

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.3
(475 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
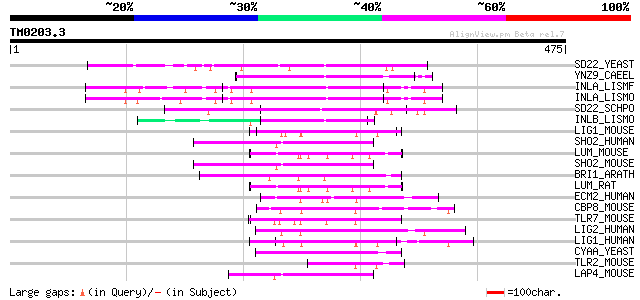
Score E
Sequences producing significant alignments: (bits) Value
SD22_YEAST (P36047) Protein phosphatases PP1 regulatory subunit ... 70 2e-11
YNZ9_CAEEL (P45969) Hypothetical protein T09A5.9 in chromosome III 63 2e-09
INLA_LISMF (Q723K6) Internalin A precursor 63 2e-09
INLA_LISMO (P25146) Internalin A precursor 62 3e-09
SD22_SCHPO (P22194) Protein phosphatases PP1 regulatory subunit ... 62 4e-09
INLB_LISMO (P25147) Internalin B precursor 55 3e-07
LIG1_MOUSE (P70193) Leucine-rich repeats and immunoglobulin-like... 55 4e-07
SHO2_HUMAN (Q9UQ13) Leucine-rich repeat protein SHOC-2 (Ras-bind... 55 5e-07
LUM_MOUSE (P51885) Lumican precursor (Keratan sulfate proteoglyc... 55 5e-07
SHO2_MOUSE (O88520) Leucine-rich repeat protein SHOC-2 (Ras-bind... 54 7e-07
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 53 1e-06
LUM_RAT (P51886) Lumican precursor (Keratan sulfate proteoglycan... 53 2e-06
ECM2_HUMAN (O94769) Extracellular matrix protein 2 precursor (Ma... 53 2e-06
CBP8_MOUSE (Q9DBB9) Carboxypeptidase N 83 kDa chain precursor (C... 52 3e-06
TLR7_MOUSE (P58681) Toll-like receptor 7 precursor 52 3e-06
LIG2_HUMAN (O94898) Leucine-rich repeats and immunoglobulin-like... 52 3e-06
LIG1_HUMAN (Q96JA1) Leucine-rich repeats and immunoglobulin-like... 51 7e-06
CYAA_YEAST (P08678) Adenylate cyclase (EC 4.6.1.1) (ATP pyrophos... 51 7e-06
TLR2_MOUSE (Q9QUN7) Toll-like receptor 2 precursor 50 1e-05
LAP4_MOUSE (Q80U72) LAP4 protein (Scribble homolog protein) 50 1e-05
>SD22_YEAST (P36047) Protein phosphatases PP1 regulatory subunit
SDS22
Length = 338
Score = 69.7 bits (169), Expect = 2e-11
Identities = 82/328 (25%), Positives = 147/328 (44%), Gaps = 50/328 (15%)
Query: 67 EDNHMGEYFDDGFDSYLLSDLEKDWVMPTTDDISEVKTLQGDNSVDCFGEFPNKDFKVKR 126
++ H E DD ++ +D E +P ++ ++ L+ S++ + K+ K
Sbjct: 14 DERHKIEVVDDTNPDFITADSELTQDLPDDVEVIDLVHLK-IKSLEDLNLYRFKNLKQLC 72
Query: 127 IEDWVVDLQHCGPPVEEINELPESVDPVVDI----NTINGVTAAGVNH--KITPGMEAAK 180
+ +++ + E+ LP D +VD+ N I +++ VN K+T ++ +
Sbjct: 73 LRQNLIE------SISEVEVLPH--DKIVDLDFYDNKIKHISS-NVNKLTKLT-SLDLSF 122
Query: 181 RYISSLTANASAAQLAN-----HGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLH 235
I + + L N + + + LS SLK L L GN + I + GL
Sbjct: 123 NKIKHIKNLENLTDLENLYFVQNSISKIENLSTLKSLKNLELGGNKVHSIEPDSF-EGLS 181
Query: 236 SLN---LSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV 292
+L L +N+I + L L L++L + N++ +I L ++L+ELYL+ N I ++
Sbjct: 182 NLEEIWLGKNSIPRLINLHPLKNLKILSIQSNKLKKI-ENLEELTNLEELYLSHNFITKI 240
Query: 293 EGLHRLLKLSILDLRFNKISTAKCLGQLA------ANYNS-----------------LQA 329
EGL + LKL+ LD+ NKI++ + L L+ A++N L+
Sbjct: 241 EGLEKNLKLTTLDVTSNKITSLENLNHLSNLTDIWASFNKIDQSFESLGENLSALSRLET 300
Query: 330 INLDGNPCQKNVGDEQLKKYLQGLLPHL 357
I L+GNP Q +K L P L
Sbjct: 301 IYLEGNPIQLENKTSYRRKLTMNLPPSL 328
>YNZ9_CAEEL (P45969) Hypothetical protein T09A5.9 in chromosome III
Length = 326
Score = 62.8 bits (151), Expect = 2e-09
Identities = 53/168 (31%), Positives = 82/168 (48%), Gaps = 8/168 (4%)
Query: 195 LANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELT 254
L ++ + + L A LK+L L N I +I L L + +N I +EG+ L
Sbjct: 132 LVSNKIEKIENLEALTQLKLLELGDNRIKKIENIGHLVNLDELFIGKNKIRQLEGVETLQ 191
Query: 255 RLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTA 314
+L VL L NRI++I + ++LKELYL+ + ++ G+ L L +LD+ N+I T
Sbjct: 192 KLSVLSLPGNRIVKI-ENVEQLNNLKELYLSDQGLQDIHGVEPLTNLLLLDVANNEIKTF 250
Query: 315 KCLGQLAANYNSLQAINLDGNPCQKNVGDEQLKKYLQGLLPHLVYYNR 362
+ +L SL + N + EQL K L+GL VY R
Sbjct: 251 SGVERL----ESLNDFWANDNKVESFSEIEQLSK-LKGL--QTVYLER 291
Score = 59.7 bits (143), Expect = 2e-08
Identities = 52/154 (33%), Positives = 79/154 (50%), Gaps = 6/154 (3%)
Query: 194 QLANHGLVVV-PFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRE 252
++ N+ LV + P +S+ V+L L+L N + I+ L SL+LS N I I GL +
Sbjct: 64 RMRNNLLVSISPTISSLVTLTSLDLYENQLTEISHLESLVNLVSLDLSYNRIRQINGLDK 123
Query: 253 LTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKIS 312
LT+L L L N+I +I L + + LK L L N+I ++E + L+ L L + NKI
Sbjct: 124 LTKLETLYLVSNKIEKI-ENLEALTQLKLLELGDNRIKKIENIGHLVNLDELFIGKNKIR 182
Query: 313 TAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQL 346
+ + L L ++L GN K EQL
Sbjct: 183 QLEGVETL----QKLSVLSLPGNRIVKIENVEQL 212
Score = 36.2 bits (82), Expect = 0.19
Identities = 30/125 (24%), Positives = 54/125 (43%), Gaps = 17/125 (13%)
Query: 237 LNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLH 296
++L+ I L ++ L + N ++ I ++S +L L L N++ E+ L
Sbjct: 41 IDLTHTRADHIPDLTGFPKIEELRMRNNLLVSISPTISSLVTLTSLDLYENQLTEISHLE 100
Query: 297 RLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQK-------------NVGD 343
L+ L LDL +N+I L +L L+ + L N +K +GD
Sbjct: 101 SLVNLVSLDLSYNRIRQINGLDKL----TKLETLYLVSNKIEKIENLEALTQLKLLELGD 156
Query: 344 EQLKK 348
++KK
Sbjct: 157 NRIKK 161
>INLA_LISMF (Q723K6) Internalin A precursor
Length = 800
Score = 62.8 bits (151), Expect = 2e-09
Identities = 93/353 (26%), Positives = 155/353 (43%), Gaps = 60/353 (16%)
Query: 66 TEDNHMGEYFDD-GFDSYLLSDLEKDWVMPTTD--DISEVKTLQGDN----SVDCFGEFP 118
T+D + + F D + + L K V T D+ +V TLQ D S+D E+
Sbjct: 39 TQDTPINQIFTDTALAEKMKTVLGKTNVTDTVSQTDLDQVTTLQADRLGIKSIDGL-EYL 97
Query: 119 NKDFKVKRIEDWVVDLQHCGPPVEEINELPE---------SVDPVVDINTINGVTAAGVN 169
N ++ + + D+ P++++ +L + + P+ +++ + G+T N
Sbjct: 98 NNLTQINFSNNQLTDIT----PLKDLTKLVDILMNNNQIADITPLANLSNLTGLTL--FN 151
Query: 170 HKITP-----------GMEAAKRYISSLTA--NASAAQLANHGLVVVPF--LSAFVSLKV 214
++IT +E + IS ++A ++ Q + G V L+ +L+
Sbjct: 152 NQITDIDPLKNLTNLNRLELSSNTISDISALSGLTSLQQLSFGNQVTDLKPLANLTTLER 211
Query: 215 LNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLA 274
L+++ N + I+ A L SL + N IS I L LT L L L+ N++ IG LA
Sbjct: 212 LDISSNKVSDISVLAKLTNLESLIATNNQISDITPLGILTNLDELSLNGNQLKDIGT-LA 270
Query: 275 SCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLAA------NYNSLQ 328
S ++L +L LA N+I + L L KL+ L L N+IS L L A N N L+
Sbjct: 271 SLTNLTDLDLANNQISNLAPLSGLTKLTELKLGANQISNISPLAGLTALTNLELNENQLE 330
Query: 329 AINLDGNPCQKNVGDEQLKKYLQGL-----------LPHLVYYNRQAMKVSTL 370
I+ N KN+ L Y + L L +YN + VS+L
Sbjct: 331 DISPISN--LKNL--TYLTLYFNNISDISPVSSLTKLQRLFFYNNKVSDVSSL 379
Score = 52.4 bits (124), Expect = 3e-06
Identities = 42/138 (30%), Positives = 64/138 (45%), Gaps = 2/138 (1%)
Query: 183 ISSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRN 242
++SLT N + LAN+ + + LS L L L N I I+ A L +L L+ N
Sbjct: 269 LASLT-NLTDLDLANNQISNLAPLSGLTKLTELKLGANQISNISPLAGLTALTNLELNEN 327
Query: 243 NISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLS 302
+ I + L L L L +N I I ++S + L+ L+ NK+ +V L L ++
Sbjct: 328 QLEDISPISNLKNLTYLTLYFNNISDISP-VSSLTKLQRLFFYNNKVSDVSSLANLTNIN 386
Query: 303 ILDLRFNKISTAKCLGQL 320
L N+IS L L
Sbjct: 387 WLSAGHNQISDLTPLANL 404
>INLA_LISMO (P25146) Internalin A precursor
Length = 800
Score = 62.0 bits (149), Expect = 3e-09
Identities = 92/349 (26%), Positives = 150/349 (42%), Gaps = 52/349 (14%)
Query: 66 TEDNHMGEYFDD-GFDSYLLSDLEKDWVMPTTD--DISEVKTLQGD-------NSVDCFG 115
T+D + + F D + + L K V T D+ +V TLQ D + V+
Sbjct: 39 TQDTPINQIFTDTALAEKMKTVLGKTNVTDTVSQTDLDQVTTLQADRLGIKSIDGVEYLN 98
Query: 116 EFPNKDFKVKRIEDWVVDLQHCGPPVEEI--NELPESVDPVVDINTINGVTAAGVNHKIT 173
+F ++ D + L++ V+ + N + P+ ++ + G+T N++IT
Sbjct: 99 NLTQINFSNNQLTD-ITPLKNLTKLVDILMNNNQIADITPLANLTNLTGLTL--FNNQIT 155
Query: 174 P-----------GMEAAKRYISSLTA--NASAAQLANHGLVVVPF--LSAFVSLKVLNLA 218
+E + IS ++A ++ Q + G V L+ +L+ L+++
Sbjct: 156 DIDPLKNLTNLNRLELSSNTISDISALSGLTSLQQLSFGNQVTDLKPLANLTTLERLDIS 215
Query: 219 GNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSS 278
N + I+ A L SL + N IS I L LT L L L+ N++ IG LAS ++
Sbjct: 216 SNKVSDISVLAKLTNLESLIATNNQISDITPLGILTNLDELSLNGNQLKDIGT-LASLTN 274
Query: 279 LKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLAA------NYNSLQAINL 332
L +L LA N+I + L L KL+ L L N+IS L L A N N L+ I+
Sbjct: 275 LTDLDLANNQISNLAPLSGLTKLTELKLGANQISNISPLAGLTALTNLELNENQLEDISP 334
Query: 333 DGNPCQKNVGDEQLKKYLQGL-----------LPHLVYYNRQAMKVSTL 370
N KN+ L Y + L L +YN + VS+L
Sbjct: 335 ISN--LKNL--TYLTLYFNNISDISPVSSLTKLQRLFFYNNKVSDVSSL 379
Score = 52.4 bits (124), Expect = 3e-06
Identities = 42/138 (30%), Positives = 64/138 (45%), Gaps = 2/138 (1%)
Query: 183 ISSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRN 242
++SLT N + LAN+ + + LS L L L N I I+ A L +L L+ N
Sbjct: 269 LASLT-NLTDLDLANNQISNLAPLSGLTKLTELKLGANQISNISPLAGLTALTNLELNEN 327
Query: 243 NISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLS 302
+ I + L L L L +N I I ++S + L+ L+ NK+ +V L L ++
Sbjct: 328 QLEDISPISNLKNLTYLTLYFNNISDISP-VSSLTKLQRLFFYNNKVSDVSSLANLTNIN 386
Query: 303 ILDLRFNKISTAKCLGQL 320
L N+IS L L
Sbjct: 387 WLSAGHNQISDLTPLANL 404
>SD22_SCHPO (P22194) Protein phosphatases PP1 regulatory subunit
sds22
Length = 332
Score = 61.6 bits (148), Expect = 4e-09
Identities = 71/269 (26%), Positives = 126/269 (46%), Gaps = 20/269 (7%)
Query: 133 DLQHCGPPVEEINELPESVDPVVDINT-INGVTAAGV----NHKITPGMEAAKRYISSLT 187
D+Q + ++++P+ VD V I + I + + G+ N + + + I S+
Sbjct: 22 DVQQIDADEDLLDDVPDDVDCVELIQSRIQSMASLGLERFKNLQSLCLRQNQIKKIESVP 81
Query: 188 ANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSI 247
+ L ++ +V + L +L L+L+ N+I I +GL +L +N I I
Sbjct: 82 ETLTELDLYDNLIVRIENLDNVKNLTYLDLSFNNIKTIRNINHLKGLENLFFVQNRIRRI 141
Query: 248 EGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLR 307
E L L RL L+L N+I R+ L + +L++L++ NKI + E +L KLS+L ++
Sbjct: 142 ENLEGLDRLTNLELGGNKI-RVIENLDTLVNLEKLWVGKNKITKFENFEKLQKLSLLSIQ 200
Query: 308 FNKIS-------TAKCLGQLAANYN---SLQAINLDGNPCQKNVGDEQLK--KYLQGL-- 353
N+I+ + CL +L ++N S I + N +V + +K YL GL
Sbjct: 201 SNRITQFENLACLSHCLRELYVSHNGLTSFSGIEVLENLEILDVSNNMIKHLSYLAGLKN 260
Query: 354 LPHLVYYNRQAMKVSTLKDGADRLVRLGT 382
L L N + ++D L +L T
Sbjct: 261 LVELWASNNELSSFQEIEDELSGLKKLET 289
Score = 46.2 bits (108), Expect = 2e-04
Identities = 41/147 (27%), Positives = 62/147 (41%), Gaps = 22/147 (14%)
Query: 215 LNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLA 274
L L GN I I L L + +N I+ E +L +L +L + NRI + +
Sbjct: 153 LELGGNKIRVIENLDTLVNLEKLWVGKNKITKFENFEKLQKLSLLSIQSNRITQFENLAC 212
Query: 275 SCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKI------STAKCLGQLAANYN--- 325
L+ELY++ N + G+ L L ILD+ N I + K L +L A+ N
Sbjct: 213 LSHCLRELYVSHNGLTSFSGIEVLENLEILDVSNNMIKHLSYLAGLKNLVELWASNNELS 272
Query: 326 -------------SLQAINLDGNPCQK 339
L+ + +GNP QK
Sbjct: 273 SFQEIEDELSGLKKLETVYFEGNPLQK 299
>INLB_LISMO (P25147) Internalin B precursor
Length = 630
Score = 55.5 bits (132), Expect = 3e-07
Identities = 37/98 (37%), Positives = 51/98 (51%), Gaps = 1/98 (1%)
Query: 215 LNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLA 274
L L GN + I A + L L L N + + L++L +L+ L L +N I I +GL
Sbjct: 103 LFLNGNKLTDIKPLANLKNLGWLFLDENKVKDLSSLKDLKKLKSLSLEHNGISDI-NGLV 161
Query: 275 SCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKIS 312
L+ LYL NKI ++ L RL KL L L N+IS
Sbjct: 162 HLPQLESLYLGNNKITDITVLSRLTKLDTLSLEDNQIS 199
Score = 50.1 bits (118), Expect = 1e-05
Identities = 57/205 (27%), Positives = 82/205 (39%), Gaps = 24/205 (11%)
Query: 110 SVDCFGEFPNKDFKVKRIEDWVVDLQHCGPPVEEINELPESVDPVVDINTINGVTAAGVN 169
S D F E + K K + D V E+N + + + DI ++ G+
Sbjct: 49 SDDAFAETIKDNLKKKSVTDAVTQ--------NELNSIDQIIANNSDIKSVQGI------ 94
Query: 170 HKITPGMEAAKRYISSLTANASAAQLANHGLVVVP--------FLSAFVSLKVLNLAGNS 221
+ P + + LT A L N G + + L LK L+L N
Sbjct: 95 -QYLPNVTKLFLNGNKLTDIKPLANLKNLGWLFLDENKVKDLSSLKDLKKLKSLSLEHNG 153
Query: 222 IVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKE 281
I I L SL L N I+ I L LT+L L L N+I I LA + L+
Sbjct: 154 ISDINGLVHLPQLESLYLGNNKITDITVLSRLTKLDTLSLEDNQISDI-VPLAGLTKLQN 212
Query: 282 LYLAGNKIGEVEGLHRLLKLSILDL 306
LYL+ N I ++ L L L +L+L
Sbjct: 213 LYLSKNHISDLRALAGLKNLDVLEL 237
Score = 31.2 bits (69), Expect = 6.1
Identities = 24/100 (24%), Positives = 47/100 (47%), Gaps = 7/100 (7%)
Query: 238 NLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHR 297
NL + +++ EL + + ++ N ++ G+ ++ +L+L GNK+ +++ L
Sbjct: 60 NLKKKSVTDAVTQNELNSIDQI-IANNSDIKSVQGIQYLPNVTKLFLNGNKLTDIKPLAN 118
Query: 298 LLKLSILDLRFNKI------STAKCLGQLAANYNSLQAIN 331
L L L L NK+ K L L+ +N + IN
Sbjct: 119 LKNLGWLFLDENKVKDLSSLKDLKKLKSLSLEHNGISDIN 158
>LIG1_MOUSE (P70193) Leucine-rich repeats and immunoglobulin-like
domains protein 1 precursor (LIG-1)
Length = 1091
Score = 55.1 bits (131), Expect = 4e-07
Identities = 49/132 (37%), Positives = 71/132 (53%), Gaps = 8/132 (6%)
Query: 212 LKVLNLAGNSIVRITAGALP--RGLHSLNLSRNNISSI--EGLRELTRLRVLDLSYNRIL 267
+ VL+L NS+V + +G+L LH L+LS N+IS I +G +L L LS+N +
Sbjct: 263 MHVLHLEYNSLVEVNSGSLYGLTALHQLHLSNNSISRIQRDGWSFCQKLHELILSFNNLT 322
Query: 268 RIG-HGLASCSSLKELYLAGNKIGEV-EGLHRLLK-LSILDLRFNKIS-TAKCLGQLAAN 323
R+ LA SSL L L+ N I + EG + LK L +LDL N+IS T +
Sbjct: 323 RLDEESLAELSSLSILRLSHNAISHIAEGAFKGLKSLRVLDLDHNEISGTIEDTSGAFTG 382
Query: 324 YNSLQAINLDGN 335
++L + L GN
Sbjct: 383 LDNLSKLTLFGN 394
Score = 51.6 bits (122), Expect = 4e-06
Identities = 49/141 (34%), Positives = 69/141 (48%), Gaps = 15/141 (10%)
Query: 206 LSAFVSLKVLNLAGNSIVRITAGALPRGL--HSLNLSRNNISSIE--GLRELTR-LRVLD 260
L +++SL+VL+L+ N+I I + P GL LNL+ N IS +E L+R L L
Sbjct: 137 LKSYLSLEVLDLSSNNITEIRSSCFPNGLRIRELNLASNRISILESGAFDGLSRSLLTLR 196
Query: 261 LSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGL--HRLLKLSILDLRFNKIST----- 313
LS NRI ++ L +L L N+I +EGL L L +L L+ N IS
Sbjct: 197 LSKNRITQLPVKAFKLPRLTQLDLNRNRIRLIEGLTFQGLDSLEVLRLQRNNISRLTDGA 256
Query: 314 ---AKCLGQLAANYNSLQAIN 331
+ L YNSL +N
Sbjct: 257 FWGLSKMHVLHLEYNSLVEVN 277
Score = 41.6 bits (96), Expect = 0.005
Identities = 38/105 (36%), Positives = 52/105 (49%), Gaps = 10/105 (9%)
Query: 218 AGNSIVRITAG--ALPRGL----HSLNLSRNNISSIEG--LRELTRLRVLDLSYNRILRI 269
AGNS+ G LPR L SLNLS N +S I+ +LT L+ + L+ N + I
Sbjct: 50 AGNSLDCSGRGLATLPRDLPSWTRSLNLSYNRLSEIDSAAFEDLTNLQEVYLNSNELTAI 109
Query: 270 GHGLASCSSLKELYLAGNKIGEVEG--LHRLLKLSILDLRFNKIS 312
+ + L+L NKI V+G L L L +LDL N I+
Sbjct: 110 PSLGTASIGVVSLFLQHNKILSVDGSQLKSYLSLEVLDLSSNNIT 154
>SHO2_HUMAN (Q9UQ13) Leucine-rich repeat protein SHOC-2 (Ras-binding
protein Sur-8)
Length = 582
Score = 54.7 bits (130), Expect = 5e-07
Identities = 50/159 (31%), Positives = 75/159 (46%), Gaps = 6/159 (3%)
Query: 158 NTINGVTAAGVNHKITPGMEAAKRYISSLTANASAAQLANHGLVVVPF-LSAFVSLKVLN 216
NTI A K + E K N+ L+ + ++P + L L
Sbjct: 70 NTIKRPNPAPGTRKKSSNAEVIKELNKCREENSMRLDLSKRSIHILPSSIKELTQLTELY 129
Query: 217 LAGNSIVRITA--GALPRGLHSLNLSRNNISSI-EGLRELTRLRVLDLSYNRILRIGHGL 273
L N + + A G L L +L LS N+++S+ + L L +LR+LDL +N++ I +
Sbjct: 130 LYSNKLQSLPAEVGCLVN-LMTLALSENSLTSLPDSLDNLKKLRMLDLRHNKLREIPSVV 188
Query: 274 ASCSSLKELYLAGNKIGEVE-GLHRLLKLSILDLRFNKI 311
SL LYL N+I VE + L KLS+L +R NKI
Sbjct: 189 YRLDSLTTLYLRFNRITTVEKDIKNLSKLSMLSIRENKI 227
Score = 41.6 bits (96), Expect = 0.005
Identities = 32/107 (29%), Positives = 54/107 (49%), Gaps = 5/107 (4%)
Query: 236 SLNLSRNNISSI-EGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEG 294
+L+L N + + + + L+ L L L YNR+ I LA CS+L+EL L N I +
Sbjct: 265 NLDLQHNELLDLPDTIGNLSSLSRLGLRYNRLSAIPRSLAKCSALEELNLENNNISTLPE 324
Query: 295 --LHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQK 339
L L+KL+ L L N G + ++++ ++N++ N K
Sbjct: 325 SLLSSLVKLNSLTLARNCFQLYPVGG--PSQFSTIYSLNMEHNRINK 369
Score = 39.3 bits (90), Expect = 0.022
Identities = 38/117 (32%), Positives = 58/117 (49%), Gaps = 7/117 (5%)
Query: 203 VPFLSAFVSLKVLNLAGNSIVRITAGALPRG----LHSLNLSRNNISSI-EGLRELTRLR 257
+PF F KVL+ ++T+ L G + LNL+ N ++ I E + L L
Sbjct: 370 IPF-GIFSRAKVLSKLNMKDNQLTSLPLDFGTWTSMVELNLATNQLTKIPEDVSGLVSLE 428
Query: 258 VLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLK-LSILDLRFNKIST 313
VL LS N + ++ HGL + L+EL L NK+ + LK L L L N+++T
Sbjct: 429 VLILSNNLLKKLPHGLGNLRKLRELDLEENKLESLPNEIAYLKDLQKLVLTNNQLTT 485
Score = 33.9 bits (76), Expect = 0.94
Identities = 30/106 (28%), Positives = 55/106 (51%), Gaps = 5/106 (4%)
Query: 212 LKVLNLAGNSIVRITA--GALPRGLHSLNLSRNNISSI-EGLRELTRLRVLDLSYNRILR 268
L +L++ N I ++ A G L L +L+++ N + + + + T++ LDL +N +L
Sbjct: 217 LSMLSIRENKIKQLPAEIGELCN-LITLDVAHNQLEHLPKEIGNCTQITNLDLQHNELLD 275
Query: 269 IGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLSILDLRFNKIST 313
+ + + SSL L L N++ + L + L L+L N IST
Sbjct: 276 LPDTIGNLSSLSRLGLRYNRLSAIPRSLAKCSALEELNLENNNIST 321
>LUM_MOUSE (P51885) Lumican precursor (Keratan sulfate proteoglycan
lumican) (KSPG lumican)
Length = 338
Score = 54.7 bits (130), Expect = 5e-07
Identities = 47/139 (33%), Positives = 75/139 (53%), Gaps = 11/139 (7%)
Query: 206 LSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSI--EGLRELTRLRVLDLSY 263
L SL+ L+L+ N + ++ AG LP L +L L N IS+I E + T L+ L LS+
Sbjct: 180 LKGLKSLEYLDLSFNQMSKLPAG-LPTSLLTLYLDNNKISNIPDEYFKRFTGLQYLRLSH 238
Query: 264 NRILRIG--HGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDL----RFNKISTAKCL 317
N + G + SSL EL L+ NK+ + ++ L+ L++ +F+ S K L
Sbjct: 239 NELADSGVPGNSFNISSLLELDLSYNKLKSIPTVNENLENYYLEVNELEKFDVKSFCKIL 298
Query: 318 GQLAANYNSLQAINLDGNP 336
G L+ Y+ ++ + LDGNP
Sbjct: 299 GPLS--YSKIKHLRLDGNP 315
Score = 45.1 bits (105), Expect = 4e-04
Identities = 47/156 (30%), Positives = 70/156 (44%), Gaps = 31/156 (19%)
Query: 207 SAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNIS---SIEGLRELT--------- 254
S LK L++ N++ + G LP+ L L L+ N IS S +GL LT
Sbjct: 113 SKLKQLKKLHINYNNLTE-SVGPLPKSLQDLQLTNNKISKLGSFDGLVNLTFIYLQHNQL 171
Query: 255 -------------RLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV--EGLHRLL 299
L LDLS+N++ ++ GL +SL LYL NKI + E R
Sbjct: 172 KEDAVSASLKGLKSLEYLDLSFNQMSKLPAGLP--TSLLTLYLDNNKISNIPDEYFKRFT 229
Query: 300 KLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGN 335
L L L N+++ + G + N +SL ++L N
Sbjct: 230 GLQYLRLSHNELADSGVPGN-SFNISSLLELDLSYN 264
>SHO2_MOUSE (O88520) Leucine-rich repeat protein SHOC-2 (Ras-binding
protein Sur-8)
Length = 582
Score = 54.3 bits (129), Expect = 7e-07
Identities = 50/159 (31%), Positives = 75/159 (46%), Gaps = 6/159 (3%)
Query: 158 NTINGVTAAGVNHKITPGMEAAKRYISSLTANASAAQLANHGL-VVVPFLSAFVSLKVLN 216
NTI A K + E K N+ L+ + ++ P + L L
Sbjct: 70 NTIKRPNPAPGTRKKSSNAEVIKELNKCREENSMRLDLSKRSIHILPPSVKELTQLTELY 129
Query: 217 LAGNSIVRITA--GALPRGLHSLNLSRNNISSI-EGLRELTRLRVLDLSYNRILRIGHGL 273
L N + + A G L L +L LS N+++S+ + L L +LR+LDL +N++ I +
Sbjct: 130 LYSNKLQSLPAEVGCLVN-LMTLALSENSLTSLPDSLDNLKKLRMLDLRHNKLREIPSVV 188
Query: 274 ASCSSLKELYLAGNKIGEVE-GLHRLLKLSILDLRFNKI 311
SL LYL N+I VE + L KLS+L +R NKI
Sbjct: 189 YRLDSLTTLYLRFNRITTVEKDIKNLPKLSMLSIRENKI 227
Score = 42.4 bits (98), Expect = 0.003
Identities = 32/107 (29%), Positives = 55/107 (50%), Gaps = 5/107 (4%)
Query: 236 SLNLSRNNISSI-EGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEG 294
+L+L N++ + + + L+ L L L YNR+ I LA CS+L+EL L N I +
Sbjct: 265 NLDLQHNDLLDLPDTIGNLSSLNRLGLRYNRLSAIPRSLAKCSALEELNLENNNISTLPE 324
Query: 295 --LHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQK 339
L L+KL+ L L N G + ++++ ++N++ N K
Sbjct: 325 SLLSSLVKLNSLTLARNCFQLYPVGG--PSQFSTIYSLNMEHNRINK 369
Score = 40.4 bits (93), Expect = 0.010
Identities = 39/117 (33%), Positives = 58/117 (49%), Gaps = 7/117 (5%)
Query: 203 VPFLSAFVSLKVLNLAGNSIVRITAGALPRG----LHSLNLSRNNISSI-EGLRELTRLR 257
+PF F KVL+ ++T+ L G + LNL+ N ++ I E + L L
Sbjct: 370 IPF-GIFSRAKVLSKLNMKDNQLTSLPLDFGTWTSMVELNLATNQLTKIPEDVSGLVSLE 428
Query: 258 VLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLK-LSILDLRFNKIST 313
VL LS N + ++ HGL + L+EL L NK+ + LK L L L N++ST
Sbjct: 429 VLILSNNLLKKLPHGLGNLRKLRELDLEENKLESLPNEIAYLKDLQKLVLTNNQLST 485
Score = 33.1 bits (74), Expect = 1.6
Identities = 30/106 (28%), Positives = 55/106 (51%), Gaps = 5/106 (4%)
Query: 212 LKVLNLAGNSIVRITA--GALPRGLHSLNLSRNNISSI-EGLRELTRLRVLDLSYNRILR 268
L +L++ N I ++ A G L L +L+++ N + + + + T++ LDL +N +L
Sbjct: 217 LSMLSIRENKIKQLPAEIGELCN-LITLDVAHNQLEHLPKEIGNCTQITNLDLQHNDLLD 275
Query: 269 IGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLSILDLRFNKIST 313
+ + + SSL L L N++ + L + L L+L N IST
Sbjct: 276 LPDTIGNLSSLNRLGLRYNRLSAIPRSLAKCSALEELNLENNNIST 321
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 53.1 bits (126), Expect = 1e-06
Identities = 58/186 (31%), Positives = 84/186 (44%), Gaps = 17/186 (9%)
Query: 163 VTAAGVNHK-ITPGMEAAKRYISSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNS 221
VT+ ++ K + G A + SLT S +H V SL L+L+ NS
Sbjct: 74 VTSIDLSSKPLNVGFSAVSSSLLSLTGLESLFLSNSHINGSVSGFKCSASLTSLDLSRNS 133
Query: 222 ----IVRITAGALPRGLHSLNLSRNNIS---SIEGLRELTRLRVLDLSYNRILR---IGH 271
+ +T+ GL LN+S N + + G +L L VLDLS N I +G
Sbjct: 134 LSGPVTTLTSLGSCSGLKFLNVSSNTLDFPGKVSGGLKLNSLEVLDLSANSISGANVVGW 193
Query: 272 GLAS-CSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTA-KCLGQLAANYNSLQA 329
L+ C LK L ++GNKI + R + L LD+ N ST LG +A LQ
Sbjct: 194 VLSDGCGELKHLAISGNKISGDVDVSRCVNLEFLDVSSNNFSTGIPFLGDCSA----LQH 249
Query: 330 INLDGN 335
+++ GN
Sbjct: 250 LDISGN 255
Score = 37.4 bits (85), Expect = 0.085
Identities = 29/84 (34%), Positives = 41/84 (48%), Gaps = 5/84 (5%)
Query: 211 SLKVLNLAGNSI--VRITAGALPRG---LHSLNLSRNNISSIEGLRELTRLRVLDLSYNR 265
SL+VL+L+ NSI + L G L L +S N IS + L LD+S N
Sbjct: 174 SLEVLDLSANSISGANVVGWVLSDGCGELKHLAISGNKISGDVDVSRCVNLEFLDVSSNN 233
Query: 266 ILRIGHGLASCSSLKELYLAGNKI 289
L CS+L+ L ++GNK+
Sbjct: 234 FSTGIPFLGDCSALQHLDISGNKL 257
>LUM_RAT (P51886) Lumican precursor (Keratan sulfate proteoglycan
lumican) (KSPG lumican)
Length = 338
Score = 52.8 bits (125), Expect = 2e-06
Identities = 46/139 (33%), Positives = 74/139 (53%), Gaps = 11/139 (7%)
Query: 206 LSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSI--EGLRELTRLRVLDLSY 263
L SL+ L+L+ N + ++ AG LP L +L L N I++I E T L+ L LS+
Sbjct: 180 LKGLKSLEYLDLSFNQMSKLPAG-LPTSLLTLYLDNNKITNIPDEYFNRFTGLQYLRLSH 238
Query: 264 NRILRIG--HGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDL----RFNKISTAKCL 317
N + G + SSL EL L+ NK+ + ++ L+ L++ +F+ S K L
Sbjct: 239 NELADSGVPGNSFNISSLLELDLSYNKLKSIPTVNENLENYYLEVNKLEKFDVKSFCKIL 298
Query: 318 GQLAANYNSLQAINLDGNP 336
G L+ Y+ ++ + LDGNP
Sbjct: 299 GPLS--YSKIKHLRLDGNP 315
Score = 45.1 bits (105), Expect = 4e-04
Identities = 47/156 (30%), Positives = 71/156 (45%), Gaps = 31/156 (19%)
Query: 207 SAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNIS---SIEGLRELT--------- 254
S LK L++ N++ + G LP+ L L L+ N IS S +GL LT
Sbjct: 113 SKLKQLKKLHINYNNLTE-SVGPLPKSLQDLQLANNKISKLGSFDGLVNLTFIYLQHNQL 171
Query: 255 -------------RLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV--EGLHRLL 299
L LDLS+N++ ++ GL +SL LYL NKI + E +R
Sbjct: 172 KEEAVSASLKGLKSLEYLDLSFNQMSKLPAGLP--TSLLTLYLDNNKITNIPDEYFNRFT 229
Query: 300 KLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGN 335
L L L N+++ + G + N +SL ++L N
Sbjct: 230 GLQYLRLSHNELADSGVPGN-SFNISSLLELDLSYN 264
>ECM2_HUMAN (O94769) Extracellular matrix protein 2 precursor
(Matrix glycoprotein SC1/ECM2)
Length = 699
Score = 52.8 bits (125), Expect = 2e-06
Identities = 50/183 (27%), Positives = 84/183 (45%), Gaps = 39/183 (21%)
Query: 215 LNLAGNSIVRITAGALPRGLHSLNLSRNNISSI--EGLRELTRLRVLDLSYNRI------ 266
LN+ GN++++I + LP L L ++ NN+ +I E L +L +L L+L N +
Sbjct: 398 LNMDGNNLIQIPS-QLPSTLEELKVNENNLQAIDEESLSDLNQLVTLELEGNNLSEANVN 456
Query: 267 ------------LRIG--------HGLASCSSLKELYLAGNKIGEVEGL--HRLLKLSIL 304
LR+G GL S++ELYL N+I E+ + + K++++
Sbjct: 457 PLAFKPLKSLAYLRLGKNKFRIIPQGLPG--SIEELYLENNQIEEITEICFNHTRKINVI 514
Query: 305 DLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQLKKYLQGLLPHLVYYNRQA 364
LR+NKI + N +L++I+L N + YL L HLV Q
Sbjct: 515 VLRYNKIEENRIAPLAWINQENLESIDLSYNKLY------HVPSYLPKSLLHLVLLGNQI 568
Query: 365 MKV 367
++
Sbjct: 569 ERI 571
>CBP8_MOUSE (Q9DBB9) Carboxypeptidase N 83 kDa chain precursor
(Carboxypeptidase N regulatory subunit)
(Carboxypeptidase N polypeptide 2)
Length = 547
Score = 52.4 bits (124), Expect = 3e-06
Identities = 60/181 (33%), Positives = 92/181 (50%), Gaps = 21/181 (11%)
Query: 212 LKVLNLAGNSIVRITAGAL---PRGLHSLNLSRNNISSIE--GLRELTRLRVLDLSYNRI 266
L+ L L GN + R G L R L +LNL++N ++ + + LT L++L LS N +
Sbjct: 147 LESLQLQGNQL-RTLPGRLFQSLRDLRTLNLAQNLLTQLPKGAFQSLTGLQMLKLSNNML 205
Query: 267 LRIGHG-LASCSSLKELYLAGNKIGEVEG--LHRLLKLSILDLRFNKISTAKCLGQLAAN 323
R+ G L S SSL+EL+L GN I E+ +L L +L L+ N I L ++
Sbjct: 206 ARLPEGALGSLSSLQELFLDGNAITELSPHLFSQLFSLEMLWLQHNAICHLPV--SLFSS 263
Query: 324 YNSLQAINLDGNPCQKNVGDEQLKKYLQGLLPHLVYYNRQAMKVSTLKDGA----DRLVR 379
++L ++L N + E L + QGLL + YN ++ T+ +GA RLV
Sbjct: 264 LHNLTFLSLKDNALR--TLPEGLFAHNQGLLHLSLSYN----QLETIPEGAFTNLSRLVS 317
Query: 380 L 380
L
Sbjct: 318 L 318
>TLR7_MOUSE (P58681) Toll-like receptor 7 precursor
Length = 1050
Score = 52.0 bits (123), Expect = 3e-06
Identities = 42/139 (30%), Positives = 73/139 (52%), Gaps = 8/139 (5%)
Query: 205 FLSAFVSLKVLNLAGNSIVRITAGA---LPRGLHSLNLSRNNISSI--EGLRELTRLRVL 259
F +L+VL+++ NS+ + +P L +L+L++N + S + L+ L L +L
Sbjct: 646 FFKNLFNLEVLDISRNSLNSLPPEVFEGMPPNLKNLSLAKNGLKSFFWDRLQLLKHLEIL 705
Query: 260 DLSYNRILRIGHGLASCS-SLKELYLAGNKIGEVEG--LHRLLKLSILDLRFNKISTAKC 316
DLS+N++ ++ LA+CS SL L L N+I ++ L L+L LD+ NKI +
Sbjct: 706 DLSHNQLTKVPERLANCSKSLTTLILKHNQIRQLTKYFLEDALQLRYLDISSNKIQVIQK 765
Query: 317 LGQLAANYNSLQAINLDGN 335
N+L+ + L N
Sbjct: 766 TSFPENVLNNLEMLVLHHN 784
Score = 44.3 bits (103), Expect = 7e-04
Identities = 45/147 (30%), Positives = 69/147 (46%), Gaps = 20/147 (13%)
Query: 207 SAFVSLKVLNLAGNSIVRI--TAGALPRGLHSLNLSRN----NISSIEGLRELTRLRVLD 260
++ LKVL L NS+ + T R L L+LS+N I + L L L LD
Sbjct: 287 NSLTELKVLRLHSNSLQHVPPTWFKNMRNLQELDLSQNYLAREIEEAKFLHFLPNLVELD 346
Query: 261 LSYNRILRI-------GHGLASCSSLKELYLAGNKIGEVEG-----LHRLLKLSILDLRF 308
S+N L++ H L+S +LK L + G E++ LH+L +L +LDL
Sbjct: 347 FSFNYELQVYHASITLPHSLSSLENLKILRVKGYVFKELKNSSLSVLHKLPRLEVLDLGT 406
Query: 309 NKISTAKCLGQLAANYNSLQAINLDGN 335
N I A + ++ +L+ I+L N
Sbjct: 407 NFIKIADL--NIFKHFENLKLIDLSVN 431
Score = 43.1 bits (100), Expect = 0.002
Identities = 45/154 (29%), Positives = 62/154 (40%), Gaps = 21/154 (13%)
Query: 207 SAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSI--EGLRELTRLRVLDLSYN 264
S LK L L GN ++ I LP LH L+L NNI SI E L EL + L L N
Sbjct: 124 SGLSDLKALYLDGNQLLEIPQD-LPSSLHLLSLEANNIFSITKENLTELVNIETLYLGQN 182
Query: 265 RILR----IGHGLASCSSL--KELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLG 318
R + + + + L + L + K V + L ++L+L K
Sbjct: 183 CYYRNPCNVSYSIEKDAFLVMRNLKVLSLKDNNVTAVPTTLPPNLLELYLYNNIIKKIQE 242
Query: 319 QLAANYNSLQAINLDGN------------PCQKN 340
N N LQ ++L GN PC+ N
Sbjct: 243 NDFNNLNELQVLDLSGNCPRCYNVPYPCTPCENN 276
>LIG2_HUMAN (O94898) Leucine-rich repeats and immunoglobulin-like
domains protein 2 precursor (LIG-2)
Length = 1065
Score = 52.0 bits (123), Expect = 3e-06
Identities = 60/190 (31%), Positives = 84/190 (43%), Gaps = 15/190 (7%)
Query: 211 SLKVLNLAGNSIVRITAGALP--RGLHSLNLSRNNISSI--EGLRELTRLRVLDLSYNRI 266
+++ L L N++ R+ G L R L L +S+N I I + RL LDLSYN++
Sbjct: 264 NMEELELEHNNLTRVNKGWLYGLRMLQQLYVSQNAIERISPDAWEFCQRLSELDLSYNQL 323
Query: 267 LRIGH-GLASCSSLKELYLAGNKIGEV-EGLHRLL-KLSILDLRFNKISTA-KCLGQLAA 322
R+ S L+ L L N++ + +G+ R L L LDLR N+IS A + + A
Sbjct: 324 TRLDESAFVGLSLLERLNLGDNRVTHIADGVFRFLSNLQTLDLRNNEISWAIEDASEAFA 383
Query: 323 NYNSLQAINLDGNPCQKNVGDEQLKKYLQGL--LPHLVYYNRQAMKVSTLKDGADRLVRL 380
SL + L GN + KK GL L HL N M + L L
Sbjct: 384 GLTSLTKLILQGNQIKSIT-----KKAFIGLESLEHLDLNNNAIMSIQENAFSQTHLKEL 438
Query: 381 GTNDRSLRVD 390
N SL D
Sbjct: 439 ILNTSSLLCD 448
Score = 43.5 bits (101), Expect = 0.001
Identities = 56/224 (25%), Positives = 91/224 (40%), Gaps = 40/224 (17%)
Query: 121 DFKVKRIEDWVVDL-----QHCGPPVEEINELPESVDPVVDINTINGVTAAGVNHKITPG 175
DF R+ +W + L Q E+ E+P +P +I ++ V H I P
Sbjct: 81 DFSHNRLSNWNISLESQTLQEVKMNYNELTEIPYFGEPTSNITLLSLV------HNIIPE 134
Query: 176 MEAAKRYISSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPR-GL 234
+ NA A Q + +L+ L+L+ N I I + PR L
Sbjct: 135 I------------NAQALQF-------------YPALESLDLSSNIISEIKTSSFPRMQL 169
Query: 235 HSLNLSRNNISSIEGL---RELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGE 291
LNLS N I+++E + L V+ L+ NR+ I + L+ L L N+I
Sbjct: 170 KYLNLSNNRITTLEAGCFDNLSSSLLVVKLNRNRMSMIPPKIFKLPHLQFLELKRNRIKI 229
Query: 292 VEGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGN 335
VEGL S+ L+ + +K N+++ + L+ N
Sbjct: 230 VEGLTFQGLDSLRSLKMQRNGISKLKDGAFFGLNNMEELELEHN 273
>LIG1_HUMAN (Q96JA1) Leucine-rich repeats and immunoglobulin-like
domains protein 1 precursor (LIG-1)
Length = 1093
Score = 50.8 bits (120), Expect = 7e-06
Identities = 52/177 (29%), Positives = 81/177 (45%), Gaps = 14/177 (7%)
Query: 228 GALPRGLHSLNLSRNNISSIE--GLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLA 285
G LP SLNLS N +S I+ G +L L+ + L+ N + + A+ S + L+L
Sbjct: 64 GDLPSWTRSLNLSYNKLSEIDPAGFEDLPNLQEVYLNNNELTAVPSLGAASSHVVSLFLQ 123
Query: 286 GNKIGEVEG--LHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQKNVGD 343
NKI VEG L L L +LDL N I+ + + ++ +NL GN +G
Sbjct: 124 HNKIRSVEGSQLKAYLSLEVLDLSLNNITEVR--NTCFPHGPPIKELNLAGN----RIGT 177
Query: 344 EQLKKYLQGLLPHLVYYNRQAMKVSTLKDGA---DRLVRLGTNDRSLRVDRKTTRKG 397
+L + GL L+ +++ L A RL +L N +R+ T +G
Sbjct: 178 LELGAF-DGLSRSLLTLRLSKNRITQLPVRAFKLPRLTQLDLNRNRIRLIEGLTFQG 233
Score = 47.8 bits (112), Expect = 6e-05
Identities = 47/141 (33%), Positives = 68/141 (47%), Gaps = 15/141 (10%)
Query: 206 LSAFVSLKVLNLAGNSIVRITAGALPRG--LHSLNLSRNNISSIE--GLRELTR-LRVLD 260
L A++SL+VL+L+ N+I + P G + LNL+ N I ++E L+R L L
Sbjct: 135 LKAYLSLEVLDLSLNNITEVRNTCFPHGPPIKELNLAGNRIGTLELGAFDGLSRSLLTLR 194
Query: 261 LSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGL--HRLLKLSILDLRFNKIST----- 313
LS NRI ++ L +L L N+I +EGL L L +L L+ N IS
Sbjct: 195 LSKNRITQLPVRAFKLPRLTQLDLNRNRIRLIEGLTFQGLNSLEVLKLQRNNISKLTDGA 254
Query: 314 ---AKCLGQLAANYNSLQAIN 331
+ L YNSL +N
Sbjct: 255 FWGLSKMHVLHLEYNSLVEVN 275
>CYAA_YEAST (P08678) Adenylate cyclase (EC 4.6.1.1) (ATP
pyrophosphate-lyase) (Adenylyl cyclase)
Length = 2026
Score = 50.8 bits (120), Expect = 7e-06
Identities = 39/127 (30%), Positives = 63/127 (48%), Gaps = 8/127 (6%)
Query: 211 SLKVLNLAGNSIVRITAGALP-RGLHSLNLSRNNISSI-EGLRELTRLRVLDLSYNRILR 268
+L +LNL N + + AG + + L L+LS N E + T L +DLSYN+I
Sbjct: 887 NLTILNLQCNELESLPAGFVELKNLQLLDLSSNKFMHYPEVINYCTNLLQIDLSYNKIQS 946
Query: 269 IGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQ 328
+ L ++ L+ NK+ + L + L L+LR+N+IS+ K N ++LQ
Sbjct: 947 LPQSTKYLVKLAKMNLSHNKLNFIGDLSEMTDLRTLNLRYNRISSIK------TNASNLQ 1000
Query: 329 AINLDGN 335
+ L N
Sbjct: 1001 NLFLTDN 1007
Score = 42.0 bits (97), Expect = 0.003
Identities = 50/196 (25%), Positives = 73/196 (36%), Gaps = 38/196 (19%)
Query: 183 ISSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRN 242
IS LT + N + P LS SL+ L+L N+I G L SLN+S N
Sbjct: 1083 ISKLTKLVFLSVARNKLEYIPPELSQLKSLRTLDLHSNNIRDFVDGMENLELTSLNISSN 1142
Query: 243 NI--SSIEG---------------------------------LRELTRLRVLDLSYNRIL 267
SS+E L+VL+LSYN
Sbjct: 1143 AFGNSSLENSFYHNMSYGSKLSKSLMFFIAADNQFDDAMWPLFNCFVNLKVLNLSYNNFS 1202
Query: 268 RIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSL 327
+ H S+ ELYL+GNK+ + G L S+ L N +L +N + L
Sbjct: 1203 DVSH--MKLESITELYLSGNKLTTLSGDTVLKWSSLKTLMLNSNQMLSLPAEL-SNLSQL 1259
Query: 328 QAINLDGNPCQKNVGD 343
++ N + N+ +
Sbjct: 1260 SVFDVGANQLKYNISN 1275
>TLR2_MOUSE (Q9QUN7) Toll-like receptor 2 precursor
Length = 784
Score = 50.1 bits (118), Expect = 1e-05
Identities = 35/88 (39%), Positives = 53/88 (59%), Gaps = 9/88 (10%)
Query: 256 LRVLDLSYNRILRIGHG-LASCSSLKELYLAGNKIGEVEG--LHRLLKLSILDLRFNKIS 312
++ LDLS+N+I IGHG L +C++L+ L L ++I +EG + L L LDL N +S
Sbjct: 54 MKSLDLSFNKITYIGHGDLRACANLQVLILKSSRINTIEGDAFYSLGSLEHLDLSDNHLS 113
Query: 313 --TAKCLGQLAANYNSLQAINLDGNPCQ 338
++ G L +SL+ +NL GNP Q
Sbjct: 114 SLSSSWFGPL----SSLKYLNLMGNPYQ 137
>LAP4_MOUSE (Q80U72) LAP4 protein (Scribble homolog protein)
Length = 1612
Score = 50.1 bits (118), Expect = 1e-05
Identities = 46/129 (35%), Positives = 70/129 (53%), Gaps = 6/129 (4%)
Query: 188 ANASAAQLANHGLVVVPF-LSAFVSLKVLNLAGNSIVRI--TAGALPRGLHSLNLSRNNI 244
AN +L + L +P LS V L+ L+L GN + + T GALP L L L RN +
Sbjct: 151 ANLVTLELRENLLKSLPASLSFLVKLEQLDLGGNDLEVLPDTLGALPN-LRELWLDRNQL 209
Query: 245 SSIEG-LRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLS 302
S++ L L RL LD+S NR+ + L + L +L L+ N + + EG+ +L +LS
Sbjct: 210 SALPPELGNLRRLVCLDVSENRLEELPVELGGLALLTDLLLSQNLLQRLPEGIGQLKQLS 269
Query: 303 ILDLRFNKI 311
IL + N++
Sbjct: 270 ILKVDQNRL 278
Score = 37.4 bits (85), Expect = 0.085
Identities = 24/80 (30%), Positives = 43/80 (53%), Gaps = 2/80 (2%)
Query: 234 LHSLNLSRNNISSI-EGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV 292
L L LS+N + + EG+ +L +L +L + NR+ + + C +L EL L N + +
Sbjct: 245 LTDLLLSQNLLQRLPEGIGQLKQLSILKVDQNRLCEVTEAIGDCENLSELILTENLLTAL 304
Query: 293 -EGLHRLLKLSILDLRFNKI 311
L +L KL+ L++ N +
Sbjct: 305 PHSLGKLTKLTNLNVDRNHL 324
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.132 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 53,626,129
Number of Sequences: 164201
Number of extensions: 2251744
Number of successful extensions: 5759
Number of sequences better than 10.0: 229
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 195
Number of HSP's that attempted gapping in prelim test: 5310
Number of HSP's gapped (non-prelim): 467
length of query: 475
length of database: 59,974,054
effective HSP length: 114
effective length of query: 361
effective length of database: 41,255,140
effective search space: 14893105540
effective search space used: 14893105540
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0203.3


