

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
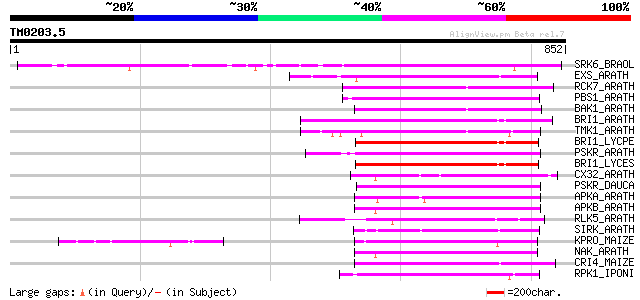
Query= TM0203.5
(852 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 495 e-139
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 232 4e-60
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 218 5e-56
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 214 6e-55
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 214 8e-55
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 209 2e-53
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 208 6e-53
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 207 7e-53
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 207 1e-52
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 206 2e-52
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 205 5e-52
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 196 3e-49
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 193 2e-48
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 190 2e-47
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 187 8e-47
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 181 9e-45
KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1 precu... 179 3e-44
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 179 4e-44
CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4 pr... 176 2e-43
RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC 2... 175 5e-43
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 495 bits (1274), Expect = e-139
Identities = 331/868 (38%), Positives = 477/868 (54%), Gaps = 74/868 (8%)
Query: 13 FLFHMHCWLLCFSQLCFAGDTLNVGQEIT-GNGTVLVSAAKKFELGFFSPDLNVTGGKGR 71
FL +L L +TL+ + +T + LVS FE+GFF +
Sbjct: 15 FLLVFVVMILIHPALSIYINTLSSTESLTISSNKTLVSPGSIFEVGFFRTNSRW------ 68
Query: 72 YLGIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLT 131
YLG+WY + VWVANRDNP+++ +IG +I+ + NLV+LD S W +NLT
Sbjct: 69 YLGMWYKKVSDR---TYVWVANRDNPLSN-AIGTLKISGN-NLVLLDHSNKPVWW-TNLT 122
Query: 132 NSSSATNRSVKLMDSGNLVLLDEH---VGMKLWESFEHPTDTFLPGMKMDKTLE------ 182
+ + +L+ +GN V+ D LW+SF++PTDT LP MK+ L+
Sbjct: 123 RGNERSPVVAELLANGNFVMRDSSNNDASEYLWQSFDYPTDTLLPEMKLGYNLKTGLNRF 182
Query: 183 LTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDISNDV 242
LT W+S DP GNF++K++ + F + + +S + +S V
Sbjct: 183 LTSWRSSDDPSSGNFSYKLETQSLPEFYLSRENFPMHRSGPWNGIRFSGIPEDQKLSYMV 242
Query: 243 YNLLTNFKELKN--KTVSSYDNTRLLLNSTGVIKVLYRVNFQSDIVW---WYQP-RTTCL 296
YN + N +E+ + ++ +RL L S G + L + S +W W P C
Sbjct: 243 YNFIENNEEVAYTFRMTNNSFYSRLTLISEGYFQRL--TWYPSIRIWNRFWSSPVDPQCD 300
Query: 297 TYNVCGNFSSCNDDNDKLCTCLPGFGRRSPLN-DYTV--GGDTSSLLCTRKSTSCGANTN 353
TY +CG ++ C+ + +C C+ GF R+ D V GG C R+ T + +
Sbjct: 301 TYIMCGPYAYCDVNTSPVCNCIQGFNPRNIQQWDQRVWAGG------CIRR-TQLSCSGD 353
Query: 354 TFLNLTMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPIPVQQRGLNLS 409
F + MK+ P+ ++ D + ECK RCIS C+ T ++ ++ G S
Sbjct: 354 GFTRMKKMKL--PETTMATVDRSIGVKECKKRCISDCNCT-----AFANADIRNGG---S 403
Query: 410 PCWIWTQNLTTLKEEYLGG-DDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVI 468
C IWT+ L ++ D + L+VR+A +DI + ++ +SL +G+++ ++I
Sbjct: 404 GCVIWTERLEDIRNYATDAIDGQDLYVRLAAADIAK--KRNASGKIISLTVGVSVLLLLI 461
Query: 469 LACICILAYVCRRKIALKLKQESESILR-QRGRFYDSERHVKDLIDKEGLEEKDNEGIEV 527
+ C+ ++ + K + SI QR + V +E E E +E+
Sbjct: 462 MFCLW-------KRKQKRAKASAISIANTQRNQNLPMNEMVLSS-KREFSGEYKFEELEL 513
Query: 528 PYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEV 587
P + E+++ AT+ FS NKLG+GG+G VYKG+L G+EIAVKRLS S QG EF NEV
Sbjct: 514 PLIEMETVVKATENFSSCNKLGQGGFGIVYKGRLLDGKEIAVKRLSKTSVQGTDEFMNEV 573
Query: 588 VLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLG 647
LIA+LQH NLV++ G CI+GDEK+LIYEY+ N SLD+++F T+ + L+W RFDI G
Sbjct: 574 TLIARLQHINLVQVLGCCIEGDEKMLIYEYLENLSLDSYLFGKTRRSKLNWNERFDITNG 633
Query: 648 IARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGT 707
+ARGLLYLHQDSR R+IHRDLK SNILLD M PKISDFG+ARIF ETEANT +VVGT
Sbjct: 634 VARGLLYLHQDSRFRIIHRDLKVSNILLDKNMIPKISDFGMARIFERDETEANTMKVVGT 693
Query: 708 YGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENK 767
YGYMSPEYA+ G FS KSD+FSFGV++LEI+SGKKN GFY LL Y W W E +
Sbjct: 694 YGYMSPEYAMYGIFSEKSDVFSFGVIVLEIVSGKKNRGFYNLDYENDLLSYVWSRWKEGR 753
Query: 768 LLDLMD-------LSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATL 820
L+++D S + + ++C +GLLCVQ+ + RP MS+VV M SE +
Sbjct: 754 ALEIVDPVIVDSLSSQPSIFQPQEVLKCIQIGLLCVQELAEHRPAMSSVVWMFGSEATEI 813
Query: 821 PTPKQPTFFTRKD-LSSTASSSLQFDSS 847
P PK P + R+ SSS Q D +
Sbjct: 814 PQPKPPGYCVRRSPYELDPSSSWQCDEN 841
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 232 bits (591), Expect = 4e-60
Identities = 141/392 (35%), Positives = 224/392 (56%), Gaps = 21/392 (5%)
Query: 430 DRKLFVRVAKSDIE-EPTRKGNPKSTLSLILGIALPGVVILACICILAYVCRRKIALKLK 488
+++L RV SD + E T+ + L+LG + +V + + + +++ +
Sbjct: 805 NKELCGRVVGSDCKIEGTKLRSAWGIAGLMLGFTI--IVFVFVFSLRRWAMTKRVKQRDD 862
Query: 489 QESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFD-------FESILVATDY 541
E R +G F D ++L G ++ I + F+ I+ ATD+
Sbjct: 863 PERMEESRLKG-FVD-----QNLYFLSGSRSREPLSINIAMFEQPLLKVRLGDIVEATDH 916
Query: 542 FSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRL 601
FS N +G GG+G VYK L G + +AVK+LS +QG +EF E+ + K++H NLV L
Sbjct: 917 FSKKNIIGDGGFGTVYKACLPGEKTVAVKKLSEAKTQGNREFMAEMETLGKVKHPNLVSL 976
Query: 602 WGYCIKGDEKILIYEYMPNKSLDAFVFDPT-KSALLDWQMRFDILLGIARGLLYLHQDSR 660
GYC +EK+L+YEYM N SLD ++ + T +LDW R I +G ARGL +LH
Sbjct: 977 LGYCSFSEEKLLVYEYMVNGSLDHWLRNQTGMLEVLDWSKRLKIAVGAARGLAFLHHGFI 1036
Query: 661 LRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQ 720
+IHRD+K SNILLDG+ +PK++DFGLAR+ E+ +T + GT+GY+ PEY +
Sbjct: 1037 PHIIHRDIKASNILLDGDFEPKVADFGLARLISACESHVSTV-IAGTFGYIPPEYGQSAR 1095
Query: 721 FSTKSDIFSFGVVLLEIISGKKNTG--FYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGE 778
+TK D++SFGV+LLE+++GK+ TG F + +G +L+G+A + + K +D++D L
Sbjct: 1096 ATTKGDVYSFGVILLELVTGKEPTGPDFKESEGG-NLVGWAIQKINQGKAVDVIDPLLVS 1154
Query: 779 AYNANQFIRCTHVGLLCVQDEPDDRPNMSNVV 810
N +R + +LC+ + P RPNM +V+
Sbjct: 1155 VALKNSQLRLLQIAMLCLAETPAKRPNMLDVL 1186
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 218 bits (555), Expect = 5e-56
Identities = 130/329 (39%), Positives = 197/329 (59%), Gaps = 6/329 (1%)
Query: 512 IDKEGLEEKDN-EGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQG-GREIAV 569
+D +GL D G + F F+ + AT F LG GG+G V+KG ++ + +A+
Sbjct: 72 LDVKGLNLNDQVTGKKAQTFTFQELAEATGNFRSDCFLGEGGFGKVFKGTIEKLDQVVAI 131
Query: 570 KRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFD 629
K+L QGI+EF EV+ ++ H NLV+L G+C +GD+++L+YEYMP SL+ +
Sbjct: 132 KQLDRNGVQGIREFVVEVLTLSLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHV 191
Query: 630 -PTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGL 688
P+ LDW R I G ARGL YLH VI+RDLK SNILL + QPK+SDFGL
Sbjct: 192 LPSGKKPLDWNTRMKIAAGAARGLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGL 251
Query: 689 ARIF-GGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFY 747
A++ G +T +T RV+GTYGY +P+YA+ GQ + KSDI+SFGVVLLE+I+G+K
Sbjct: 252 AKVGPSGDKTHVST-RVMGTYGYCAPDYAMTGQLTFKSDIYSFGVVLLELITGRKAIDNT 310
Query: 748 QYKGTLSLLGYAWKLWTENK-LLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNM 806
+ + +L+G+A L+ + + ++D L Y + + +CVQ++P RP +
Sbjct: 311 KTRKDQNLVGWARPLFKDRRNFPKMVDPLLQGQYPVRGLYQALAISAMCVQEQPTMRPVV 370
Query: 807 SNVVIMLDSETATLPTPKQPTFFTRKDLS 835
S+VV+ L+ ++ P P+ + K+ S
Sbjct: 371 SDVVLALNFLASSKYDPNSPSSSSGKNPS 399
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 214 bits (546), Expect = 6e-55
Identities = 123/306 (40%), Positives = 179/306 (58%), Gaps = 8/306 (2%)
Query: 511 LIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQG-GREIAV 569
L+ ++GL + I F F + AT F LG GG+G VYKG+L G+ +AV
Sbjct: 60 LLPRDGLGQ-----IAAHTFAFRELAAATMNFHPDTFLGEGGFGRVYKGRLDSTGQVVAV 114
Query: 570 KRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFD 629
K+L QG +EF EV++++ L H NLV L GYC GD+++L+YE+MP SL+ + D
Sbjct: 115 KQLDRNGLQGNREFLVEVLMLSLLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHD 174
Query: 630 -PTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGL 688
P LDW MR I G A+GL +LH + VI+RD K+SNILLD PK+SDFGL
Sbjct: 175 LPPDKEALDWNMRMKIAAGAAKGLEFLHDKANPPVIYRDFKSSNILLDEGFHPKLSDFGL 234
Query: 689 ARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQ 748
A++ + + RV+GTYGY +PEYA+ GQ + KSD++SFGVV LE+I+G+K
Sbjct: 235 AKLGPTGDKSHVSTRVMGTYGYCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDSEM 294
Query: 749 YKGTLSLLGYAWKLWTE-NKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMS 807
G +L+ +A L+ + K + L D L + + V +C+Q++ RP ++
Sbjct: 295 PHGEQNLVAWARPLFNDRRKFIKLADPRLKGRFPTRALYQALAVASMCIQEQAATRPLIA 354
Query: 808 NVVIML 813
+VV L
Sbjct: 355 DVVTAL 360
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 214 bits (545), Expect = 8e-55
Identities = 122/291 (41%), Positives = 175/291 (59%), Gaps = 5/291 (1%)
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQ-EFKNEVV 588
F + VA+D FS+ N LGRGG+G VYKG+L G +AVKRL +QG + +F+ EV
Sbjct: 277 FSLRELQVASDNFSNKNILGRGGFGKVYKGRLADGTLVAVKRLKEERTQGGELQFQTEVE 336
Query: 589 LIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFD-PTKSALLDWQMRFDILLG 647
+I+ HRNL+RL G+C+ E++L+Y YM N S+ + + + P LDW R I LG
Sbjct: 337 MISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPESQPPLDWPKRQRIALG 396
Query: 648 IARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGT 707
ARGL YLH ++IHRD+K +NILLD E + + DFGLA++ K+T T V GT
Sbjct: 397 SARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTT-AVRGT 455
Query: 708 YGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKG--TLSLLGYAWKLWTE 765
G+++PEY G+ S K+D+F +GV+LLE+I+G++ + + LL + L E
Sbjct: 456 IGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVMLLDWVKGLLKE 515
Query: 766 NKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSE 816
KL L+D+ L Y + + V LLC Q P +RP MS VV ML+ +
Sbjct: 516 KKLEALVDVDLQGNYKDEEVEQLIQVALLCTQSSPMERPKMSEVVRMLEGD 566
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 209 bits (532), Expect = 2e-53
Identities = 135/396 (34%), Positives = 214/396 (53%), Gaps = 11/396 (2%)
Query: 447 RKGNPKSTLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSER 506
++ + + SL +A+ + CI L V R + K+E+E + G +R
Sbjct: 782 QRSHGRRPASLAGSVAMGLLFSFVCIFGLILVGREMRKRRRKKEAELEMYAEGHGNSGDR 841
Query: 507 HVKDLIDK-EGLEEK---DNEGIEVPY--FDFESILVATDYFSDANKLGRGGYGPVYKGK 560
+ K G++E + E P F +L AT+ F + + +G GG+G VYK
Sbjct: 842 TANNTNWKLTGVKEALSINLAAFEKPLRKLTFADLLQATNGFHNDSLIGSGGFGDVYKAI 901
Query: 561 LQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPN 620
L+ G +A+K+L VS QG +EF E+ I K++HRNLV L GYC GDE++L+YE+M
Sbjct: 902 LKDGSAVAIKKLIHVSGQGDREFMAEMETIGKIKHRNLVPLLGYCKVGDERLLVYEFMKY 961
Query: 621 KSLDAFVFDPTKSAL-LDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEM 679
SL+ + DP K+ + L+W R I +G ARGL +LH + +IHRD+K+SN+LLD +
Sbjct: 962 GSLEDVLHDPKKAGVKLNWSTRRKIAIGSARGLAFLHHNCSPHIIHRDMKSSNVLLDENL 1021
Query: 680 QPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIIS 739
+ ++SDFG+AR+ +T + + GT GY+ PEY + STK D++S+GVVLLE+++
Sbjct: 1022 EARVSDFGMARLMSAMDTHLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLT 1081
Query: 740 GKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTH--VGLLCVQ 797
GK+ T + G +L+G+ K + ++ D+ D L + A + H V + C+
Sbjct: 1082 GKRPTDSPDF-GDNNLVGWV-KQHAKLRISDVFDPELMKEDPALEIELLQHLKVAVACLD 1139
Query: 798 DEPDDRPNMSNVVIMLDSETATLPTPKQPTFFTRKD 833
D RP M V+ M A Q T + +D
Sbjct: 1140 DRAWRRPTMVQVMAMFKEIQAGSGIDSQSTIRSIED 1175
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 208 bits (529), Expect = 6e-53
Identities = 138/400 (34%), Positives = 223/400 (55%), Gaps = 35/400 (8%)
Query: 447 RKGNPKST-LSLILGIALPGVVILACICILAYVCRRKIALKLKQESES----ILRQRGRF 501
R+G ST + +I+G L G++ + I +L + C K K SES ++ R
Sbjct: 472 RRGMKSSTFIGIIVGSVLGGLLSIFLIGLLVF-CWYKKRQKRFSGSESSNAVVVHPRHSG 530
Query: 502 YDSER----------HVKDLIDKEGLEEKDNEGIEVPYFDFESILVA-------TDYFSD 544
D+E V + D L G + + ++L++ T+ FS
Sbjct: 531 SDNESVKITVAGSSVSVGGISDTYTLPGTSEVGDNIQMVEAGNMLISIQVLRSVTNNFSS 590
Query: 545 ANKLGRGGYGPVYKGKLQGGREIAVKRLSS--VSSQGIQEFKNEVVLIAKLQHRNLVRLW 602
N LG GG+G VYKG+L G +IAVKR+ + ++ +G EFK+E+ ++ K++HR+LV L
Sbjct: 591 DNILGSGGFGVVYKGELHDGTKIAVKRMENGVIAGKGFAEFKSEIAVLTKVRHRHLVTLL 650
Query: 603 GYCIKGDEKILIYEYMPNKSLDAFVFDPTKSAL--LDWQMRFDILLGIARGLLYLHQDSR 660
GYC+ G+EK+L+YEYMP +L +F+ ++ L L W+ R + L +ARG+ YLH +
Sbjct: 651 GYCLDGNEKLLVYEYMPQGTLSRHLFEWSEEGLKPLLWKQRLTLALDVARGVEYLHGLAH 710
Query: 661 LRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQ 720
IHRDLK SNILL +M+ K++DFGL R+ + T R+ GT+GY++PEYA+ G+
Sbjct: 711 QSFIHRDLKPSNILLGDDMRAKVADFGLVRLAPEGKGSIET-RIAGTFGYLAPEYAVTGR 769
Query: 721 FSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTE-----NKLLDLMDLS 775
+TK D++SFGV+L+E+I+G+K+ Q + ++ L+ + +++ K +D +
Sbjct: 770 VTTKVDVYSFGVILMELITGRKSLDESQPEESIHLVSWFKRMYINKEASFKKAID-TTID 828
Query: 776 LGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDS 815
L E A+ G C + EP RP+M + V +L S
Sbjct: 829 LDEETLASVHTVAELAGHCCAR-EPYQRPDMGHAVNILSS 867
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 207 bits (528), Expect = 7e-53
Identities = 113/284 (39%), Positives = 175/284 (60%), Gaps = 5/284 (1%)
Query: 532 FESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIA 591
F +L AT+ F + + +G GG+G VYK +L+ G +A+K+L VS QG +EF E+ I
Sbjct: 878 FADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDREFTAEMETIG 937
Query: 592 KLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSAL-LDWQMRFDILLGIAR 650
K++HRNLV L GYC G+E++L+YEYM SL+ + D K+ + L+W R I +G AR
Sbjct: 938 KIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIKLNWPARRKIAIGAAR 997
Query: 651 GLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGY 710
GL +LH + +IHRD+K+SN+LLD ++ ++SDFG+AR+ +T + + GT GY
Sbjct: 998 GLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGTPGY 1057
Query: 711 MSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLD 770
+ PEY + STK D++S+GVVLLE+++GK+ T + G +L+G+ KL + K+ D
Sbjct: 1058 VPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADF-GDNNLVGWV-KLHAKGKITD 1115
Query: 771 LMDLSL--GEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIM 812
+ D L +A + ++ V C+ D RP M V+ M
Sbjct: 1116 VFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAM 1159
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 207 bits (526), Expect = 1e-52
Identities = 124/363 (34%), Positives = 196/363 (53%), Gaps = 17/363 (4%)
Query: 455 LSLILGIALPGVVILACICILAYVCRRKIALKLKQ--ESESILRQRGRFYDSERHVKDLI 512
+ + +GIA V +L + ++ RR+ + ESES+ R+ S+ V
Sbjct: 658 IGMAIGIAFGSVFLLTLLSLIVLRARRRSGEVDPEIEESESMNRKELGEIGSKLVVL--- 714
Query: 513 DKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRL 572
+ D E ++ +L +T+ F AN +G GG+G VYK L G+++A+K+L
Sbjct: 715 ----FQSNDKE------LSYDDLLDSTNSFDQANIIGCGGFGMVYKATLPDGKKVAIKKL 764
Query: 573 SSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTK 632
S Q +EF+ EV +++ QH NLV L G+C ++++LIY YM N SLD ++ +
Sbjct: 765 SGDCGQIEREFEAEVETLSRAQHPNLVLLRGFCFYKNDRLLIYSYMENGSLDYWLHERND 824
Query: 633 S-ALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARI 691
ALL W+ R I G A+GLLYLH+ ++HRD+K+SNILLD ++DFGLAR+
Sbjct: 825 GPALLKWKTRLRIAQGAAKGLLYLHEGCDPHILHRDIKSSNILLDENFNSHLADFGLARL 884
Query: 692 FGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKG 751
ET +T +VGT GY+ PEY + K D++SFGVVLLE+++ K+ + KG
Sbjct: 885 MSPYETHVSTD-LVGTLGYIPPEYGQASVATYKGDVYSFGVVLLELLTDKRPVDMCKPKG 943
Query: 752 TLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVI 811
L+ + K+ E++ ++ D + N + R + LC+ + P RP +V
Sbjct: 944 CRDLISWVVKMKHESRASEVFDPLIYSKENDKEMFRVLEIACLCLSENPKQRPTTQQLVS 1003
Query: 812 MLD 814
LD
Sbjct: 1004 WLD 1006
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 206 bits (525), Expect = 2e-52
Identities = 113/284 (39%), Positives = 174/284 (60%), Gaps = 5/284 (1%)
Query: 532 FESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIA 591
F +L AT+ F + + +G GG+G VYK +L+ G +A+K+L VS QG +EF E+ I
Sbjct: 878 FADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDREFTAEMETIG 937
Query: 592 KLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSAL-LDWQMRFDILLGIAR 650
K++HRNLV L GYC G+E++L+YEYM SL+ + D K + L+W R I +G AR
Sbjct: 938 KIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIKLNWPARRKIAIGAAR 997
Query: 651 GLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGY 710
GL +LH + +IHRD+K+SN+LLD ++ ++SDFG+AR+ +T + + GT GY
Sbjct: 998 GLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGTPGY 1057
Query: 711 MSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLD 770
+ PEY + STK D++S+GVVLLE+++GK+ T + G +L+G+ KL + K+ D
Sbjct: 1058 VPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADF-GDNNLVGWV-KLHAKGKITD 1115
Query: 771 LMDLSL--GEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIM 812
+ D L +A + ++ V C+ D RP M V+ M
Sbjct: 1116 VFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAM 1159
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 205 bits (521), Expect = 5e-52
Identities = 118/330 (35%), Positives = 188/330 (56%), Gaps = 16/330 (4%)
Query: 523 EGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKG----------KLQGGREIAVKRL 572
E + ++F + AT F + LG+GG+G VY+G ++ G +A+KRL
Sbjct: 67 ESPNLKVYNFLDLKTATKNFKPDSMLGQGGFGKVYRGWVDATTLAPSRVGSGMIVAIKRL 126
Query: 573 SSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTK 632
+S S QG E+++EV + L HRNLV+L GYC + E +L+YE+MP SL++ +F +
Sbjct: 127 NSESVQGFAEWRSEVNFLGMLSHRNLVKLLGYCREDKELLLVYEFMPKGSLESHLF--RR 184
Query: 633 SALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIF 692
+ W +R I++G ARGL +LH R VI+RD K SNILLD K+SDFGLA++
Sbjct: 185 NDPFPWDLRIKIVIGAARGLAFLHSLQR-EVIYRDFKASNILLDSNYDAKLSDFGLAKLG 243
Query: 693 GGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGT 752
E T R++GTYGY +PEY G KSD+F+FGVVLLEI++G + +G
Sbjct: 244 PADEKSHVTTRIMGTYGYAAPEYMATGHLYVKSDVFAFGVVLLEIMTGLTAHNTKRPRGQ 303
Query: 753 LSLLGYAW-KLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVI 811
SL+ + +L ++++ +MD + Y + L C++ +P +RP+M VV
Sbjct: 304 ESLVDWLRPELSNKHRVKQIMDKGIKGQYTTKVATEMARITLSCIEPDPKNRPHMKEVVE 363
Query: 812 MLDSETATLPTPKQPTFFTRKDLSSTASSS 841
+L+ P + + T++ +++++ SS
Sbjct: 364 VLEHIQGLNVVPNRSS--TKQAVANSSRSS 391
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 196 bits (497), Expect = 3e-49
Identities = 105/284 (36%), Positives = 165/284 (57%), Gaps = 2/284 (0%)
Query: 533 ESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAK 592
+ IL +T F+ AN +G GG+G VYK L G ++A+KRLS + Q +EF+ EV +++
Sbjct: 734 DDILKSTSSFNQANIIGCGGFGLVYKATLPDGTKVAIKRLSGDTGQMDREFQAEVETLSR 793
Query: 593 LQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKS-ALLDWQMRFDILLGIARG 651
QH NLV L GYC ++K+LIY YM N SLD ++ + LDW+ R I G A G
Sbjct: 794 AQHPNLVHLLGYCNYKNDKLLIYSYMDNGSLDYWLHEKVDGPPSLDWKTRLRIARGAAEG 853
Query: 652 LLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYM 711
L YLHQ ++HRD+K+SNILL ++DFGLAR+ +T T +VGT GY+
Sbjct: 854 LAYLHQSCEPHILHRDIKSSNILLSDTFVAHLADFGLARLILPYDTHVTTD-LVGTLGYI 912
Query: 712 SPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDL 771
PEY + K D++SFGVVLLE+++G++ + +G+ L+ + ++ TE + ++
Sbjct: 913 PPEYGQASVATYKGDVYSFGVVLLELLTGRRPMDVCKPRGSRDLISWVLQMKTEKRESEI 972
Query: 772 MDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDS 815
D + + +A + + + C+ + P RP +V L++
Sbjct: 973 FDPFIYDKDHAEEMLLVLEIACRCLGENPKTRPTTQQLVSWLEN 1016
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 193 bits (490), Expect = 2e-48
Identities = 117/300 (39%), Positives = 174/300 (58%), Gaps = 19/300 (6%)
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQG----------GREIAVKRLSSVSSQG 579
F F + AT F + LG GG+G V+KG + G IAVK+L+ QG
Sbjct: 56 FSFAELKSATRNFRPDSVLGEGGFGCVFKGWIDEKSLTASRPGTGLVIAVKKLNQDGWQG 115
Query: 580 IQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSAL---- 635
QE+ EV + + HR+LV+L GYC++ + ++L+YE+MP SL+ +F + L
Sbjct: 116 HQEWLAEVNYLGQFSHRHLVKLIGYCLEDEHRLLVYEFMPRGSLENHLF---RRGLYFQP 172
Query: 636 LDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGK 695
L W++R + LG A+GL +LH S RVI+RD KTSNILLD E K+SDFGLA+
Sbjct: 173 LSWKLRLKVALGAAKGLAFLH-SSETRVIYRDFKTSNILLDSEYNAKLSDFGLAKDGPIG 231
Query: 696 ETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSL 755
+ + RV+GT+GY +PEY G +TKSD++SFGVVLLE++SG++ + G +L
Sbjct: 232 DKSHVSTRVMGTHGYAAPEYLATGHLTTKSDVYSFGVVLLELLSGRRAVDKNRPSGERNL 291
Query: 756 LGYAWK-LWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLD 814
+ +A L + K+ ++D L + Y+ + + + L C+ E RPNMS VV L+
Sbjct: 292 VEWAKPYLVNKRKIFRVIDNRLQDQYSMEEACKVATLSLRCLTTEIKLRPNMSEVVSHLE 351
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 190 bits (482), Expect = 2e-47
Identities = 114/297 (38%), Positives = 170/297 (56%), Gaps = 13/297 (4%)
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKG----------KLQGGREIAVKRLSSVSSQG 579
F F + AT F + LG GG+G V+KG K G IAVK+L+ QG
Sbjct: 57 FTFAELKAATRNFRPDSVLGEGGFGSVFKGWIDEQTLTASKPGTGVVIAVKKLNQDGWQG 116
Query: 580 IQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDP-TKSALLDW 638
QE+ EV + + H NLV+L GYC++ + ++L+YE+MP SL+ +F + L W
Sbjct: 117 HQEWLAEVNYLGQFSHPNLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGSYFQPLSW 176
Query: 639 QMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETE 698
+R + LG A+GL +LH ++ VI+RD KTSNILLD E K+SDFGLA+ +
Sbjct: 177 TLRLKVALGAAKGLAFLH-NAETSVIYRDFKTSNILLDSEYNAKLSDFGLAKDGPTGDKS 235
Query: 699 ANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGY 758
+ R++GTYGY +PEY G +TKSD++S+GVVLLE++SG++ + G L+ +
Sbjct: 236 HVSTRIMGTYGYAAPEYLATGHLTTKSDVYSYGVVLLEVLSGRRAVDKNRPPGEQKLVEW 295
Query: 759 AWKLW-TENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLD 814
A L + KL ++D L + Y+ + + + L C+ E RPNM+ VV L+
Sbjct: 296 ARPLLANKRKLFRVIDNRLQDQYSMEEACKVATLALRCLTFEIKLRPNMNEVVSHLE 352
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 187 bits (476), Expect = 8e-47
Identities = 126/389 (32%), Positives = 190/389 (48%), Gaps = 46/389 (11%)
Query: 446 TRKGNPKSTLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSE 505
TR N L+ L G+V + I + CR+ ALK + S R + + SE
Sbjct: 617 TRSKNIGYVWILLTIFLLAGLVFVVGIVMFIAKCRKLRALKSSTLAASKWRSFHKLHFSE 676
Query: 506 RHVKDLIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGR 565
+ D +D++ N +G G G VYK +L+GG
Sbjct: 677 HEIADCLDEK------------------------------NVIGFGSSGKVYKVELRGGE 706
Query: 566 EIAVKRLSSVSSQGIQEFKN----------EVVLIAKLQHRNLVRLWGYCIKGDEKILIY 615
+AVK+L+ G E+ + EV + ++H+++VRLW C GD K+L+Y
Sbjct: 707 VVAVKKLNKSVKGGDDEYSSDSLNRDVFAAEVETLGTIRHKSIVRLWCCCSSGDCKLLVY 766
Query: 616 EYMPNKSL-DAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNIL 674
EYMPN SL D D +L W R I L A GL YLH D ++HRD+K+SNIL
Sbjct: 767 EYMPNGSLADVLHGDRKGGVVLGWPERLRIALDAAEGLSYLHHDCVPPIVHRDVKSSNIL 826
Query: 675 LDGEMQPKISDFGLARI--FGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGV 732
LD + K++DFG+A++ G +T + G+ GY++PEY + + KSDI+SFGV
Sbjct: 827 LDSDYGAKVADFGIAKVGQMSGSKTPEAMSGIAGSCGYIAPEYVYTLRVNEKSDIYSFGV 886
Query: 733 VLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVG 792
VLLE+++GK+ T G + + + L ++D L + + + H+G
Sbjct: 887 VLLELVTGKQPTD--SELGDKDMAKWVCTALDKCGLEPVIDPKLDLKFK-EEISKVIHIG 943
Query: 793 LLCVQDEPDDRPNMSNVVIMLDSETATLP 821
LLC P +RP+M VVIML + +P
Sbjct: 944 LLCTSPLPLNRPSMRKVVIMLQEVSGAVP 972
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 181 bits (458), Expect = 9e-45
Identities = 101/285 (35%), Positives = 165/285 (57%), Gaps = 5/285 (1%)
Query: 529 YFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVV 588
YF + ++ T+ F +G+GG+G VY G + G ++AVK LS S+QG +EF+ EV
Sbjct: 563 YFKYSEVVNITNNFERV--IGKGGFGKVYHGVING-EQVAVKVLSEESAQGYKEFRAEVD 619
Query: 589 LIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGI 648
L+ ++ H NL L GYC + + +LIYEYM N++L ++ +S +L W+ R I L
Sbjct: 620 LLMRVHHTNLTSLVGYCNEINHMVLIYEYMANENLGDYLAGK-RSFILSWEERLKISLDA 678
Query: 649 ARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTY 708
A+GL YLH + ++HRD+K +NILL+ ++Q K++DFGL+R F + + + V G+
Sbjct: 679 AQGLEYLHNGCKPPIVHRDVKPTNILLNEKLQAKMADFGLSRSFSVEGSGQISTVVAGSI 738
Query: 709 GYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKL 768
GY+ PEY Q + KSD++S GVVLLE+I+G+ + + + + + + +
Sbjct: 739 GYLDPEYYSTRQMNEKSDVYSLGVVLLEVITGQPAIASSKTE-KVHISDHVRSILANGDI 797
Query: 769 LDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIML 813
++D L E Y+ + + + L C + RP MS VV+ L
Sbjct: 798 RGIVDQRLRERYDVGSAWKMSEIALACTEHTSAQRPTMSQVVMEL 842
>KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1
precursor (EC 2.7.1.37)
Length = 817
Score = 179 bits (454), Expect = 3e-44
Identities = 100/289 (34%), Positives = 162/289 (55%), Gaps = 11/289 (3%)
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVL 589
+ + ++ AT F +LGRG G VYKG L+ R +AVK+L +V QG + F+ E+ +
Sbjct: 524 YSYRELVKATRKFKV--ELGRGESGTVYKGVLEDDRHVAVKKLENVR-QGKEVFQAELSV 580
Query: 590 IAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIA 649
I ++ H NLVR+WG+C +G ++L+ EY+ N SL +F + LLDW+ RF+I LG+A
Sbjct: 581 IGRINHMNLVRIWGFCSEGSHRLLVSEYVENGSLANILFSEGGNILLDWEGRFNIALGVA 640
Query: 650 RGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYG 709
+GL YLH + VIH D+K NILLD +PKI+DFGL ++ + N V GT G
Sbjct: 641 KGLAYLHHECLEWVIHCDVKPENILLDQAFEPKITDFGLVKLLNRGGSTQNVSHVRGTLG 700
Query: 710 YMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFY--------QYKGTLSLLGYAWK 761
Y++PE+ + K D++S+GVVLLE+++G + + + + +L +
Sbjct: 701 YIAPEWVSSLPITAKVDVYSYGVVLLELLTGTRVSELVGGTDEVHSMLRKLVRMLSAKLE 760
Query: 762 LWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVV 810
++ + +D L N Q + + C++++ RP M + V
Sbjct: 761 GEEQSWIDGYLDSKLNRPVNYVQARTLIKLAVSCLEEDRSKRPTMEHAV 809
Score = 84.3 bits (207), Expect = 1e-15
Identities = 74/266 (27%), Positives = 110/266 (40%), Gaps = 19/266 (7%)
Query: 75 IWYYREEGSGLS--PVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTN 132
+WY + E + + +VW AN D PV + DGN+V+ D G W A N
Sbjct: 71 VWYSKTEAAAANNKTIVWSANPDRPVHARR-SALTLQKDGNMVLTDYDGAAVWRADG--N 127
Query: 133 SSSATNRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMDKTLELTCWKSLSDP 192
+ + R+ +L+D+GNLV+ D G +W+SF+ PTDTFLP + L P
Sbjct: 128 NFTGVQRA-RLLDTGNLVIEDSG-GNTVWQSFDSPTDTFLPTQLITAATRLVPTTQSRSP 185
Query: 193 GRGNFTFKMDKKWENRFAILNQGQLYWQSEEQG---DGVMNPESNPDDISNDVYNL---- 245
G F F + + +YW +Q DG S + D L
Sbjct: 186 GNYIFRFSDLSVLSLIYHVPQVSDIYWPDPDQNLYQDGRNQYNSTRLGMLTDSGVLASSD 245
Query: 246 LTNFKELKNKTVSSYDNTRLLLNSTGVIKVLYRVNFQSDIVW---WYQPRTTCLTYNVCG 302
+ + L V RL L+ G ++ LY +N SD W C + +CG
Sbjct: 246 FADGQALVASDVGPGVKRRLTLDPDGNLR-LYSMN-DSDGSWSVSMVAMTQPCNIHGLCG 303
Query: 303 NFSSCNDDNDKLCTCLPGFGRRSPLN 328
C+ C+C PG+ R+P N
Sbjct: 304 PNGICHYSPTPTCSCPPGYATRNPGN 329
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 179 bits (453), Expect = 4e-44
Identities = 111/293 (37%), Positives = 165/293 (55%), Gaps = 13/293 (4%)
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKG----------KLQGGREIAVKRLSSVSSQG 579
F + AT F + +G GG+G V+KG K G IAVKRL+ QG
Sbjct: 56 FSLSELKSATRNFRPDSVVGEGGFGCVFKGWIDESSLAPSKPGTGIVIAVKRLNQEGFQG 115
Query: 580 IQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDP-TKSALLDW 638
+E+ E+ + +L H NLV+L GYC++ + ++L+YE+M SL+ +F T L W
Sbjct: 116 HREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLVYEFMTRGSLENHLFRRGTFYQPLSW 175
Query: 639 QMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETE 698
R + LG ARGL +LH +++ +VI+RD K SNILLD K+SDFGLAR +
Sbjct: 176 NTRVRMALGAARGLAFLH-NAQPQVIYRDFKASNILLDSNYNAKLSDFGLARDGPMGDNS 234
Query: 699 ANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGY 758
+ RV+GT GY +PEY G S KSD++SFGVVLLE++SG++ Q G +L+ +
Sbjct: 235 HVSTRVMGTQGYAAPEYLATGHLSVKSDVYSFGVVLLELLSGRRAIDKNQPVGEHNLVDW 294
Query: 759 AWK-LWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVV 810
A L + +LL +MD L Y+ + ++ + L C+ + RP M+ +V
Sbjct: 295 ARPYLTNKRRLLRVMDPRLQGQYSLTRALKIAVLALDCISIDAKSRPTMNEIV 347
>CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4
precursor (EC 2.7.1.-)
Length = 901
Score = 176 bits (446), Expect = 2e-43
Identities = 110/315 (34%), Positives = 171/315 (53%), Gaps = 9/315 (2%)
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSS--QGIQEFKNEV 587
F +E + AT FS+ +++G+G + V+KG L+ G +AVKR S + +EF NE+
Sbjct: 493 FSYEELEQATGGFSEDSQVGKGSFSCVFKGILRDGTVVAVKRAIKASDVKKSSKEFHNEL 552
Query: 588 VLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVF--DPTKSALLDWQMRFDIL 645
L+++L H +L+ L GYC G E++L+YE+M + SL + DP L+W R I
Sbjct: 553 DLLSRLNHAHLLNLLGYCEDGSERLLVYEFMAHGSLYQHLHGKDPNLKKRLNWARRVTIA 612
Query: 646 LGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVV 705
+ ARG+ YLH + VIHRD+K+SNIL+D + +++DFGL+ + ++
Sbjct: 613 VQAARGIEYLHGYACPPVIHRDIKSSNILIDEDHNARVADFGLSILGPADSGTPLSELPA 672
Query: 706 GTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTE 765
GT GY+ PEY +TKSD++SFGVVLLEI+SG+K +G +++ +A L
Sbjct: 673 GTLGYLDPEYYRLHYLTTKSDVYSFGVVLLEILSGRKAIDMQFEEG--NIVEWAVPLIKA 730
Query: 766 NKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATL---PT 822
+ ++D L + + V CV+ DRP+M V L+ A L P
Sbjct: 731 GDIFAILDPVLSPPSDLEALKKIASVACKCVRMRGKDRPSMDKVTTALEHALALLMGSPC 790
Query: 823 PKQPTFFTRKDLSST 837
+QP T L S+
Sbjct: 791 IEQPILPTEVVLGSS 805
>RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC
2.7.1.37)
Length = 1109
Score = 175 bits (443), Expect = 5e-43
Identities = 104/314 (33%), Positives = 167/314 (53%), Gaps = 14/314 (4%)
Query: 507 HVKDLIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGRE 566
H K + + + ++ +G + +L AT+ +D +G+G +G +YK L +
Sbjct: 786 HCKKSVQEIAISAQEGDGSLL-----NKVLEATENLNDKYVIGKGAHGTIYKATLSPDKV 840
Query: 567 IAVKRLSSVS-SQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDA 625
AVK+L G E+ I K++HRNL++L + ++ + +++Y YM N SL
Sbjct: 841 YAVKKLVFTGIKNGSVSMVREIETIGKVRHRNLIKLEEFWLRKEYGLILYTYMENGSLHD 900
Query: 626 FVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISD 685
+ + LDW R +I +G A GL YLH D ++HRD+K NILLD +++P ISD
Sbjct: 901 ILHETNPPKPLDWSTRHNIAVGTAHGLAYLHFDCDPAIVHRDIKPMNILLDSDLEPHISD 960
Query: 686 FGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTG 745
FG+A++ T + V GT GYM+PE A S +SD++S+GVVLLE+I+ KK
Sbjct: 961 FGIAKLLDQSATSIPSNTVQGTIGYMAPENAFTTVKSRESDVYSYGVVLLELITRKKALD 1020
Query: 746 FYQYKGTLSLLGYAWKLWTE----NKLLD--LMDLSLGEAYNANQFIRCTHVGLLCVQDE 799
+ G ++G+ +WT+ K++D L+D L ++ Q + L C + E
Sbjct: 1021 -PSFNGETDIVGWVRSVWTQTGEIQKIVDPSLLD-ELIDSSVMEQVTEALSLALRCAEKE 1078
Query: 800 PDDRPNMSNVVIML 813
D RP M +VV L
Sbjct: 1079 VDKRPTMRDVVKQL 1092
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 106,150,424
Number of Sequences: 164201
Number of extensions: 4851487
Number of successful extensions: 14556
Number of sequences better than 10.0: 1638
Number of HSP's better than 10.0 without gapping: 1080
Number of HSP's successfully gapped in prelim test: 558
Number of HSP's that attempted gapping in prelim test: 10733
Number of HSP's gapped (non-prelim): 2072
length of query: 852
length of database: 59,974,054
effective HSP length: 119
effective length of query: 733
effective length of database: 40,434,135
effective search space: 29638220955
effective search space used: 29638220955
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0203.5


