSEGSKQSRSEKKSRK 89
ED + ++ E++S K
Sbjct: 645 EDEDEDEEDEEEEEEEEEESEK 666
Score = 57.4 bits (137), Expect = 3e-08
Identities = 31/86 (36%), Positives = 48/86 (55%), Gaps = 3/86 (3%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDK 67
+ E E E+ E+ + + EDDA ED ++D DE DDDE++DD+DED+D+
Sbjct: 592 EEEEEEEEEEEEEEDEDEEDEDDAEEDEDDAEEDEDDAEEDDDEEDDDEEDDDEDEDEDE 651
Query: 68 EDGALGGSEGSKQSRSEKKSRKAMLK 93
ED E ++ SEKK ++ + K
Sbjct: 652 EDEE---EEEEEEEESEKKIKRNLRK 674
Score = 51.6 bits (122), Expect = 2e-06
Identities = 27/85 (31%), Positives = 47/85 (54%), Gaps = 3/85 (3%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDED---EDDDDED 64
+ E E E+ EE +++ + +ED+ ED ++D+D+A++DED EDDD+ED
Sbjct: 578 EEEEEEEEEEEEEEEEEEEEEEEEEEEEDEDEEDEDDAEEDEDDAEEDEDDAEEDDDEED 637
Query: 65 DDKEDGALGGSEGSKQSRSEKKSRK 89
DD+ED E + E++ +
Sbjct: 638 DDEEDDDEDEDEDEEDEEEEEEEEE 662
Score = 47.4 bits (111), Expect = 3e-05
Identities = 23/62 (37%), Positives = 37/62 (59%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDK 67
+ E E E+ EE +++ + +E++ ED D ++DE D +EDEDD +EDDD+
Sbjct: 576 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDEDEEDEDDAEEDEDDAEEDEDDAEEDDDE 635
Query: 68 ED 69
ED
Sbjct: 636 ED 637
Score = 45.4 bits (106), Expect = 1e-04
Identities = 19/62 (30%), Positives = 39/62 (62%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDK 67
+ + E E+ EE +++ + +E++ ED ++D+D+A++DED+ ++DEDD +
Sbjct: 571 EVEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDEDEEDEDDAEEDEDDAEEDEDDAE 630
Query: 68 ED 69
ED
Sbjct: 631 ED 632
Score = 43.1 bits (100), Expect = 5e-04
Identities = 17/62 (27%), Positives = 40/62 (64%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDK 67
+ + E +E+ EE +++ + +E++ E+ +++++DE ++DED+ ++DEDD +
Sbjct: 564 EVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDEDEEDEDDAEEDEDDAE 623
Query: 68 ED 69
ED
Sbjct: 624 ED 625
Score = 42.7 bits (99), Expect = 7e-04
Identities = 18/64 (28%), Positives = 43/64 (67%), Gaps = 3/64 (4%)
Query: 6 VVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDD 65
V + S E + E+ + +E +++ + +E++ E+ ++++++E +++EDED++DEDD
Sbjct: 558 VQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEE---EEEEEEEEEEEEEEEDEDEEDEDD 614
Query: 66 DKED 69
+ED
Sbjct: 615 AEED 618
Score = 42.4 bits (98), Expect = 0.001
Identities = 18/64 (28%), Positives = 39/64 (60%)
Query: 6 VVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDD 65
V + E E E+ EE +++ + +E++ E+ ++D+ +E +DD +ED+DD ++
Sbjct: 565 VQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDEDEEDEDDAEEDEDDAEE 624
Query: 66 DKED 69
D++D
Sbjct: 625 DEDD 628
Score = 39.3 bits (90), Expect = 0.008
Identities = 18/75 (24%), Positives = 45/75 (60%), Gaps = 1/75 (1%)
Query: 12 EAEALEQV-LSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDG 70
EAE +Q + EE ++ + QE+ + ED ++ ++DE +++E+E++++E++++E+
Sbjct: 536 EAELQKQKHVDKEEDKKEESKEVQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEE 595
Query: 71 ALGGSEGSKQSRSEK 85
E ++ E+
Sbjct: 596 EEEEEEEEEEDEDEE 610
Score = 37.7 bits (86), Expect = 0.023
Identities = 13/62 (20%), Positives = 42/62 (66%), Gaps = 3/62 (4%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDK 67
+ S E + + + +E +++ + +E++ E+ ++++++E +++E+E++++ED+D+
Sbjct: 553 EESKEVQEESKEVQEDEEEVEEDEEEEEEEE---EEEEEEEEEEEEEEEEEEEEEEDEDE 609
Query: 68 ED 69
ED
Sbjct: 610 ED 611
Score = 35.8 bits (81), Expect = 0.086
Identities = 14/71 (19%), Positives = 41/71 (57%)
Query: 16 LEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGS 75
+E+ ++ + K+ +E+ + E+ K+ +DE + +EDE++++E++++E+
Sbjct: 534 IEEAELQKQKHVDKEEDKKEESKEVQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEE 593
Query: 76 EGSKQSRSEKK 86
E ++ E++
Sbjct: 594 EEEEEEEEEEE 604
Score = 34.3 bits (77), Expect = 0.25
Identities = 15/75 (20%), Positives = 43/75 (57%), Gaps = 2/75 (2%)
Query: 17 EQVLSAEETTLKKKPQ--PQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGG 74
+++ EE L+K+ +ED ++V+++ K+ +D+E+ ++D+E++++E+
Sbjct: 529 KKMAKIEEAELQKQKHVDKEEDKKEESKEVQEESKEVQEDEEEVEEDEEEEEEEEEEEEE 588
Query: 75 SEGSKQSRSEKKSRK 89
E ++ E++ +
Sbjct: 589 EEEEEEEEEEEEEEE 603
Score = 33.9 bits (76), Expect = 0.33
Identities = 10/63 (15%), Positives = 40/63 (62%)
Query: 7 VDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDD 66
VD + + + + E +++ + E+D E+ ++++++E +++E+E++++E+++
Sbjct: 545 VDKEEDKKEESKEVQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 604
Query: 67 KED 69
+++
Sbjct: 605 EDE 607
Score = 32.3 bits (72), Expect = 0.95
Identities = 17/51 (33%), Positives = 27/51 (52%), Gaps = 4/51 (7%)
Query: 43 DVKDDDKDEADDDEDEDDDDEDD----DKEDGALGGSEGSKQSRSEKKSRK 89
D KD+DK+ +D+ ED E+ DK++ E +Q + EKK +K
Sbjct: 234 DTKDNDKENISEDKKEDHQQEEMLKTLDKKERKQKEKEMKEQEKIEKKKKK 284
>TOP1_USTMA (P41511) DNA topoisomerase I (EC 5.99.1.2)
Length = 1019
Score = 54.3 bits (129), Expect = 2e-07
Identities = 35/106 (33%), Positives = 50/106 (47%), Gaps = 15/106 (14%)
Query: 1 MAPGPVVDASSEAEALEQVLSAEETTLKKKPQPQEDD----------APIVEDVK----- 45
+A P V+ S L +S++ K+ DD AP+ VK
Sbjct: 48 LAKRPKVEDSDSDAPLTSTVSSQNGVQKRSGSSNNDDNDDDSDSDSDAPLTALVKKSNGS 107
Query: 46 DDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAM 91
DDD+D+ DDD++ DDDDE+DD +D SK S+S +K K M
Sbjct: 108 DDDEDDDDDDDEGDDDDEEDDDDDDDDDDKPLSKSSKSNRKPPKTM 153
>CAQ1_RANES (P31231) Calsequestrin, skeletal muscle isoform
precursor
Length = 420
Score = 53.1 bits (126), Expect = 5e-07
Identities = 24/54 (44%), Positives = 32/54 (58%)
Query: 16 LEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+E VL E T +DD +D DDD D+ DDD+D+DDDD+DDD +D
Sbjct: 365 IEDVLEGEVNTEDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDD 418
Score = 50.8 bits (120), Expect = 3e-06
Identities = 22/63 (34%), Positives = 38/63 (59%)
Query: 7 VDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDD 66
+D + ++++ E L+ + ++DD +D DDD D+ DDD+D+DDDD+DDD
Sbjct: 349 MDDEEDLPTVDELEDWIEDVLEGEVNTEDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDD 408
Query: 67 KED 69
+D
Sbjct: 409 DDD 411
Score = 47.4 bits (111), Expect = 3e-05
Identities = 24/64 (37%), Positives = 35/64 (54%), Gaps = 2/64 (3%)
Query: 5 PVVDASSE--AEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDD 62
P VD + + LE ++ E+ +DD +D DDD D+ DDD+D+DDDD
Sbjct: 356 PTVDELEDWIEDVLEGEVNTEDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDD 415
Query: 63 EDDD 66
+DDD
Sbjct: 416 DDDD 419
>YHT1_YEAST (P38835) Hypothetical 95.1 kDa protein in ACT5-YCK1
intergenic region
Length = 840
Score = 52.4 bits (124), Expect = 9e-07
Identities = 22/38 (57%), Positives = 28/38 (72%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGAL 72
EDD +D DDD DE DDD+D+DDDD+DDD +DG +
Sbjct: 801 EDDDDDDDDDDDDDDDEDDDDDDDDDDDDDDDDDDGQI 838
Score = 45.8 bits (107), Expect = 8e-05
Identities = 18/34 (52%), Positives = 25/34 (72%)
Query: 36 DDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
DD +D DDD D+ DD++D+DDDD+DDD +D
Sbjct: 798 DDGEDDDDDDDDDDDDDDDEDDDDDDDDDDDDDD 831
Score = 45.8 bits (107), Expect = 8e-05
Identities = 20/44 (45%), Positives = 27/44 (60%)
Query: 26 TLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
T K EDD + DDD D+ DDD+D+D+DD+DDD +D
Sbjct: 783 TTKDAEDHGEDDGDGDDGEDDDDDDDDDDDDDDDEDDDDDDDDD 826
Score = 43.5 bits (101), Expect = 4e-04
Identities = 17/35 (48%), Positives = 25/35 (70%)
Query: 32 QPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDD 66
+ +DD +D DD+ D+ DDD+D+DDDD+DDD
Sbjct: 801 EDDDDDDDDDDDDDDDEDDDDDDDDDDDDDDDDDD 835
Score = 32.0 bits (71), Expect = 1.2
Identities = 18/68 (26%), Positives = 29/68 (42%)
Query: 21 SAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQ 80
S+ L + DD E +D++ E D+ +D +D+ DD G GSE
Sbjct: 603 SSRSLCLTNRDAEINDDESATETDEDENDGETDEYAGDDTNDDTDDSNIGYAYGSESDYS 662
Query: 81 SRSEKKSR 88
E++ R
Sbjct: 663 CLIEERIR 670
Score = 29.6 bits (65), Expect = 6.1
Identities = 12/30 (40%), Positives = 18/30 (60%)
Query: 40 IVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
IV KD + DD + +D +D+DDD +D
Sbjct: 780 IVGTTKDAEDHGEDDGDGDDGEDDDDDDDD 809
>VHS3_YEAST (Q08438) Protein VHS3 (Viable in a HAL3 SIT4 background
protein 3)
Length = 674
Score = 51.6 bits (122), Expect = 2e-06
Identities = 27/66 (40%), Positives = 37/66 (55%)
Query: 4 GPVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDE 63
G D S+ E E + EE KK +D+ +D DDD D+ DDD+D+DDDD+
Sbjct: 570 GGYPDVSAGKEEEEDEDNDEEDDNKKNDTGGKDEDNDDDDDDDDDDDDDDDDDDDDDDDD 629
Query: 64 DDDKED 69
DDD +D
Sbjct: 630 DDDDDD 635
Score = 50.8 bits (120), Expect = 3e-06
Identities = 23/59 (38%), Positives = 34/59 (56%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLK 93
+DD +D DDD D+ DDD+D+DDDD+DDD +D + + +KK K L+
Sbjct: 614 DDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDEDDEDEDEDDEGKKKEDKGGLQ 672
Score = 42.0 bits (97), Expect = 0.001
Identities = 18/44 (40%), Positives = 29/44 (65%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGS 78
+DD +D DDD D+ DD++DED+D++D+ K+ GG + S
Sbjct: 631 DDDDDDDDDDDDDDDDDDDDEDDEDEDEDDEGKKKEDKGGLQRS 674
>NUCL_HUMAN (P19338) Nucleolin (Protein C23)
Length = 706
Score = 51.2 bits (121), Expect = 2e-06
Identities = 26/83 (31%), Positives = 47/83 (56%)
Query: 11 SEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDG 70
SE EA+E + + K P ++ A ++ +DD+ ++ DDDED++DDD++DD+E+
Sbjct: 205 SEEEAMETTPAKGKKAAKVVPVKAKNVAEDEDEEEDDEDEDDDDDEDDEDDDDEDDEEEE 264
Query: 71 ALGGSEGSKQSRSEKKSRKAMLK 93
E K++ ++K A K
Sbjct: 265 EEEEEEPVKEAPGKRKKEMAKQK 287
Score = 42.4 bits (98), Expect = 0.001
Identities = 24/73 (32%), Positives = 37/73 (49%), Gaps = 1/73 (1%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKD-DDKDEADDDEDEDDDDEDDD 66
D S E E ++ +E ++ AP ED D DD+D+ DDD+DE+DD E++
Sbjct: 150 DDSEEDEEDDEDEDEDEDEIEPAAMKAAAAAPASEDEDDEDDEDDEDDDDDEEDDSEEEA 209
Query: 67 KEDGALGGSEGSK 79
E G + +K
Sbjct: 210 METTPAKGKKAAK 222
Score = 41.6 bits (96), Expect = 0.002
Identities = 26/85 (30%), Positives = 39/85 (45%), Gaps = 26/85 (30%)
Query: 28 KKKPQPQEDDAPIVEDVKDDDKDEADDDE----------------------DEDDDDEDD 65
KK+ +E+D ED +DD+ ++ D+DE DEDD+D+DD
Sbjct: 140 KKEDSDEEEDDDSEEDEEDDEDEDEDEDEIEPAAMKAAAAAPASEDEDDEDDEDDEDDDD 199
Query: 66 DKEDGALGGSEGSKQSRSEKKSRKA 90
D+ED SE + K +KA
Sbjct: 200 DEED----DSEEEAMETTPAKGKKA 220
Score = 37.4 bits (85), Expect = 0.029
Identities = 17/79 (21%), Positives = 41/79 (51%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDK 67
D E + ++ +EE ++ P + A +V + ++ D++ED++D+D+DDD+
Sbjct: 191 DEDDEDDDDDEEDDSEEEAMETTPAKGKKAAKVVPVKAKNVAEDEDEEEDDEDEDDDDDE 250
Query: 68 EDGALGGSEGSKQSRSEKK 86
+D + ++ E++
Sbjct: 251 DDEDDDDEDDEEEEEEEEE 269
Score = 34.7 bits (78), Expect = 0.19
Identities = 10/28 (35%), Positives = 23/28 (81%)
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKED 69
++ K +D DE +DD+ E+D+++D+D+++
Sbjct: 137 KNAKKEDSDEEEDDDSEEDEEDDEDEDE 164
Score = 32.7 bits (73), Expect = 0.72
Identities = 22/102 (21%), Positives = 47/102 (45%), Gaps = 20/102 (19%)
Query: 28 KKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDD----------------DEDDDKEDGA 71
K +++D+ ED ++ +E D+DEDED+D +++DD++D
Sbjct: 134 KNGKNAKKEDSDEEEDDDSEEDEEDDEDEDEDEDEIEPAAMKAAAAAPASEDEDDEDDED 193
Query: 72 LGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNI 113
+ ++ SE+++ + G K ++V + KN+
Sbjct: 194 DEDDDDDEEDDSEEEAMETTPAKGKK----AAKVVPVKAKNV 231
>ABF1_HUMAN (Q15911) Alpha-fetoprotein enhancer binding protein (AT
motif-binding factor) (AT-binding transcription factor
1)
Length = 3703
Score = 50.8 bits (120), Expect = 3e-06
Identities = 41/174 (23%), Positives = 84/174 (47%), Gaps = 19/174 (10%)
Query: 29 KKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSR 88
+K +P E++A ++++++EA+++E+E++++E++++++G G E +
Sbjct: 454 EKVEPAEEEAE-----EEEEEEEAEEEEEEEEEEEEEEEDEGCKGLFPSELDEELEDRPH 508
Query: 89 K---AMLKLGLKPVTGVSRVTIKRT---KNILFFISKPDVFKSPNS-ETYVIFGEA---- 137
+ A K +S +I + N+L +S+ S NS ++V+F A
Sbjct: 509 EEPGAAAGSSSKKDLALSNQSISNSPLMPNVLQTLSRGTASTSSNSASSFVVFDGANRRN 568
Query: 138 KIEDLSSQLQTQAAQQFRMPDMGSVMAKQDQGTAADGGQPEEEEEVDETGVEPH 191
++ S ++T A+ R D A +D TA +P E E D+ G PH
Sbjct: 569 RLSFNSEGVRTNVAEGGRRLDFADESANKDNATAP---EPNESTEGDDGGFVPH 619
>YGX7_YEAST (P53076) Hypothetical 108.2 kDa protein in SAP4-OST5
intergenic region
Length = 958
Score = 50.4 bits (119), Expect = 3e-06
Identities = 29/69 (42%), Positives = 38/69 (55%), Gaps = 12/69 (17%)
Query: 6 VVDASSEA-----EALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDD 60
V+ ASS+ E LE VLS E L + + D DDD D+ DDD+D+DD
Sbjct: 127 VMKASSDMRKNAQELLEPVLSEREMALNS-------NTSLENDRNDDDDDDDDDDDDDDD 179
Query: 61 DDEDDDKED 69
DD+DDD+ D
Sbjct: 180 DDDDDDESD 188
Score = 31.6 bits (70), Expect = 1.6
Identities = 17/60 (28%), Positives = 31/60 (51%), Gaps = 11/60 (18%)
Query: 44 VKDDDKDEADDDEDEDDDDE-----------DDDKEDGALGGSEGSKQSRSEKKSRKAML 92
+++ D DE+++DE E+ +E D D+EDGA G + K + + + K +L
Sbjct: 622 IEEFDSDESEEDETENGPEENKSTNVNEDLMDIDQEDGAAGNKDTKKLNDEKDNNLKFLL 681
>SPT5_YEAST (P27692) Transcription initiation protein SPT5
Length = 1063
Score = 50.1 bits (118), Expect = 4e-06
Identities = 25/85 (29%), Positives = 49/85 (57%), Gaps = 6/85 (7%)
Query: 4 GPVVD-----ASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDE 58
GP D ++++ +++ EE + ++K +P+E+D + D+ D D+D+DE
Sbjct: 95 GPATDDAQATLNTDSSEANEIVKKEEGSDERK-RPREEDTKNSDGDTKDEGDNKDEDDDE 153
Query: 59 DDDDEDDDKEDGALGGSEGSKQSRS 83
DDDD+DDD++D ++ +Q R+
Sbjct: 154 DDDDDDDDEDDDDEAPTKRRRQERN 178
Score = 43.1 bits (100), Expect = 5e-04
Identities = 22/87 (25%), Positives = 45/87 (51%), Gaps = 1/87 (1%)
Query: 9 ASSEAEALEQVLSAEETTLKKKPQPQED-DAPIVEDVKDDDKDEADDDEDEDDDDEDDDK 67
A+ +A+A S+E + KK + ++ P ED K+ D D D+ +++D+DD++DD
Sbjct: 97 ATDDAQATLNTDSSEANEIVKKEEGSDERKRPREEDTKNSDGDTKDEGDNKDEDDDEDDD 156
Query: 68 EDGALGGSEGSKQSRSEKKSRKAMLKL 94
+D + ++ ++ R L +
Sbjct: 157 DDDDDEDDDDEAPTKRRRQERNRFLDI 183
Score = 30.0 bits (66), Expect = 4.7
Identities = 18/55 (32%), Positives = 23/55 (41%), Gaps = 17/55 (30%)
Query: 29 KKPQPQEDDAPIVEDVKDDDKD-----------------EADDDEDEDDDDEDDD 66
K EDD +D DDD+ E DDEDED+D+ED +
Sbjct: 147 KDEDDDEDDDDDDDDEDDDDEAPTKRRRQERNRFLDIEAEVSDDEDEDEDEEDSE 201
>UBF1_XENLA (P25979) Nucleolar transcription factor 1 (Upstream
binding factor-1) (UBF-1) (xUBF-1)
Length = 677
Score = 49.7 bits (117), Expect = 6e-06
Identities = 30/79 (37%), Positives = 42/79 (52%), Gaps = 3/79 (3%)
Query: 9 ASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDD-EDDDK 67
+S + A ++ S + + K P + + DDD ++ DDDEDEDDDD ED+DK
Sbjct: 556 SSQDRAAYKEQNSNKRKSTAKIQVPVAKPKLVAQSKSDDDDEDDDDDEDEDDDDDEDEDK 615
Query: 68 EDGALGG--SEGSKQSRSE 84
ED + G SE S SE
Sbjct: 616 EDSSEDGDSSESSSDEDSE 634
Score = 40.0 bits (92), Expect = 0.005
Identities = 19/65 (29%), Positives = 31/65 (47%)
Query: 23 EETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSR 82
+E K+ D + D +D +E +D+E E+D+DE++D +D G S S S
Sbjct: 610 DEDEDKEDSSEDGDSSESSSDEDSEDGEENEDEEGEEDEDEEEDDDDNESGSSSSSSSSA 669
Query: 83 SEKKS 87
S
Sbjct: 670 DSSDS 674
Score = 37.7 bits (86), Expect = 0.023
Identities = 22/66 (33%), Positives = 32/66 (48%), Gaps = 13/66 (19%)
Query: 32 QPQEDDAPIVEDVKDDDKDEADDDEDEDDDD-----------EDDDKEDGALGGSEGSKQ 80
Q + DD ED DD+ ++ DDDEDED +D D+D EDG E ++
Sbjct: 589 QSKSDDDD--EDDDDDEDEDDDDDEDEDKEDSSEDGDSSESSSDEDSEDGEENEDEEGEE 646
Query: 81 SRSEKK 86
E++
Sbjct: 647 DEDEEE 652
Score = 36.6 bits (83), Expect = 0.050
Identities = 14/46 (30%), Positives = 27/46 (58%)
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKS 87
ED +D +++E ++ E+++D++EDDD + S S S+ S
Sbjct: 631 EDSEDGEENEDEEGEEDEDEEEDDDDNESGSSSSSSSSADSSDSDS 676
Score = 35.8 bits (81), Expect = 0.086
Identities = 14/39 (35%), Positives = 24/39 (60%)
Query: 49 KDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKS 87
+ ++DDD+++DDDDED+D +D E S + +S
Sbjct: 589 QSKSDDDDEDDDDDEDEDDDDDEDEDKEDSSEDGDSSES 627
Score = 32.0 bits (71), Expect = 1.2
Identities = 18/64 (28%), Positives = 28/64 (43%), Gaps = 8/64 (12%)
Query: 29 KKPQPQEDDAPIVEDVKDDDKDEADDDEDED--------DDDEDDDKEDGALGGSEGSKQ 80
K EDD ++ DDD+DE +D ED D+D +D +E+ G E +
Sbjct: 591 KSDDDDEDDDDDEDEDDDDDEDEDKEDSSEDGDSSESSSDEDSEDGEENEDEEGEEDEDE 650
Query: 81 SRSE 84
+
Sbjct: 651 EEDD 654
Score = 30.4 bits (67), Expect = 3.6
Identities = 11/47 (23%), Positives = 25/47 (52%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQS 81
++D+ E+ +D++ +E +D+E++DDD+E + S
Sbjct: 630 DEDSEDGEENEDEEGEEDEDEEEDDDDNESGSSSSSSSSADSSDSDS 676
>SIS2_CANTR (Q12600) SIS2 protein (Halotolerance protein HAL3)
Length = 531
Score = 49.7 bits (117), Expect = 6e-06
Identities = 18/34 (52%), Positives = 28/34 (81%)
Query: 36 DDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
D++ I++D DDD D+ DDD+D+DDDD+DDD ++
Sbjct: 478 DESAIIDDDDDDDDDDDDDDDDDDDDDDDDDDDE 511
Score = 47.4 bits (111), Expect = 3e-05
Identities = 19/35 (54%), Positives = 27/35 (76%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
++ A I +D DDD D+ DDD+D+DDDD+DDD E+
Sbjct: 478 DESAIIDDDDDDDDDDDDDDDDDDDDDDDDDDDEE 512
Score = 46.6 bits (109), Expect = 5e-05
Identities = 23/64 (35%), Positives = 35/64 (53%), Gaps = 7/64 (10%)
Query: 33 PQEDDAPIVEDVKDD-------DKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEK 85
P+++D +D KD+ D D+ DDD+D+DDDD+DDD +D E Q +S
Sbjct: 462 PEDEDEDEADDSKDNIDESAIIDDDDDDDDDDDDDDDDDDDDDDDDDDDEEDPPQQQSTT 521
Query: 86 KSRK 89
+ K
Sbjct: 522 DNSK 525
Score = 31.6 bits (70), Expect = 1.6
Identities = 19/72 (26%), Positives = 34/72 (46%), Gaps = 3/72 (4%)
Query: 1 MAPGPVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDD 60
M G + E EA + + +E+ + +DD +D DDD D+ DDDE++
Sbjct: 456 MKLGGYPEDEDEDEADDSKDNIDESAIIDDDDDDDDDDDDDDDDDDDDDDDDDDDEEDPP 515
Query: 61 DDE---DDDKED 69
+ D+ K++
Sbjct: 516 QQQSTTDNSKDE 527
>ASF1_YEAST (P32447) Anti-silencing protein 1
Length = 279
Score = 49.7 bits (117), Expect = 6e-06
Identities = 22/50 (44%), Positives = 35/50 (70%), Gaps = 1/50 (2%)
Query: 37 DAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKK 86
+ P V+D +++D +E DDDED D+DDEDDD+EDG E +++ E++
Sbjct: 165 EQPGVDDEEEEDDEEEDDDED-DEDDEDDDQEDGEGEAEEAAEEEEEEEE 213
Score = 43.1 bits (100), Expect = 5e-04
Identities = 16/44 (36%), Positives = 30/44 (67%)
Query: 46 DDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRK 89
DD+++E D++ED+D+DDEDD+ +D G E + + E++ +
Sbjct: 170 DDEEEEDDEEEDDDEDDEDDEDDDQEDGEGEAEEAAEEEEEEEE 213
Score = 32.3 bits (72), Expect = 0.95
Identities = 17/66 (25%), Positives = 38/66 (56%), Gaps = 4/66 (6%)
Query: 24 ETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRS 83
E ++ + +E++ ED + + ++E +D E+ D D+E+ ++E G++ +E +
Sbjct: 198 EGEAEEAAEEEEEEEEKTEDNETNLEEEEEDIENSDGDEEEGEEEVGSVDKNEDG----N 253
Query: 84 EKKSRK 89
+KK RK
Sbjct: 254 DKKRRK 259
Score = 31.2 bits (69), Expect = 2.1
Identities = 19/63 (30%), Positives = 33/63 (52%), Gaps = 5/63 (7%)
Query: 8 DASSEAE-ALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDD 66
D EAE A E+ EE T + +E++ ED+++ D DE + +E+ D+++D
Sbjct: 196 DGEGEAEEAAEEEEEEEEKTEDNETNLEEEE----EDIENSDGDEEEGEEEVGSVDKNED 251
Query: 67 KED 69
D
Sbjct: 252 GND 254
>UBF2_XENLA (P25980) Nucleolar transcription factor 2 (Upstream
binding factor-2) (UBF-2) (xUBF-2)
Length = 701
Score = 49.3 bits (116), Expect = 7e-06
Identities = 27/71 (38%), Positives = 41/71 (57%), Gaps = 8/71 (11%)
Query: 23 EETTLKKKPQ-----PQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDD-DKEDGALGG-- 74
E+ T K+K P +++ DDD+D+ DD+++EDDDD+DD DKED + G
Sbjct: 587 EQNTNKRKSTTKIQAPSSKSKLVIQSKSDDDEDDEDDEDEEDDDDDDDEDKEDSSEDGDS 646
Query: 75 SEGSKQSRSEK 85
S+ S SE+
Sbjct: 647 SDSSSDEDSEE 657
Score = 42.0 bits (97), Expect = 0.001
Identities = 22/66 (33%), Positives = 36/66 (54%), Gaps = 5/66 (7%)
Query: 21 SAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALG-GSEGSK 79
S + ++ K EDD ED +D++ D+ DDDED++D ED D D + SE +
Sbjct: 604 SKSKLVIQSKSDDDEDD----EDDEDEEDDDDDDDEDKEDSSEDGDSSDSSSDEDSEEGE 659
Query: 80 QSRSEK 85
++ E+
Sbjct: 660 ENEDEE 665
Score = 40.0 bits (92), Expect = 0.005
Identities = 16/51 (31%), Positives = 30/51 (58%)
Query: 37 DAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKS 87
D+ ED ++ +++E ++DE++DD+D ++D +D G S S S S
Sbjct: 648 DSSSDEDSEEGEENEDEEDEEDDDEDNEEDDDDNESGSSSSSSSSADSSDS 698
Score = 38.1 bits (87), Expect = 0.017
Identities = 22/68 (32%), Positives = 36/68 (52%), Gaps = 5/68 (7%)
Query: 20 LSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSK 79
+ A + K Q + DD ED +DD+ +E DDD +DD+D++D EDG S +
Sbjct: 599 IQAPSSKSKLVIQSKSDDD---EDDEDDEDEEDDDD--DDDEDKEDSSEDGDSSDSSSDE 653
Query: 80 QSRSEKKS 87
S +++
Sbjct: 654 DSEEGEEN 661
Score = 37.0 bits (84), Expect = 0.038
Identities = 13/42 (30%), Positives = 26/42 (60%)
Query: 46 DDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKS 87
++++DE D+++D++D++EDDD + S S S+ S
Sbjct: 659 EENEDEEDEEDDDEDNEEDDDDNESGSSSSSSSSADSSDSDS 700
Score = 35.0 bits (79), Expect = 0.15
Identities = 14/49 (28%), Positives = 30/49 (60%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRS 83
++D+ E+ +D++ +E DD+++E+DDD+++ + S S S S
Sbjct: 652 DEDSEEGEENEDEEDEEDDDEDNEEDDDDNESGSSSSSSSSADSSDSDS 700
>YKK5_YEAST (P34250) Hypothetical 125.6 kDa protein in AAT1-GFA1
intergenic region
Length = 1132
Score = 48.9 bits (115), Expect = 1e-05
Identities = 23/41 (56%), Positives = 28/41 (68%), Gaps = 1/41 (2%)
Query: 30 KPQPQEDDAPIVEDVKDDDKDE-ADDDEDEDDDDEDDDKED 69
K +DD +DV DDDKDE DDDE+ DDDD+DDD +D
Sbjct: 601 KDNDNDDDDDKDDDVNDDDKDENVDDDENVDDDDDDDDDDD 641
Score = 37.4 bits (85), Expect = 0.029
Identities = 15/35 (42%), Positives = 24/35 (67%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+++ P + DDDKD +DD+D+ DDD +DD +D
Sbjct: 587 KNNGPNNDKDDDDDKDNDNDDDDDKDDDVNDDDKD 621
Score = 36.6 bits (83), Expect = 0.050
Identities = 16/36 (44%), Positives = 25/36 (69%), Gaps = 1/36 (2%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDD-DEDDDKED 69
ED+ V D + ++ DDD+D+D+D D+DDDK+D
Sbjct: 578 EDETGSVMDKNNGPNNDKDDDDDKDNDNDDDDDKDD 613
Score = 35.0 bits (79), Expect = 0.15
Identities = 17/40 (42%), Positives = 27/40 (67%), Gaps = 5/40 (12%)
Query: 31 PQPQEDDAPIVEDVKDDDKDEADD-DEDEDDDDEDDDKED 69
P +DD +D KD+D D+ DD D+D +DDD+D++ +D
Sbjct: 591 PNNDKDD----DDDKDNDNDDDDDKDDDVNDDDKDENVDD 626
>SMI1_YARLI (Q6CDX0) KNR4/SMI1 homolog
Length = 713
Score = 48.9 bits (115), Expect = 1e-05
Identities = 32/92 (34%), Positives = 51/92 (54%), Gaps = 4/92 (4%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVK-DDDKDEADD-DEDEDDDDEDD 65
DAS + EA + +AEE+ K + +E++K D++ A++ DE+ DDDDEDD
Sbjct: 624 DASKKVEA--EKAAAEESKESKAESEESKVERDLEELKIDEENGNAEEADEEADDDDEDD 681
Query: 66 DKEDGALGGSEGSKQSRSEKKSRKAMLKLGLK 97
++E + G E + S+ KS+K K G K
Sbjct: 682 EEEGDSKEGEETKSTTASKSKSKKKNKKKGKK 713
>CAQ1_CHICK (P19204) Calsequestrin, skeletal muscle isoform
precursor (Calsequestrin 1) (Aspartactin)
(Laminin-binding protein)
Length = 406
Score = 48.9 bits (115), Expect = 1e-05
Identities = 26/57 (45%), Positives = 35/57 (60%), Gaps = 5/57 (8%)
Query: 13 AEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
AE LE + E L K ++DD +D DDD D+ DDD+D+DDDD+DDD +D
Sbjct: 355 AEELEDWI---EDVLSGKINTEDDDDD--DDDDDDDDDDDDDDDDDDDDDDDDDDDD 406
Score = 35.8 bits (81), Expect = 0.086
Identities = 16/38 (42%), Positives = 25/38 (65%), Gaps = 3/38 (7%)
Query: 35 EDDAPIVEDVKDDDKDEAD---DDEDEDDDDEDDDKED 69
+DD P E+++D +D + ED+DDDD+DDD +D
Sbjct: 349 DDDLPTAEELEDWIEDVLSGKINTEDDDDDDDDDDDDD 386
>UBF1_HUMAN (P17480) Nucleolar transcription factor 1 (Upstream
binding factor 1) (UBF-1) (Autoantigen NOR-90)
Length = 764
Score = 48.5 bits (114), Expect = 1e-05
Identities = 24/63 (38%), Positives = 37/63 (58%), Gaps = 9/63 (14%)
Query: 22 AEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQS 81
+ TTL+ K + +EDD ++D+D+ D+DE+E+DD+ D EDG SE S +
Sbjct: 666 SSRTTLQSKSESEEDD--------EEDEDDEDEDEEEEDDENGDSSEDGG-DSSESSSED 716
Query: 82 RSE 84
SE
Sbjct: 717 ESE 719
Score = 41.6 bits (96), Expect = 0.002
Identities = 15/40 (37%), Positives = 29/40 (72%)
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQS 81
ED ++++D+ D+D+DEDDD+++D++ +G+ S S S
Sbjct: 719 EDGDENEEDDEDEDDDEDDDEDEDNESEGSSSSSSSSGDS 758
Score = 41.2 bits (95), Expect = 0.002
Identities = 15/46 (32%), Positives = 31/46 (66%)
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKS 87
++ +D D++E DD++++DD+D+D+D+++ + G S S S S
Sbjct: 716 DESEDGDENEEDDEDEDDDEDDDEDEDNESEGSSSSSSSSGDSSDS 761
Score = 40.4 bits (93), Expect = 0.003
Identities = 18/61 (29%), Positives = 34/61 (55%), Gaps = 1/61 (1%)
Query: 26 TLKKKPQPQEDDAPIVEDVKDDDKDEADDD-EDEDDDDEDDDKEDGALGGSEGSKQSRSE 84
T + P P+ + + ++ DE D+D EDED+++EDD+ D + G + S+ S +
Sbjct: 657 TKLRGPNPKSSRTTLQSKSESEEDDEEDEDDEDEDEEEEDDENGDSSEDGGDSSESSSED 716
Query: 85 K 85
+
Sbjct: 717 E 717
Score = 40.4 bits (93), Expect = 0.003
Identities = 18/44 (40%), Positives = 26/44 (58%)
Query: 44 VKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKS 87
++ + E DD+EDEDD+DED+++ED G S SE S
Sbjct: 671 LQSKSESEEDDEEDEDDEDEDEEEEDDENGDSSEDGGDSSESSS 714
Score = 37.0 bits (84), Expect = 0.038
Identities = 21/73 (28%), Positives = 35/73 (47%), Gaps = 2/73 (2%)
Query: 8 DASSEAEALEQVLSAEE--TTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDD 65
D E E E S+E+ + + + + +D E+ +D+ D+ DDDEDED++ E
Sbjct: 689 DEDEEEEDDENGDSSEDGGDSSESSSEDESEDGDENEEDDEDEDDDEDDDEDEDNESEGS 748
Query: 66 DKEDGALGGSEGS 78
+ G S S
Sbjct: 749 SSSSSSSGDSSDS 761
Score = 37.0 bits (84), Expect = 0.038
Identities = 17/75 (22%), Positives = 41/75 (54%), Gaps = 2/75 (2%)
Query: 15 ALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGG 74
A ++ +S + ++ K P + K + +++ ++DED++D+DE+++ ++
Sbjct: 644 AYKEYISNKRKSMTKLRGPNPKSSRTTLQSKSESEEDDEEDEDDEDEDEEEEDDENGDSS 703
Query: 75 SEG--SKQSRSEKKS 87
+G S +S SE +S
Sbjct: 704 EDGGDSSESSSEDES 718
>NUCL_MOUSE (P09405) Nucleolin (Protein C23)
Length = 706
Score = 48.5 bits (114), Expect = 1e-05
Identities = 37/127 (29%), Positives = 58/127 (45%), Gaps = 3/127 (2%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDK 67
D E ++ E+V+ E TT K K P + + V +++ DE +D++DED+DDE++D
Sbjct: 204 DEEEEDDSEEEVM--EITTAKGKKTPAKVVPMKAKSVAEEEDDEEEDEDDEDEDDEEEDD 261
Query: 68 EDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVT-IKRTKNILFFISKPDVFKSP 126
ED E + K +K M K P +V + T FI + KS
Sbjct: 262 EDDDEEEEEEEPVKAAPGKRKKEMTKQKEAPEAKKQKVEGSEPTTPFNLFIGNLNPNKSV 321
Query: 127 NSETYVI 133
N + I
Sbjct: 322 NELKFAI 328
Score = 40.4 bits (93), Expect = 0.003
Identities = 24/67 (35%), Positives = 35/67 (51%), Gaps = 6/67 (8%)
Query: 3 PGPVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDD 62
P P +++A+AL T KK A ++ K +D DE +D+EDEDD D
Sbjct: 104 PTPGKKGAAQAKALVP------TPGKKGAATPAKGAKNGKNAKKEDSDEDEDEEDEDDSD 157
Query: 63 EDDDKED 69
ED+D E+
Sbjct: 158 EDEDDEE 164
Score = 40.0 bits (92), Expect = 0.005
Identities = 25/63 (39%), Positives = 35/63 (54%), Gaps = 4/63 (6%)
Query: 8 DASSEAEALEQVLSAEETTLKK-KPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDD 66
D S E E E+ E +K KP AP ED +DD E +DDE++DD++E+DD
Sbjct: 154 DDSDEDEDDEEEDEFEPPIVKGVKPAKAAPAAPASEDEEDD---EDEDDEEDDDEEEEDD 210
Query: 67 KED 69
E+
Sbjct: 211 SEE 213
Score = 39.7 bits (91), Expect = 0.006
Identities = 14/25 (56%), Positives = 22/25 (88%)
Query: 45 KDDDKDEADDDEDEDDDDEDDDKED 69
+D D+DE ++DED+ D+DEDD++ED
Sbjct: 142 EDSDEDEDEEDEDDSDEDEDDEEED 166
Score = 36.6 bits (83), Expect = 0.050
Identities = 13/26 (50%), Positives = 21/26 (80%)
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDK 67
ED +D+ +E +DD DED+DDE++D+
Sbjct: 142 EDSDEDEDEEDEDDSDEDEDDEEEDE 167
Score = 32.0 bits (71), Expect = 1.2
Identities = 18/67 (26%), Positives = 34/67 (49%), Gaps = 4/67 (5%)
Query: 32 QPQEDDAPIVEDVKDDDKDEA----DDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKS 87
+ E + PIV+ VK A +D+ED++D+D+++D ++ SE + K
Sbjct: 164 EEDEFEPPIVKGVKPAKAAPAAPASEDEEDDEDEDDEEDDDEEEEDDSEEEVMEITTAKG 223
Query: 88 RKAMLKL 94
+K K+
Sbjct: 224 KKTPAKV 230
>YU0O_CAEEL (Q19753) Hypothetical protein F23B12.7 in chromosome V
Length = 953
Score = 48.1 bits (113), Expect = 2e-05
Identities = 22/45 (48%), Positives = 30/45 (65%), Gaps = 1/45 (2%)
Query: 35 EDDAPIVEDVKDDDKDEAD-DDEDEDDDDEDDDKEDGALGGSEGS 78
EDD D+ DDD+D+ D +D+DE DDD++DD EDG GG +
Sbjct: 843 EDDDEDDGDMDDDDEDDEDAEDDDEVDDDDEDDDEDGGFGGKSSA 887
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.305 0.127 0.336
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,397,300
Number of Sequences: 164201
Number of extensions: 1261695
Number of successful extensions: 28224
Number of sequences better than 10.0: 1196
Number of HSP's better than 10.0 without gapping: 820
Number of HSP's successfully gapped in prelim test: 410
Number of HSP's that attempted gapping in prelim test: 10865
Number of HSP's gapped (non-prelim): 6542
length of query: 227
length of database: 59,974,054
effective HSP length: 106
effective length of query: 121
effective length of database: 42,568,748
effective search space: 5150818508
effective search space used: 5150818508
T: 11
A: 40
X1: 16 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0207.3


