

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.8
(124 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
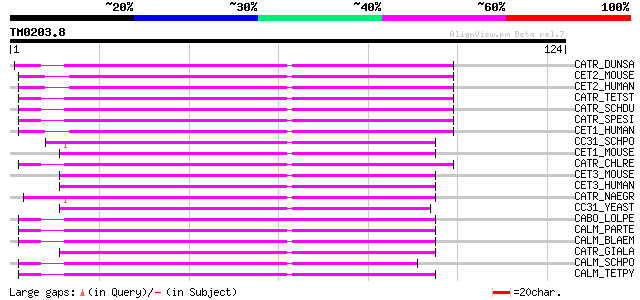
Score E
Sequences producing significant alignments: (bits) Value
CATR_DUNSA (P54213) Caltractin (Centrin) 58 4e-09
CET2_MOUSE (Q9R1K9) Centrin 2 (Caltractin isoform 1) 57 6e-09
CET2_HUMAN (P41208) Centrin 2 (Caltractin isoform 1) 57 7e-09
CATR_TETST (P43646) Caltractin (Centrin) (Fragment) 57 7e-09
CATR_SCHDU (Q06827) Caltractin (Centrin) 57 7e-09
CATR_SPESI (P43645) Caltractin (Centrin) (Fragment) 56 1e-08
CET1_HUMAN (Q12798) Centrin 1 (Caltractin isoform 2) 55 4e-08
CC31_SCHPO (O74435) Cell division control protein 31 54 8e-08
CET1_MOUSE (P41209) Centrin 1 (Caltractin) 53 1e-07
CATR_CHLRE (P05434) Caltractin (Centrin) (20 kDa calcium-binding... 52 3e-07
CET3_MOUSE (O35648) Centrin 3 51 4e-07
CET3_HUMAN (O15182) Centrin 3 51 4e-07
CATR_NAEGR (P53441) Caltractin (Centrin) 49 2e-06
CC31_YEAST (P06704) Cell division control protein 31 (Nucleopori... 49 3e-06
CABO_LOLPE (P14533) Squidulin (Optic LOBE calcium-binding protei... 49 3e-06
CALM_PARTE (P07463) Calmodulin (CaM) 48 3e-06
CALM_BLAEM (Q9HFY6) Calmodulin (CaM) 48 3e-06
CATR_GIALA (Q24956) Caltractin (Centrin) 48 4e-06
CALM_SCHPO (P05933) Calmodulin (CaM) 47 1e-05
CALM_TETPY (P02598) Calmodulin (CaM) 46 1e-05
>CATR_DUNSA (P54213) Caltractin (Centrin)
Length = 169
Score = 57.8 bits (138), Expect = 4e-09
Identities = 34/98 (34%), Positives = 57/98 (57%), Gaps = 6/98 (6%)
Query: 2 AGTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVL 61
AG+GT+ +FE+ L +M +K+G +E+ F + D + G ITL++LK A L
Sbjct: 76 AGSGTI-----DFEEFLQMMTSKMGERDSREEIIKAFKLFDDDNTGFITLKNLKRVAKEL 130
Query: 62 GLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
G +++ ++EL M E D +GDG + + EF +M + S
Sbjct: 131 G-ENLTDEELQEMTDEADRNGDGQIDEDEFYRIMKKTS 167
Score = 31.6 bits (70), Expect = 0.33
Identities = 20/64 (31%), Positives = 29/64 (45%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I + LK LG + KE E+ MI + D G G + EF
Sbjct: 28 QEIREAFDLFDTDGSGTIDAKELKVAMRALGFEPKKE-EIKKMIADIDKAGSGTIDFEEF 86
Query: 92 CVLM 95
+M
Sbjct: 87 LQMM 90
>CET2_MOUSE (Q9R1K9) Centrin 2 (Caltractin isoform 1)
Length = 172
Score = 57.4 bits (137), Expect = 6e-09
Identities = 34/97 (35%), Positives = 53/97 (54%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
GTG +N F D L VM K+ + +E+ F + D + G I+ ++LK A LG
Sbjct: 80 GTGKMN-----FSDFLTVMTQKMSEKDTKEEILKAFKLFDDDETGKISFKNLKRVAKELG 134
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++EL MI E D DGDG + + EF +M + S
Sbjct: 135 -ENLTDEELQEMIDEADRDGDGEVNEQEFLRIMKKTS 170
Score = 34.3 bits (77), Expect = 0.051
Identities = 20/64 (31%), Positives = 31/64 (48%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I ++ LK LG + KE E+ MI E D +G G + +F
Sbjct: 31 QEIREAFDLFDADGTGTIDIKELKVAMRALGFEPKKE-EIKKMISEIDKEGTGKMNFSDF 89
Query: 92 CVLM 95
+M
Sbjct: 90 LTVM 93
>CET2_HUMAN (P41208) Centrin 2 (Caltractin isoform 1)
Length = 172
Score = 57.0 bits (136), Expect = 7e-09
Identities = 34/97 (35%), Positives = 54/97 (55%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
GTG +N F D L VM K+ + +E+ F + D + G I+ ++LK A LG
Sbjct: 80 GTGKMN-----FGDFLTVMTQKMSEKDTKEEILKAFKLFDDDETGKISFKNLKRVAKELG 134
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++EL MI E D DGDG +++ EF +M + S
Sbjct: 135 -ENLTDEELQEMIDEADRDGDGEVSEQEFLRIMKKTS 170
Score = 33.1 bits (74), Expect = 0.11
Identities = 20/64 (31%), Positives = 31/64 (48%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I ++ LK LG + KE E+ MI E D +G G + +F
Sbjct: 31 QEIREAFDLFDADGTGTIDVKELKVAMRALGFEPKKE-EIKKMISEIDKEGTGKMNFGDF 89
Query: 92 CVLM 95
+M
Sbjct: 90 LTVM 93
>CATR_TETST (P43646) Caltractin (Centrin) (Fragment)
Length = 148
Score = 57.0 bits (136), Expect = 7e-09
Identities = 34/97 (35%), Positives = 55/97 (56%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G+GT+ +FE+ L +M K+G +E+ F + D + G I+ ++LK A LG
Sbjct: 56 GSGTI-----DFEEFLQMMTAKMGERDSREEIMKAFRLFDDDETGKISFKNLKRVAKELG 110
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
++M ++EL MI E D DGDG + + EF +M + S
Sbjct: 111 -ENMTDEELQEMIDEADRDGDGEVNEEEFFRIMKKTS 146
Score = 33.5 bits (75), Expect = 0.086
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+++ FD+ G I + LK LG + KE E+ MI + D DG G + EF
Sbjct: 7 QDIREAFDLFDTDGSGTIDAKELKVAMRALGFEPKKE-EIKKMIADIDKDGSGTIDFEEF 65
Query: 92 CVLM 95
+M
Sbjct: 66 LQMM 69
>CATR_SCHDU (Q06827) Caltractin (Centrin)
Length = 168
Score = 57.0 bits (136), Expect = 7e-09
Identities = 34/97 (35%), Positives = 55/97 (56%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G+GT+ +FE+ L +M K+G +E+ F + D + G I+ ++LK A LG
Sbjct: 76 GSGTI-----DFEEFLQMMTAKMGERDSREEIMKAFRLFDDDETGKISFKNLKRVAKELG 130
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
++M ++EL MI E D DGDG + + EF +M + S
Sbjct: 131 -ENMTDEELQEMIDEADRDGDGEVNEEEFFRIMKKTS 166
Score = 34.7 bits (78), Expect = 0.039
Identities = 21/64 (32%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I + LK LG + KE E+ MI + D DG G + EF
Sbjct: 27 QEIREAFDLFDTDGSGTIDAKELKVAMRALGFEPKKE-EIKKMIADIDKDGSGTIDFEEF 85
Query: 92 CVLM 95
+M
Sbjct: 86 LQMM 89
>CATR_SPESI (P43645) Caltractin (Centrin) (Fragment)
Length = 148
Score = 56.2 bits (134), Expect = 1e-08
Identities = 33/97 (34%), Positives = 54/97 (55%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G+GT+ +FE+ L +M K+G +E+ F + D G IT ++LK A LG
Sbjct: 56 GSGTI-----DFEEFLQMMTAKMGERDSREEIMKAFRLFDDDQTGKITFKNLKRVAKELG 110
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++E+ MI E D DGDG + + EF +M + S
Sbjct: 111 -ENLTDEEIQEMIDEADRDGDGEINEEEFFRIMKKTS 146
Score = 34.3 bits (77), Expect = 0.051
Identities = 21/64 (32%), Positives = 29/64 (44%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E FD+ G I + LK LG + KE E+ MI + D DG G + EF
Sbjct: 7 QEXREAFDLFDTDGSGTIDAKELKVXMXALGFEPKKE-EIQKMISDIDKDGSGTIDFEEF 65
Query: 92 CVLM 95
+M
Sbjct: 66 LQMM 69
>CET1_HUMAN (Q12798) Centrin 1 (Caltractin isoform 2)
Length = 172
Score = 54.7 bits (130), Expect = 4e-08
Identities = 33/97 (34%), Positives = 53/97 (54%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
GTG ++ F D L VM K+ + +E+ F + D + G I+ ++LK A LG
Sbjct: 80 GTGKIS-----FNDFLAVMTQKMSEKDTKEEILKAFRLFDDDETGKISFKNLKRVANELG 134
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++EL MI E D DGDG + + EF +M + S
Sbjct: 135 -ENLTDEELQEMIDEADRDGDGEVNEEEFLRIMKKTS 170
Score = 32.7 bits (73), Expect = 0.15
Identities = 20/64 (31%), Positives = 31/64 (48%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I + LK LG + KE E+ MI E D +G G ++ +F
Sbjct: 31 QEVREAFDLFDVDGSGTIDAKELKVAMRALGFEPRKE-EMKKMISEVDREGTGKISFNDF 89
Query: 92 CVLM 95
+M
Sbjct: 90 LAVM 93
>CC31_SCHPO (O74435) Cell division control protein 31
Length = 176
Score = 53.5 bits (127), Expect = 8e-08
Identities = 31/88 (35%), Positives = 51/88 (57%), Gaps = 2/88 (2%)
Query: 9 GKG-LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMK 67
GKG L+ ED + VM K+ ++E+ F++ D + G I+L +L+ A L +++
Sbjct: 86 GKGYLQMEDFVRVMTEKIVERDPLEEIKRAFELFDDDETGKISLRNLRRVAKELN-ENID 144
Query: 68 EDELVSMIREGDLDGDGALTQMEFCVLM 95
+ EL +MI E DLD DG + + EF +M
Sbjct: 145 DQELEAMIEEFDLDQDGEINEQEFIAIM 172
>CET1_MOUSE (P41209) Centrin 1 (Caltractin)
Length = 172
Score = 53.1 bits (126), Expect = 1e-07
Identities = 29/84 (34%), Positives = 46/84 (54%), Gaps = 1/84 (1%)
Query: 12 LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDEL 71
+ F D L VM K+ + +E+ F + D + G I+ ++LK A LG + + ++EL
Sbjct: 84 ISFNDFLAVMTQKMAEKDTKEEILKAFRLFDDDETGKISFKNLKRVANELG-ESLTDEEL 142
Query: 72 VSMIREGDLDGDGALTQMEFCVLM 95
MI E D DGDG + + EF +M
Sbjct: 143 QEMIDEADRDGDGEVNEEEFLKIM 166
Score = 32.3 bits (72), Expect = 0.19
Identities = 19/64 (29%), Positives = 31/64 (47%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I ++ LK LG + KE E+ MI E D + G ++ +F
Sbjct: 31 QEVREAFDLFDSDGSGTIDVKELKVAMRALGFEPRKE-EMKKMISEVDKEATGKISFNDF 89
Query: 92 CVLM 95
+M
Sbjct: 90 LAVM 93
>CATR_CHLRE (P05434) Caltractin (Centrin) (20 kDa calcium-binding
protein)
Length = 169
Score = 51.6 bits (122), Expect = 3e-07
Identities = 31/97 (31%), Positives = 53/97 (53%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G+GT+ +FE+ L +M K+G +E+ F + D + G IT++ L+ A LG
Sbjct: 77 GSGTI-----DFEEFLTMMTAKMGERDSREEILKAFRLFDDDNSGTITIKDLRRVAKELG 131
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ E+EL MI E D + D + + EF +M + S
Sbjct: 132 -ENLTEEELQEMIAEADRNDDNEIDEDEFIRIMKKTS 167
Score = 36.6 bits (83), Expect = 0.010
Identities = 22/64 (34%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I + LK LG + KE E+ MI E D DG G + EF
Sbjct: 28 QEIREAFDLFDTDGSGTIDAKELKVAMRALGFEPKKE-EIKKMISEIDKDGSGTIDFEEF 86
Query: 92 CVLM 95
+M
Sbjct: 87 LTMM 90
>CET3_MOUSE (O35648) Centrin 3
Length = 167
Score = 51.2 bits (121), Expect = 4e-07
Identities = 30/84 (35%), Positives = 47/84 (55%), Gaps = 1/84 (1%)
Query: 12 LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDEL 71
+ FED V+ + + +E+ F + D D G I+L +L+ A LG ++M ++EL
Sbjct: 81 ITFEDFNEVVTDWILERDPHEEILKAFKLFDDDDSGKISLRNLRRVARELG-ENMSDEEL 139
Query: 72 VSMIREGDLDGDGALTQMEFCVLM 95
+MI E D DGDG + Q EF +M
Sbjct: 140 RAMIEEFDKDGDGEINQEEFIAIM 163
>CET3_HUMAN (O15182) Centrin 3
Length = 167
Score = 51.2 bits (121), Expect = 4e-07
Identities = 30/84 (35%), Positives = 47/84 (55%), Gaps = 1/84 (1%)
Query: 12 LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDEL 71
+ FED V+ + + +E+ F + D D G I+L +L+ A LG ++M ++EL
Sbjct: 81 ITFEDFNEVVTDWILERDPHEEILKAFKLFDDDDSGKISLRNLRRVARELG-ENMSDEEL 139
Query: 72 VSMIREGDLDGDGALTQMEFCVLM 95
+MI E D DGDG + Q EF +M
Sbjct: 140 RAMIEEFDKDGDGEINQEEFIAIM 163
>CATR_NAEGR (P53441) Caltractin (Centrin)
Length = 172
Score = 49.3 bits (116), Expect = 2e-06
Identities = 30/93 (32%), Positives = 49/93 (52%), Gaps = 2/93 (2%)
Query: 4 TGTVNGKG-LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
+G NG G ++F D L +M K+ + E+ F + + D G IT +LK A LG
Sbjct: 75 SGIDNGSGKIDFNDFLQLMTAKMSEKDSHAEIMKAFRLFDEDDSGFITFANLKRVAKDLG 134
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
++M ++EL MI E D G +++ +F +M
Sbjct: 135 -ENMTDEELREMIEEADRSNQGQISKEDFLRIM 166
>CC31_YEAST (P06704) Cell division control protein 31 (Nucleoporin
CDC31) (Nuclear pore protein CDC31)
Length = 161
Score = 48.5 bits (114), Expect = 3e-06
Identities = 26/83 (31%), Positives = 47/83 (56%), Gaps = 1/83 (1%)
Query: 12 LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDEL 71
++++D VM K+ + E+ F + D G I++++L+ A LG + + ++EL
Sbjct: 76 MKYDDFYIVMGEKILKRDPLDEIKRAFQLFDDDHTGKISIKNLRRVAKELG-ETLTDEEL 134
Query: 72 VSMIREGDLDGDGALTQMEFCVL 94
+MI E DLDGDG + + EF +
Sbjct: 135 RAMIEEFDLDGDGEINENEFIAI 157
>CABO_LOLPE (P14533) Squidulin (Optic LOBE calcium-binding protein)
(SCABP)
Length = 149
Score = 48.5 bits (114), Expect = 3e-06
Identities = 28/93 (30%), Positives = 46/93 (49%), Gaps = 5/93 (5%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ E+ + + +MA ++G KE+ F + G+IT L+ A
Sbjct: 59 GNGTI-----EYAEFVEMMAKQMGPTDPEKEMREAFRVFDKDGNGLITAAELRQVMANFS 113
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + +E+ MIRE D+DGDG + EF +M
Sbjct: 114 DEKLTSEEISEMIREADIDGDGMVNYEEFVKMM 146
Score = 37.4 bits (85), Expect = 0.006
Identities = 25/68 (36%), Positives = 35/68 (50%), Gaps = 1/68 (1%)
Query: 28 EGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALT 87
E I E+ + FDM G IT + L++ LG + + EL MIRE D DG+G +
Sbjct: 6 EKQIAEIKDAFDMFDIDGDGQITSKELRSVMKSLG-RTPSDAELEEMIREVDTDGNGTIE 64
Query: 88 QMEFCVLM 95
EF +M
Sbjct: 65 YAEFVEMM 72
>CALM_PARTE (P07463) Calmodulin (CaM)
Length = 148
Score = 48.1 bits (113), Expect = 3e-06
Identities = 30/93 (32%), Positives = 48/93 (51%), Gaps = 6/93 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ + +EL F + G+I+ L+ LG
Sbjct: 59 GNGTI-----DFPEFLSLMARKMKEQDSEEELIEAFKVFDRDGNGLISAAELRHVMTNLG 113
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + +DE+ MIRE D+DGDG + EF +M
Sbjct: 114 -EKLTDDEVDEMIREADIDGDGHINYEEFVRMM 145
Score = 37.4 bits (85), Expect = 0.006
Identities = 28/62 (45%), Positives = 32/62 (51%), Gaps = 2/62 (3%)
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
L DKD G IT + L T LG Q+ E EL MI E D DG+G + EF LM R
Sbjct: 18 LFDKDGDGTITTKELGTVMRSLG-QNPTEAELQDMINEVDADGNGTIDFPEFLSLMARKM 76
Query: 100 PE 101
E
Sbjct: 77 KE 78
>CALM_BLAEM (Q9HFY6) Calmodulin (CaM)
Length = 148
Score = 48.1 bits (113), Expect = 3e-06
Identities = 30/93 (32%), Positives = 46/93 (49%), Gaps = 6/93 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +E+ F + G I+ L+ LG
Sbjct: 59 GNGTI-----DFPEFLTMMARKMKDSDSEEEIKEAFKVFDKDGNGYISAAELRHVMTNLG 113
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + EDE+ MIRE D+DGDG + EF +M
Sbjct: 114 -EKLSEDEVEEMIREADVDGDGQINYEEFVKMM 145
Score = 37.4 bits (85), Expect = 0.006
Identities = 26/58 (44%), Positives = 32/58 (54%), Gaps = 2/58 (3%)
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFR 97
L DKD G IT + L T LG Q+ E EL+ MI E D DG+G + EF +M R
Sbjct: 18 LFDKDGDGTITTKELGTVMRSLG-QNPTEAELLVMINEVDADGNGTIDFPEFLTMMAR 74
>CATR_GIALA (Q24956) Caltractin (Centrin)
Length = 176
Score = 47.8 bits (112), Expect = 4e-06
Identities = 25/84 (29%), Positives = 48/84 (56%), Gaps = 1/84 (1%)
Query: 12 LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDEL 71
+ F+D VM K+ +E+ F + D G I+L++L+ A L +++ ++EL
Sbjct: 87 ITFQDFEEVMIEKISNRDPTEEILKAFRLFDDDATGRISLKNLRRVAKELS-ENISDEEL 145
Query: 72 VSMIREGDLDGDGALTQMEFCVLM 95
++MI+E D DGDG + + +F ++
Sbjct: 146 LAMIQEFDRDGDGEIDEEDFIAIL 169
>CALM_SCHPO (P05933) Calmodulin (CaM)
Length = 150
Score = 46.6 bits (109), Expect = 1e-05
Identities = 29/89 (32%), Positives = 44/89 (48%), Gaps = 6/89 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +E+ F + G IT+E L LG
Sbjct: 61 GNGTI-----DFTEFLTMMARKMKDTDNEEEVREAFKVFDKDGNGYITVEELTHVLTSLG 115
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEF 91
+ + ++E+ MIRE D DGDG + EF
Sbjct: 116 -ERLSQEEVADMIREADTDGDGVINYEEF 143
Score = 30.8 bits (68), Expect = 0.56
Identities = 23/58 (39%), Positives = 28/58 (47%), Gaps = 2/58 (3%)
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFR 97
L D+D+ G IT L LG Q EL MI E D DG+G + EF +M R
Sbjct: 20 LFDRDQDGNITSNELGVVMRSLG-QSPTAAELQDMINEVDADGNGTIDFTEFLTMMAR 76
>CALM_TETPY (P02598) Calmodulin (CaM)
Length = 148
Score = 46.2 bits (108), Expect = 1e-05
Identities = 29/93 (31%), Positives = 47/93 (50%), Gaps = 6/93 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +EL F + G+I+ L+ LG
Sbjct: 59 GNGTI-----DFPEFLSLMARKMKDTDTEEELIEAFKVFDRDGNGLISAAELRHVMTNLG 113
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MIRE D+DGDG + EF +M
Sbjct: 114 -EKLTDEEVDEMIREADIDGDGHINYEEFVRMM 145
Score = 37.0 bits (84), Expect = 0.008
Identities = 27/58 (46%), Positives = 31/58 (52%), Gaps = 2/58 (3%)
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFR 97
L DKD G IT + L T LG Q+ E EL MI E D DG+G + EF LM R
Sbjct: 18 LFDKDGDGTITTKELGTVMRSLG-QNPTEAELQDMINEVDADGNGTIDFPEFLSLMAR 74
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.138 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,524,435
Number of Sequences: 164201
Number of extensions: 614558
Number of successful extensions: 2161
Number of sequences better than 10.0: 267
Number of HSP's better than 10.0 without gapping: 176
Number of HSP's successfully gapped in prelim test: 91
Number of HSP's that attempted gapping in prelim test: 1757
Number of HSP's gapped (non-prelim): 477
length of query: 124
length of database: 59,974,054
effective HSP length: 100
effective length of query: 24
effective length of database: 43,553,954
effective search space: 1045294896
effective search space used: 1045294896
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0203.8


