

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.14
(896 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
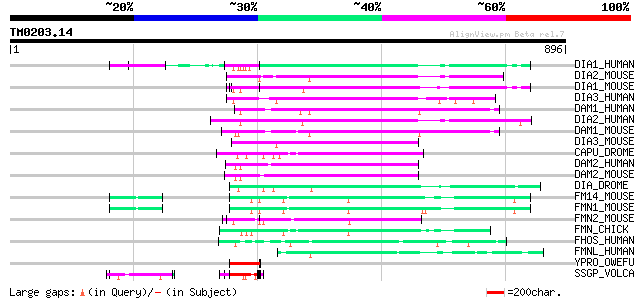
DIA1_HUMAN (O60610) Diaphanous protein homolog 1 (Diaphanous-rel... 169 2e-41
DIA2_MOUSE (O70566) Diaphanous protein homolog 2 (Diaphanous-rel... 164 1e-39
DIA1_MOUSE (O08808) Diaphanous protein homolog 1 (Diaphanous-rel... 163 2e-39
DIA3_HUMAN (Q9NSV4) Diaphanous protein homolog 3 (Diaphanous-rel... 162 3e-39
DAM1_HUMAN (Q9Y4D1) Disheveled associated activator of morphogen... 159 4e-38
DIA2_HUMAN (O60879) Diaphanous protein homolog 2 (Diaphanous-rel... 153 2e-36
DAM1_MOUSE (Q8BPM0) Disheveled associated activator of morphogen... 152 3e-36
DIA3_MOUSE (Q9Z207) Diaphanous protein homolog 3 (Diaphanous-rel... 149 2e-35
CAPU_DROME (Q24120) Cappuccino protein 135 6e-31
DAM2_HUMAN (Q86T65) Disheveled associated activator of morphogen... 134 8e-31
DAM2_MOUSE (Q80U19) Disheveled associated activator of morphogen... 133 2e-30
DIA_DROME (P48608) Diaphanous protein 132 3e-30
FM14_MOUSE (Q05859) Formin 1 isoform IV (Limb deformity protein) 122 3e-27
FMN1_MOUSE (Q05860) Formin 1 isoforms I/II/III (Limb deformity p... 120 2e-26
FMN2_MOUSE (Q9JL04) Formin 2 119 3e-26
FMN_CHICK (Q05858) Formin (Limb deformity protein) 117 2e-25
FHOS_HUMAN (Q9Y613) FH1/FH2 domains-containing protein (Formin h... 91 1e-17
FMNL_HUMAN (O95466) Formin-like protein (Protein C17orf1) 84 1e-15
YPRO_OWEFU (P21260) Hypothetical proline-rich protein (Fragment) 79 4e-14
SSGP_VOLCA (P21997) Sulfated surface glycoprotein 185 precursor ... 75 8e-13
>DIA1_HUMAN (O60610) Diaphanous protein homolog 1 (Diaphanous-related
formin 1) (DRF1)
Length = 1248
Score = 169 bits (429), Expect = 2e-41
Identities = 177/681 (25%), Positives = 263/681 (37%), Gaps = 148/681 (21%)
Query: 192 PANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKGD 251
P+ +P P P S + P P A T+PP PPP P P P PP G
Sbjct: 568 PSRAPVPPAPPLPGDSGTIIPPPPAPGDSTTPP---PPPPP---------PPPPPPLPG- 614
Query: 252 RKQKAIILAVTVSGIIIMIGLALCYRESRRSDRADKDEGPLLILTSSDYSGGKQKVVRLG 311
G A+ D PL
Sbjct: 615 -------------------GTAISPPPPLSGDATIPPPPPL------------------- 636
Query: 312 NANNEGSGISNGKKPSNVRNWSMKAGDNNTSVVEATSSEGVGQVPEPPN------GKPPP 365
EG GI + PS++ T++ G ++P PP G PPP
Sbjct: 637 ---PEGVGIPS---PSSL--------PGGTAIPPPPPLPGSARIPPPPPPLPGSAGIPPP 682
Query: 366 PPP-----GPPPPPPPRK------PPAPAPR----PPPPPRAGQPPPAPPKPVKVGNQGA 410
PPP G PPPPPP PP P P PPPPP G PPP PP V
Sbjct: 683 PPPLPGEAGMPPPPPPLPGGPGIPPPPPFPGGPGIPPPPPGMGMPPP-PPFGFGVPAAPV 741
Query: 411 NQGGTSDGDAPKPK--LKPFFWDKVAAKP-DQQMVWHEISAGSFVFNE--EKMESLFGCA 465
G + KP+ L+ W K+ A+ Q W ++ F NE K+ F
Sbjct: 742 LPFGLTPKKLYKPEVQLRRPNWSKLVAEDLSQDCFWTKVKEDRFENNELFAKLTLTFSAQ 801
Query: 466 NQNRNERK--KDSPSVDTS-VQYIQIIDPKKAQNLSILLRALNVTTDEVIDALREGNE-- 520
+ + +++ ++ SV V+ ++++D K AQNLSI L + + E+ + + E NE
Sbjct: 802 TKTKKDQEGGEEKKSVQKKKVKELKVLDSKTAQNLSIFLGSFRMPYQEIKNVILEVNEAV 861
Query: 521 IPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFLKVLVDIPFAFKRLETLLFMFI 580
+ +IQ L+K P E+ L E L +E+F V+ +P RL +LF
Sbjct: 862 LTESMIQNLIKQMPEPEQLKMLSELKDEYDDLAESEQFGVVMGTVPRLRPRLNAILFKLQ 921
Query: 581 LREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLL 640
E+ +IK ++ A ++LRKS F LLE L GN MN G+ GA F + L
Sbjct: 922 FSEQVENIKPEIVSVTAACEELRKSESFSNLLEITLLVGNYMNAGSRNAGAFGFNISFLC 981
Query: 641 KLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERESRSVVSSVGTDQDVDESPEASEEH 700
KL D K TD K TLLHF+ E E
Sbjct: 982 KLRDTKSTDQKMTLLHFLA-----------------------------------ELCEND 1006
Query: 701 YRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLGHSLAKTQEFMNTDLKSLDEESE 760
Y V +EL V++A+ + + L + ++ ++ + + + DE+ +
Sbjct: 1007 YPD-----VLKFPDELAHVEKASRVSAENLQKNLDQMKKQISDVERDVQNFPAATDEKDK 1061
Query: 761 FQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADYFHGNAGRDEGLRLFLIVRDFLIIL 820
F FV+ A+E+ L+ + K +YF + + L V +F + L
Sbjct: 1062 FVEKMTSFVKDAQEQYNKLRMMHSNMETLYKELGEYFLFDPKK-------LSVEEFFMDL 1114
Query: 821 DKVCKEVRDKTLRAVKSSQRK 841
R+ L+AVK +Q++
Sbjct: 1115 ----HNFRNMFLQAVKENQKR 1131
Score = 47.8 bits (112), Expect = 1e-04
Identities = 27/68 (39%), Positives = 32/68 (46%), Gaps = 12/68 (17%)
Query: 348 SSEGVGQVPEPPNGKP-PPPPPGP-------PPPPPP----RKPPAPAPRPPPPPRAGQP 395
S+ + P P+ P PP PP P PPPP P PP P P PPPPP G
Sbjct: 557 SAAAITVPPSVPSRAPVPPAPPLPGDSGTIIPPPPAPGDSTTPPPPPPPPPPPPPLPGGT 616
Query: 396 PPAPPKPV 403
+PP P+
Sbjct: 617 AISPPPPL 624
Score = 47.4 bits (111), Expect = 2e-04
Identities = 33/97 (34%), Positives = 40/97 (41%), Gaps = 7/97 (7%)
Query: 162 PHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSS---PTPTPEN---SSPSSTPSPT 215
P + +P P P P P L T P P + ++ P P PE SPSS P T
Sbjct: 593 PGDSTTPPPPPPPPPPPPPLPGGTAISPPPPLSGDATIPPPPPLPEGVGIPSPSSLPGGT 652
Query: 216 AMVPPTS-PPRQHPPPSPEKLQHVTYAPDPSPPDKGD 251
A+ PP P PP P L P P PP G+
Sbjct: 653 AIPPPPPLPGSARIPPPPPPLPGSAGIPPPPPPLPGE 689
Score = 38.1 bits (87), Expect = 0.11
Identities = 32/98 (32%), Positives = 37/98 (37%), Gaps = 24/98 (24%)
Query: 161 PPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPP 220
PP L P +GT+ P PA P +S + P P P P P TA+ PP
Sbjct: 574 PPAPPL-PGDSGTIIPPPPA-----------PGDSTTPPPPPPPPPPPPPLPGGTAISPP 621
Query: 221 -------TSPPRQHPP-----PSPEKLQHVTYAPDPSP 246
T PP P PSP L T P P P
Sbjct: 622 PPLSGDATIPPPPPLPEGVGIPSPSSLPGGTAIPPPPP 659
Score = 36.6 bits (83), Expect = 0.31
Identities = 29/94 (30%), Positives = 34/94 (35%), Gaps = 12/94 (12%)
Query: 158 ISDPPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAM 217
+ D S S A P +P P P P +S + P P S+TP
Sbjct: 546 LEDAKKEMASLSAAAITVPPSVPSRAPVPPAPPLPGDS-GTIIPPPPAPGDSTTP----- 599
Query: 218 VPPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKGD 251
PP PP PPP P A P PP GD
Sbjct: 600 -PPPPPPPPPPPPLPGGT-----AISPPPPLSGD 627
Score = 35.0 bits (79), Expect = 0.89
Identities = 22/65 (33%), Positives = 23/65 (34%), Gaps = 1/65 (1%)
Query: 161 PPHNELSPSPAGTVPSPGPAL-ESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVP 219
PP P AG P P P E+ P P P P P P P P P M
Sbjct: 667 PPPPPPLPGSAGIPPPPPPLPGEAGMPPPPPPLPGGPGIPPPPPFPGGPGIPPPPPGMGM 726
Query: 220 PTSPP 224
P PP
Sbjct: 727 PPPPP 731
Score = 34.3 bits (77), Expect = 1.5
Identities = 27/78 (34%), Positives = 28/78 (35%), Gaps = 12/78 (15%)
Query: 167 SPS--PAGTV-----PSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVP 219
SPS P GT P PG A P P P A P P P P + P P P
Sbjct: 644 SPSSLPGGTAIPPPPPLPGSARIPPPPPPLPGSAGIPPPPPPLPGEAGMPPPPPPLPGGP 703
Query: 220 PTSPPRQHP-----PPSP 232
PP P PP P
Sbjct: 704 GIPPPPPFPGGPGIPPPP 721
Score = 33.9 bits (76), Expect = 2.0
Identities = 30/90 (33%), Positives = 33/90 (36%), Gaps = 16/90 (17%)
Query: 154 GRNLISDPPHNELSPSPAGT-VPSP-----GPALESPTPNHGPTPANSPSSPTPTPENSS 207
G I PP P P G +PSP G A+ P P G P P P
Sbjct: 626 GDATIPPPP-----PLPEGVGIPSPSSLPGGTAIPPPPPLPGSARIPPPPPPLPGSAGIP 680
Query: 208 PSSTPSP-TAMVPPTSPPRQH----PPPSP 232
P P P A +PP PP PPP P
Sbjct: 681 PPPPPLPGEAGMPPPPPPLPGGPGIPPPPP 710
>DIA2_MOUSE (O70566) Diaphanous protein homolog 2
(Diaphanous-related formin 2) (DRF2) (mDia3)
Length = 1098
Score = 164 bits (415), Expect = 1e-39
Identities = 126/467 (26%), Positives = 199/467 (41%), Gaps = 64/467 (13%)
Query: 351 GVGQVPEPPNGKPPPPP-PGPPPPPPPRKPPAPA-PRPPPPPRAGQPPPAPPK------- 401
G G P PP PPPPP PG PPPPP P P P PPPPP +G PPP PP
Sbjct: 554 GAGPCPPPPPPPPPPPPLPGVVPPPPPPLPGMPGIPPPPPPPLSGVPPPPPPPGGVFPLL 613
Query: 402 --PVKVGNQGANQGGTSDGDAPKPKLKPFFWDKVAAKP-DQQMVWHEISAGSFVFNEEKM 458
P+++ G Q D P ++ W K+ K + VW ++ + +
Sbjct: 614 SGPIELP-YGMKQKKLYKPDIPMKRIN---WSKIEPKELSENCVWLKLKEEKYENADLFA 669
Query: 459 ESLFGCANQNRNERKKDSPSVDTS------VQYIQIIDPKKAQNLSILLRALNVTTDEVI 512
+ +Q + +R ++ + S V+ ++I+D K AQNLSI L + + +E+
Sbjct: 670 KLALTFPSQMKGQRNTEAAEENRSGPPKKKVKELRILDTKTAQNLSIFLGSYRMPYEEIK 729
Query: 513 DALREGNE--IPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFLKVLVDIPFAFK 570
+ + E NE + LIQ L+K P +L E L E+F V+ +
Sbjct: 730 NIILEVNEEMLSEALIQNLVKYLPDQNALRELAQLKSEYDDLCEPEQFGVVMSTVKMLRP 789
Query: 571 RLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGG 630
RL ++LF E ++IK S + +A ++L+KS F +LLE +L GN MN G+
Sbjct: 790 RLTSILFKLTFEEHVNNIKPSIIAVTLACEELKKSESFKRLLELILLVGNYMNSGSRNAQ 849
Query: 631 AQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERESRSVVSSVGTDQDV 690
+ F+++ L K+ D K D K+TLLHF+
Sbjct: 850 SLGFKINFLCKIKDTKSADQKSTLLHFLA------------------------------- 878
Query: 691 DESPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLGHSLAKTQEFMNT 750
E +E YR + +ELE V+ A + L S + + S+ ++ +
Sbjct: 879 ----EICDEKYRD-----ILKFPDELEHVESAGKVSAQILKSNLVAMEQSILHLEKNIKN 929
Query: 751 DLKSLDEESEFQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADYF 797
+ +F F + ARE+ L ++ +S +YF
Sbjct: 930 FPPAESHHDKFVEKMMSFTQNAREQYDKLSTMHSNMLKLYESLGEYF 976
Score = 43.5 bits (101), Expect = 0.003
Identities = 31/81 (38%), Positives = 34/81 (41%), Gaps = 15/81 (18%)
Query: 174 VPS--PGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVP--PTSPPRQHPP 229
VPS PGP P P GP P P P P P P P P +P P PP PP
Sbjct: 539 VPSAIPGPPPPPPLPGAGPCPPPPPPPPPPPP---LPGVVPPPPPPLPGMPGIPP---PP 592
Query: 230 PSPEKLQHVTYAPDPSPPDKG 250
P P ++ P P PP G
Sbjct: 593 PPP-----LSGVPPPPPPPGG 608
Score = 36.2 bits (82), Expect = 0.40
Identities = 24/67 (35%), Positives = 25/67 (36%), Gaps = 8/67 (11%)
Query: 168 PSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTS--PPR 225
P P +P GP P P P P P P P P P P PP S PP
Sbjct: 546 PPPPPPLPGAGPCPPPPPPPPPPPPLPGVVPPPPPPLPGMPGIPPPPP---PPLSGVPP- 601
Query: 226 QHPPPSP 232
PPP P
Sbjct: 602 --PPPPP 606
>DIA1_MOUSE (O08808) Diaphanous protein homolog 1 (Diaphanous-related
formin 1) (DRF1) (mDIA1) (p140mDIA)
Length = 1255
Score = 163 bits (412), Expect = 2e-39
Identities = 146/512 (28%), Positives = 218/512 (42%), Gaps = 74/512 (14%)
Query: 351 GVGQVPEPPN-----GKPPPPP-PGPP--PPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
G G P PP G PPPPP PG P PPPPP P AP PPPPP G PPP PP
Sbjct: 680 GTGIPPPPPPLPGSVGVPPPPPLPGGPGLPPPPPPFPGAPGI-PPPPPGMGVPPP-PPFG 737
Query: 403 VKVGNQGANQGGTSDGDAPKPK--LKPFFWDKVAAKP-DQQMVWHEISAGSFVFNE--EK 457
V G + KP+ L+ W K A+ Q W ++ F NE K
Sbjct: 738 FGVPAAPVLPFGLTPKKVYKPEVQLRRPNWSKFVAEDLSQDCFWTKVKEDRFENNELFAK 797
Query: 458 MESLFGCANQNRNERK-----KDSPSVDTS-VQYIQIIDPKKAQNLSILLRALNVTTDEV 511
+ F + +K ++ SV V+ ++++D K AQNLSI L + + E+
Sbjct: 798 LTLAFSAQTKTSKAKKDQEGGEEKKSVQKKKVKELKVLDSKTAQNLSIFLGSFRMPYQEI 857
Query: 512 IDALREGNE--IPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFLKVLVDIPFAF 569
+ + E NE + +IQ L+K P E+ L E L +E+F V+ +P
Sbjct: 858 KNVILEVNEAVLTESMIQNLIKQMPEPEQLKMLSELKEEYDDLAESEQFGVVMGTVPRLR 917
Query: 570 KRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRG 629
RL +LF E+ +IK ++ A ++LRKS F LLE L GN MN G+
Sbjct: 918 PRLNAILFKLQFSEQVENIKPEIVSVTAACEELRKSENFSSLLELTLLVGNYMNAGSRNA 977
Query: 630 GAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERESRSVVSSVGTDQD 689
GA F + L KL D K D K TLLHF+ +
Sbjct: 978 GAFGFNISFLCKLRDTKSADQKMTLLHFLAELC--------------------------- 1010
Query: 690 VDESPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLGHSLAKTQEFMN 749
++ PE V +EL V++A+ + + L ++ ++ +A + +
Sbjct: 1011 ENDHPE-------------VLKFPDELAHVEKASRVSAENLQKSLDQMKKQIADVERDVQ 1057
Query: 750 TDLKSLDEESEFQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADYFHGNAGRDEGLRL 809
+ DE+ +F FV+ A+E+ L+ + K DYF + +
Sbjct: 1058 NFPAATDEKDKFVEKMTSFVKDAQEQYNKLRMMHSNMETLYKELGDYFVFDPKK------ 1111
Query: 810 FLIVRDFLIILDKVCKEVRDKTLRAVKSSQRK 841
L V +F + L R+ L+AVK +Q++
Sbjct: 1112 -LSVEEFFMDL----HNFRNMFLQAVKENQKR 1138
Score = 47.4 bits (111), Expect = 2e-04
Identities = 24/51 (47%), Positives = 25/51 (48%), Gaps = 4/51 (7%)
Query: 356 PEPPNGKPPPPPPGPP----PPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
P P G PP PP PP PPPPP A P PP P + PP PP P
Sbjct: 592 PPLPGGVVPPSPPLPPGTCIPPPPPLPGGACIPPPPQLPGSAAIPPPPPLP 642
Score = 46.2 bits (108), Expect = 4e-04
Identities = 25/51 (49%), Positives = 26/51 (50%), Gaps = 4/51 (7%)
Query: 356 PEPPNG--KPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAG--QPPPAPPKP 402
P PP PPPP PG PPP + P A PPPPP G PP PP P
Sbjct: 604 PLPPGTCIPPPPPLPGGACIPPPPQLPGSAAIPPPPPLPGVASIPPPPPLP 654
Score = 45.1 bits (105), Expect = 9e-04
Identities = 21/45 (46%), Positives = 21/45 (46%)
Query: 358 PPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
PP PPP P G PP PP P P PPP P PP P P
Sbjct: 586 PPPPPPPPLPGGVVPPSPPLPPGTCIPPPPPLPGGACIPPPPQLP 630
Score = 43.1 bits (100), Expect = 0.003
Identities = 36/107 (33%), Positives = 39/107 (35%), Gaps = 17/107 (15%)
Query: 161 PPHNELSPSPAGTVPS-----PGPALESPTPNHG----PTPANSPSS-----PTPTPENS 206
PP P P G VP PG + P P G P P P S P P P +
Sbjct: 586 PPPPPPPPLPGGVVPPSPPLPPGTCIPPPPPLPGGACIPPPPQLPGSAAIPPPPPLPGVA 645
Query: 207 S---PSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKG 250
S P P TA+ PP P P P L T P P PP G
Sbjct: 646 SIPPPPPLPGATAIPPPPPLPGATAIPPPPPLPGGTGIPPPPPPLPG 692
Score = 40.8 bits (94), Expect = 0.016
Identities = 30/70 (42%), Positives = 32/70 (44%), Gaps = 16/70 (22%)
Query: 341 TSVVEATSSEGVGQVPEPPNGKPPPPPPGPP----PPPPPRKPPAPA----PRPPPPPRA 392
++VV A S VP P PP PG PPPPP PP P P PP PP
Sbjct: 557 SAVVVAPSVSSSAAVP------PAPPLPGDSGTVIPPPPPP-PPLPGGVVPPSPPLPPGT 609
Query: 393 GQPPPAPPKP 402
PPP PP P
Sbjct: 610 CIPPP-PPLP 618
Score = 36.2 bits (82), Expect = 0.40
Identities = 25/74 (33%), Positives = 27/74 (35%), Gaps = 8/74 (10%)
Query: 154 GRNLISDPPHNELSPSPAGT-VPSPGPALESPT--PNHGPTPANSPSSPTPTPENSSPSS 210
G I PP P P GT +P P P L P P P P P P +P
Sbjct: 667 GATAIPPPP-----PLPGGTGIPPPPPPLPGSVGVPPPPPLPGGPGLPPPPPPFPGAPGI 721
Query: 211 TPSPTAMVPPTSPP 224
P P M P PP
Sbjct: 722 PPPPPGMGVPPPPP 735
Score = 33.9 bits (76), Expect = 2.0
Identities = 30/94 (31%), Positives = 35/94 (36%), Gaps = 9/94 (9%)
Query: 161 PPHNELSPSPAGTV-PSPGPALESPTPNHGPTPANSPSS----PTPTPENS---SPSSTP 212
PP L P +GTV P P P P P+P P + P P P + P P
Sbjct: 572 PPAPPL-PGDSGTVIPPPPPPPPLPGGVVPPSPPLPPGTCIPPPPPLPGGACIPPPPQLP 630
Query: 213 SPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSP 246
A+ PP P P P L T P P P
Sbjct: 631 GSAAIPPPPPLPGVASIPPPPPLPGATAIPPPPP 664
Score = 33.5 bits (75), Expect = 2.6
Identities = 27/82 (32%), Positives = 30/82 (35%), Gaps = 7/82 (8%)
Query: 167 SPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSP--TAMVPPTSPP 224
S S + VP P P P P T P P P P P S P P T + PP P
Sbjct: 564 SVSSSAAVP-PAP----PLPGDSGTVIPPPPPPPPLPGGVVPPSPPLPPGTCIPPPPPLP 618
Query: 225 RQHPPPSPEKLQHVTYAPDPSP 246
P P +L P P P
Sbjct: 619 GGACIPPPPQLPGSAAIPPPPP 640
Score = 32.3 bits (72), Expect = 5.8
Identities = 31/103 (30%), Positives = 36/103 (34%), Gaps = 12/103 (11%)
Query: 154 GRNLISDPPHNELSPSPA-GTVPSPGP-----ALESPTPNHGPTPANSPSSPTPTPENSS 207
G I PP P P ++P P P A+ P P G T A P P P
Sbjct: 631 GSAAIPPPP-----PLPGVASIPPPPPLPGATAIPPPPPLPGAT-AIPPPPPLPGGTGIP 684
Query: 208 PSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKG 250
P P P ++ P PP P P AP PP G
Sbjct: 685 PPPPPLPGSVGVPPPPPLPGGPGLPPPPPPFPGAPGIPPPPPG 727
Score = 32.3 bits (72), Expect = 5.8
Identities = 20/58 (34%), Positives = 22/58 (37%), Gaps = 3/58 (5%)
Query: 345 EATSSEGVGQVPEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
E S V P + PP P P PP P P PP G PP+PP P
Sbjct: 552 EMASLSAVVVAPSVSSSAAVPPAPPLPGDSGTVIPPPPPP---PPLPGGVVPPSPPLP 606
>DIA3_HUMAN (Q9NSV4) Diaphanous protein homolog 3
(Diaphanous-related formin 3) (DRF3) (Fragment)
Length = 853
Score = 162 bits (411), Expect = 3e-39
Identities = 136/478 (28%), Positives = 214/478 (44%), Gaps = 70/478 (14%)
Query: 351 GVGQVPEPP-----NGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRA---GQ-PPPAPPK 401
G +P PP G PPPPPP PPPP P + P P PPPPP GQ PP P
Sbjct: 314 GHSALPPPPPLPSGGGVPPPPPPPPPPPLPGMRMPFSGPVPPPPPLGFLGGQNSPPLPIL 373
Query: 402 PVKVGNQGANQGGTSDGDAPKPKLKPFF------WDKVAA-KPDQQMVWHEISAGSFVFN 454
P G PK + KP W K+ + + W +++ +
Sbjct: 374 PF--------------GLKPKKEFKPEISMRRLNWLKIRPHEMTENCFWIKVNENKYENV 419
Query: 455 EE--KMESLFGCANQNRNERK--KDSPSVDTSVQYIQIIDPKKAQNLSILLRALNVTTDE 510
+ K+E+ F C + R E + ++ S+ ++ ++ +D K AQNLSI L + V +E
Sbjct: 420 DLLCKLENTFCCQQKERREEEDIEEKKSIKKKIKELKFLDSKIAQNLSIFLSSFRVPYEE 479
Query: 511 VIDALREGNE--IPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFLKVLVDIPFA 568
+ + E +E + +IQ L+K P E+ L F E S L E+F+ V+ ++
Sbjct: 480 IRMMILEVDETRLAESMIQNLIKHLPDQEQLNSLSQFKSEYSNLCEPEQFVVVMSNVKRL 539
Query: 569 FKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRMNDGTYR 628
RL +LF E+ ++IK + A ++++KS+ F KLLE VL GN MN G+
Sbjct: 540 RPRLSAILFKLQFEEQVNNIKPDIMAVSTACEEIKKSKSFSKLLELVLLMGNYMNAGSRN 599
Query: 629 GGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERESRSVVSSVGTDQ 688
F L +L KL D K D KTTLLHF+V+ E + +++ V +
Sbjct: 600 AQTFGFNLSSLCKLKDTKSADQKTTLLHFLVE-----------ICEEKYPDILNFVDDLE 648
Query: 689 DVDE----SPEASEEHYRSLGLQVVSSLSNELED-----------VKRAALIDGDALSSA 733
+D+ S E E++ R +G Q + L ELE V + +I
Sbjct: 649 PLDKASKVSVETLEKNLRQMGRQ-LQQLEKELETFPPPEDLHDKFVTKIFVISAKEQYET 707
Query: 734 VSKLGHSLAKTQE----FMNTDLKSLDEE---SEFQRCTERFVEKAREEVTWLQEEEK 784
+SKL ++ K + + D+K + E ++ F++ +E + + EEK
Sbjct: 708 LSKLHENMEKLYQSIIGYYAIDVKKVSVEDFLTDLNNFRTTFMQAIKENIKKREAEEK 765
>DAM1_HUMAN (Q9Y4D1) Disheveled associated activator of morphogenesis
1
Length = 1078
Score = 159 bits (401), Expect = 4e-38
Identities = 133/475 (28%), Positives = 215/475 (45%), Gaps = 60/475 (12%)
Query: 363 PPPPPPGP----PPPPPPRKPPAPAPRPPPPPRAG-QPPPAPPKPVKVGNQGANQGGTSD 417
PPPPPP P PPPPPP P P P P PPP PPP P + + + Q
Sbjct: 551 PPPPPPLPGGMLPPPPPPLPPGGPPPPPGPPPLGAIMPPPGAPMGLALKKKSIPQ----- 605
Query: 418 GDAPKPKLKPFFWDKVAAKPDQQMVWHEISAGSF--VFNEEKMESLFGCANQ------NR 469
P LK F W K+ + VW EI + + E +E F + N
Sbjct: 606 ---PTNALKSFNWSKLPENKLEGTVWTEIDDTKVFKILDLEDLERTFSAYQRQQDFFVNS 662
Query: 470 NERKKDSPSVDTS------VQYIQIIDPKKAQNLSILLRALNVTTDEV---IDALREGNE 520
N ++K++ ++D + V+ + +ID ++AQN +ILL L ++ DE+ I + E +
Sbjct: 663 NSKQKEADAIDDTLSSKLKVKELSVIDGRRAQNCNILLSRLKLSNDEIKRAILTMDEQED 722
Query: 521 IPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFLKVLVDIPFAFKRLETLLFMFI 580
+P ++++ LLK P + L EL ++ A+RFL + I +RL++L F
Sbjct: 723 LPKDMLEQLLKFVPEKSDIDLLEEHKHELDRMAKADRFLFEMSRINHYQQRLQSLYFKKK 782
Query: 581 LREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLL 640
E + +K + S+++ +S +LLE VL GN MN G RG A F++ +L
Sbjct: 783 FAERVAEVKPKVEAIRSGSEEVFRSGALKQLLEVVLAFGNYMNKG-QRGNAYGFKISSLN 841
Query: 641 KLSDVKGT-DGKTTLLHFVV-----------------QEIIRSEGIRAVRTERE---SRS 679
K++D K + D TLLH+++ ++I ++ + ++E RS
Sbjct: 842 KIADTKSSIDKNITLLHYLITIVENKYPSVLNLNEELRDIPQAAKVNMTELDKEISTLRS 901
Query: 680 VVSSVGTDQDVDES-PEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLG 738
+ +V T+ + +S P + + S+ Q ++ S DV+ D + AV G
Sbjct: 902 GLKAVETELEYQKSQPPQPGDKFVSVVSQFITVASFSFSDVEDLLAEAKDLFTKAVKHFG 961
Query: 739 HSLAKTQ--EFMNTDLKSLDEESEFQRCTERFVEKAREEVTWLQEEEKRIIAQVK 791
K Q EF + L SE ++ E +K EE E R+ AQ+K
Sbjct: 962 EEAGKIQPDEFFGIFDQFLQAVSEAKQENENMRKKKEEE-----ERRARMEAQLK 1011
Score = 33.1 bits (74), Expect = 3.4
Identities = 21/56 (37%), Positives = 23/56 (40%), Gaps = 4/56 (7%)
Query: 175 PSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPP 230
PSPG A P P+ P P P P P P P P + P PP PPP
Sbjct: 531 PSPG-APGGPFPSSVPGSLLPPPPPPPLPGGMLP---PPPPPLPPGGPPPPPGPPP 582
>DIA2_HUMAN (O60879) Diaphanous protein homolog 2 (Diaphanous-related
formin 2) (DRF2)
Length = 1101
Score = 153 bits (386), Expect = 2e-36
Identities = 141/552 (25%), Positives = 227/552 (40%), Gaps = 74/552 (13%)
Query: 324 KKPSNVRNWSMKAGDNNTSVVEATSSEGVG---QVPEPPNGKPPPPPPGPP--------P 372
K+ ++ + T +SS G+ P P PPPPPP PP P
Sbjct: 520 KRDEKIKELEAEIQQLRTQAQVLSSSSGIPGPPAAPPLPGVGPPPPPPAPPLPGGAPLPP 579
Query: 373 PPPPRKPPAPAPRPPPPPRA-GQPPPAPPK---PVKVGNQGANQGGTSDGDAPKPK--LK 426
PPPP P PPPPP G PPP PP P G G KP+ +K
Sbjct: 580 PPPPLPGMMGIPPPPPPPLLFGGPPPPPPLGGVPPPPGISLNLPYGMKQKKMYKPEVSMK 639
Query: 427 PFFWDKVA-AKPDQQMVWHEISAGSFVFNEEKMESLFGCANQ---NRN----ERKKDSPS 478
W K+ + + W + F + + A Q +N E KK P+
Sbjct: 640 RINWSKIEPTELSENCFWLRVKEDKFENPDLFAKLALNFATQIKVQKNAEALEEKKTGPT 699
Query: 479 VDTSVQYIQIIDPKKAQNLSILLRALNVTTDEVIDALREGNE--IPVELIQTLLKMAPTT 536
V+ ++I+DPK AQNLSI L + + +++ + + E NE + LIQ L+K P
Sbjct: 700 -KKKVKELRILDPKTAQNLSIFLGSYRMPYEDIRNVILEVNEDMLSEALIQNLVKHLPEQ 758
Query: 537 EEELKLRLFTGELSQLGPAERFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATLE 596
+ +L E L E+F V+ + RL ++LF E ++IK S +
Sbjct: 759 KILNELAELKNEYDDLCEPEQFGVVMSSVKMLQPRLSSILFKLTFEEHINNIKPSIIAVT 818
Query: 597 VASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLH 656
+A ++L+KS F +LLE VL GN MN G+ + F+++ L K+ D K D KTTLLH
Sbjct: 819 LACEELKKSESFNRLLELVLLVGNYMNSGSRNAQSLGFKINFLCKIRDTKSADQKTTLLH 878
Query: 657 FVVQEIIRSEGIRAVRTERESRSVVSSVGTDQDVDESPEASEEHYRSLGLQVVSSLSNEL 716
F+ + EE YR + EL
Sbjct: 879 FIA-----------------------------------DICEEKYRD-----ILKFPEEL 898
Query: 717 EDVKRAALIDGDALSSAVSKLGHSLAKTQEFMNTDLKSLDEESEFQRCTERFVEKAREEV 776
E V+ A+ + L S ++ + + + + ++ ++ +F F + ARE+
Sbjct: 899 EHVESASKVSAQILKSNLASMEQQIVHLERDIKKFPQAENQHDKFVEKMTSFTKTAREQY 958
Query: 777 TWLQEEEKRIIAQVKSTADYFHGNAGRDEGLRLFLIVRDF-LIILDKV-----CKEVRDK 830
L ++ ++ +YF ++ F + +F + L+ V +E+ +K
Sbjct: 959 EKLSTMHNNMMKLYENLGEYFIFDSKTVSIEEFFGDLNNFRTLFLEAVRENNKRREMEEK 1018
Query: 831 TLRAVKSSQRKE 842
T RA + ++ E
Sbjct: 1019 TRRAKLAKEKAE 1030
Score = 35.4 bits (80), Expect = 0.68
Identities = 21/59 (35%), Positives = 23/59 (38%), Gaps = 3/59 (5%)
Query: 177 PGPALESPTPNHGPTPAN-SPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQH--PPPSP 232
PGP P P GP P +P P P P P + PP PP PPP P
Sbjct: 549 PGPPAAPPLPGVGPPPPPPAPPLPGGAPLPPPPPPLPGMMGIPPPPPPPLLFGGPPPPP 607
Score = 35.0 bits (79), Expect = 0.89
Identities = 22/69 (31%), Positives = 24/69 (33%), Gaps = 5/69 (7%)
Query: 168 PSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMV----PPTSP 223
P P P PG P P P P +P P P P P P + PP P
Sbjct: 549 PGPPAAPPLPGVG-PPPPPPAPPLPGGAPLPPPPPPLPGMMGIPPPPPPPLLFGGPPPPP 607
Query: 224 PRQHPPPSP 232
P PP P
Sbjct: 608 PLGGVPPPP 616
>DAM1_MOUSE (Q8BPM0) Disheveled associated activator of morphogenesis
1
Length = 1071
Score = 152 bits (385), Expect = 3e-36
Identities = 136/495 (27%), Positives = 220/495 (43%), Gaps = 60/495 (12%)
Query: 342 SVVEATSSEGVGQVPEPPNGK----PPPPPP-----GPPPPPPPRKPPAPAPRPPPPPRA 392
+ V S G P P +G PPPPPP PPPPPP P P P PPP
Sbjct: 525 AAVPGGPSPGAPGGPFPSSGLGSLLPPPPPPLLSGGALPPPPPPLPAGGPPPPPGPPPLG 584
Query: 393 GQ-PPPAPPKPVKVGNQGANQGGTSDGDAPKPKLKPFFWDKVAAKPDQQMVWHEISAGSF 451
G PPP P + + + Q P LK F W K+ VW EI
Sbjct: 585 GVLPPPGAPVSLTLKKKNIPQ--------PTNALKSFNWSKLPENKLDGTVWTEIDDTKV 636
Query: 452 --VFNEEKMESLFGCANQNRNERKKDSPSVDTS------VQYIQIIDPKKAQNLSILLRA 503
+ + E +E F A Q + K++ ++D + V+ + +ID ++AQN +ILL
Sbjct: 637 FKILDLEDLERTFS-AYQRQQVTTKEADAIDDTLSSKLKVKELSVIDGRRAQNCNILLSR 695
Query: 504 LNVTTDEV---IDALREGNEIPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFLK 560
L ++ DE+ I + E ++P ++++ LLK P + L EL ++ A+RFL
Sbjct: 696 LKLSNDEIKRAILTMDEQEDLPKDMLEQLLKFVPEKSDIDLLEEHKHELDRMAKADRFLF 755
Query: 561 VLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGN 620
+ I +RL++L F E + +K + S+++ +SR +LLE VL GN
Sbjct: 756 EMSRINHYQQRLQSLYFKKKFAERVAEVKPKVEAIRSGSEEVFRSRALKQLLEVVLAFGN 815
Query: 621 RMNDGTYRGGAQAFRLDTLLKLSDVKGT-DGKTTLLHFVV-----------------QEI 662
MN G RG A F++ +L K++D K + D TLLH+++ ++I
Sbjct: 816 YMNKG-QRGNAYGFKISSLNKIADTKSSIDKNITLLHYLITIVENKYPKVLNLSEELRDI 874
Query: 663 IRSEGIRAVRTERE---SRSVVSSVGTDQDVDES-PEASEEHYRSLGLQVVSSLSNELED 718
++ + ++E RS + +V T+ + +S P + + S+ Q ++ S D
Sbjct: 875 PQAAKVNMTELDKEISTLRSGLKAVETELEYQKSQPPQPGDKFVSVVSQFITLASFSFSD 934
Query: 719 VKRAALIDGDALSSAVSKLGHSLAKTQ--EFMNTDLKSLDEESEFQRCTERFVEKAREEV 776
V+ + + AV G K Q EF + L +E ++ E ++ EE
Sbjct: 935 VEDLLAEAKELFTKAVKHFGEEAGKIQPDEFFGIFDQFLQAVAEAKQENENMRKRKEEE- 993
Query: 777 TWLQEEEKRIIAQVK 791
E R+ AQ+K
Sbjct: 994 ----ERRARLEAQLK 1004
>DIA3_MOUSE (Q9Z207) Diaphanous protein homolog 3
(Diaphanous-related formin 3) (DRF3) (mDIA2) (p134mDIA2)
Length = 1171
Score = 149 bits (377), Expect = 2e-35
Identities = 103/316 (32%), Positives = 151/316 (47%), Gaps = 15/316 (4%)
Query: 359 PNGKPPPPPP--GPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKPVKVGNQGANQGGTS 416
P+ PP PP G PPPPP PP P P P P G P P PP +G Q +
Sbjct: 551 PSALPPAPPALSGGVPPPPPPPPPPPPPLPGMPMPFGGPVPPPPPLGFLGGQSSIPLNLP 610
Query: 417 DGDAPKPKLKPFF------WDKVAAKP-DQQMVWHEISAGSFVFNEE--KMESLFGCAN- 466
G PK + KP W K+ + W +++ + + K+E+ F C
Sbjct: 611 FGLKPKKEFKPEISMRRLNWLKIGPNEMSENCFWIKVNENKYENRDLLCKLENTFCCQEK 670
Query: 467 QNRNERKKDSPSV-DTSVQYIQIIDPKKAQNLSILLRALNVTTDEVIDALREGNEIPVE- 524
+ RN D V ++ ++ +DPK AQNLSI L + V +++ + E +E +
Sbjct: 671 EKRNTNDFDEKKVIKKRMKELKFLDPKIAQNLSIFLSSFRVPYEKIRTMILEVDETQLSE 730
Query: 525 -LIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFLKVLVDIPFAFKRLETLLFMFILRE 583
+IQ L+K P E+ L F + + L E+F V+ ++ RL +LF E
Sbjct: 731 SMIQNLIKHLPDEEQLKSLSQFRSDYNSLCEPEQFAVVMSNVKRLRPRLSAILFKLQFEE 790
Query: 584 EASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLLKLS 643
+ ++IK + A ++++KS+ F KLLE VL GN MN G+ F L +L KL
Sbjct: 791 QVNNIKPDIMAVSTACEEIKKSKGFSKLLELVLLMGNYMNAGSRNAQTFGFDLSSLCKLK 850
Query: 644 DVKGTDGKTTLLHFVV 659
D K D KTTLLHF+V
Sbjct: 851 DTKSADQKTTLLHFLV 866
Score = 32.7 bits (73), Expect = 4.4
Identities = 20/64 (31%), Positives = 25/64 (38%), Gaps = 3/64 (4%)
Query: 190 PTPANSPSSPTPTPENSSPSSTPSPTAM---VPPTSPPRQHPPPSPEKLQHVTYAPDPSP 246
P P P+ E + P+P A+ VPP PP PPP + P P P
Sbjct: 534 PPGTKIPLQPSVEGEAGPSALPPAPPALSGGVPPPPPPPPPPPPPLPGMPMPFGGPVPPP 593
Query: 247 PDKG 250
P G
Sbjct: 594 PPLG 597
>CAPU_DROME (Q24120) Cappuccino protein
Length = 1059
Score = 135 bits (339), Expect = 6e-31
Identities = 114/426 (26%), Positives = 179/426 (41%), Gaps = 95/426 (22%)
Query: 335 KAGDNNTSVVEATSSEGVGQVPE---PPNGKPPPP------PPGPPPPPPPRKPP----- 380
K G ++T V + S++ G+ P PP PPPP PP PPPPPPP PP
Sbjct: 459 KCGQSSTKVSDNESAKEDGEKPHAVAPPPPPPPPPLHAFVAPPPPPPPPPPPPPPLANYG 518
Query: 381 APAPRPPPPPRAGQ-PPPAPPKPVKVGN--------QGANQGGTSDGDAPKPK------- 424
AP P PPPPP +G PPP PP P++ G A+ T+ AP P
Sbjct: 519 APPPPPPPPPGSGSAPPPPPPAPIEGGGGIPPPPPPMSASPSKTTISPAPLPDPAEGNWF 578
Query: 425 ----------------LKPFFWDKVA---------------------------------- 434
++P +W ++
Sbjct: 579 HRTNTMRKSAVNPPKPMRPLYWTRIVTSAPPAPRPPSVANSTDSTENSGSSPDEPPAANG 638
Query: 435 ------AKPDQQMVWHEISAGSFVFNEEKMESLFGCANQNRNERKKDSPSVDTSVQYIQI 488
A P + +W EI + N ++ LF + Q K + I++
Sbjct: 639 ADAPPTAPPATKEIWTEIEETP-LDNIDEFTELF--SRQAIAPVSKPKELKVKRAKSIKV 695
Query: 489 IDPKKAQNLSILLRALNVTTDEVIDALR--EGNEIPVELIQTLLKMAPTTEEELKLRLFT 546
+DP++++N+ I+ R+L+V + E+ A+ + + + +E +Q + + T +E +++
Sbjct: 696 LDPERSRNVGIIWRSLHVPSSEIEHAIYHIDTSVVSLEALQHMSNIQATEDELQRIKEAA 755
Query: 547 GELSQLGPAERFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSR 606
G L E+FL + I A +R+ ++F E + + T+ S +L +S
Sbjct: 756 GGDIPLDHPEQFLLDISLISMASERISCIVFQAEFEESVTLLFRKLETVSQLSQQLIESE 815
Query: 607 LFLKLLEAVLKTGNRMNDGT-YRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEII-- 663
+ +L GN MN G RG A F LD L KL DVK + TTLLHF+V+ I
Sbjct: 816 DLKLVFSIILTLGNYMNGGNRQRGQADGFNLDILGKLKDVKSKESHTTLLHFIVRTYIAQ 875
Query: 664 -RSEGI 668
R EG+
Sbjct: 876 RRKEGV 881
Score = 38.9 bits (89), Expect = 0.062
Identities = 33/120 (27%), Positives = 44/120 (36%), Gaps = 15/120 (12%)
Query: 139 TKHSRSTLGLESGHHGRNLISDPP------HNELSPSPAGTVPSPGPALESPTPNHGPTP 192
TK S + E G + PP H ++P P P P P P N+G
Sbjct: 465 TKVSDNESAKEDGEKPHAVAPPPPPPPPPLHAFVAPPPPPPPPPPPPP---PLANYG--- 518
Query: 193 ANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSP-PDKGD 251
+P P P P S + P P A + PPP T +P P P P +G+
Sbjct: 519 --APPPPPPPPPGSGSAPPPPPPAPIEGGGGIPPPPPPMSASPSKTTISPAPLPDPAEGN 576
>DAM2_HUMAN (Q86T65) Disheveled associated activator of
morphogenesis 2
Length = 1068
Score = 134 bits (338), Expect = 8e-31
Identities = 101/336 (30%), Positives = 156/336 (46%), Gaps = 29/336 (8%)
Query: 349 SEGVGQVPEPPNGK------------PPPPPPGP----PPPPPPRKPPAPAPRPP-PPPR 391
S G P PP G PPPPPP P PPPPPP PP P PP PP
Sbjct: 515 STGPVSSPPPPGGPLTLSSSMTTNDLPPPPPPLPFACCPPPPPPPLPPGGPPTPPGAPPC 574
Query: 392 AGQPPPAPPKPVKVGNQGANQGGTSDGDAPKPKLKPFFWDKVAAKPDQQMVWHEISAGSF 451
G P P P + + P LK F W K+ + VW+EI
Sbjct: 575 LGMGLPLPQDPYPSSDVPLRKKRVPQ---PSHPLKSFNWVKLNEERVPGTVWNEIDDMQV 631
Query: 452 --VFNEEKMESLFGCANQNRNE--RKKDSPSVDTSVQYIQIIDPKKAQNLSILLRALNVT 507
+ + E E +F +++ E +D V+ + +ID ++AQN ILL L ++
Sbjct: 632 FRILDLEDFEKMFSAYQRHQKELGSTEDIYLASRKVKELSVIDGRRAQNCIILLSKLKLS 691
Query: 508 TDEVIDALR---EGNEIPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFLKVLVD 564
+E+ A+ E ++ ++++ LLK P + L E+ ++ A+RFL +
Sbjct: 692 NEEIRQAILKMDEQEDLAKDMLEQLLKFIPEKSDIDLLEEHKHEIERMARADRFLYEMSR 751
Query: 565 IPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRMND 624
I +RL+ L F +E + K + +AS +L +S+ ++LE +L GN MN
Sbjct: 752 IDHYQQRLQALFFKKKFQERLAEAKPKVEAILLASRELVRSKRLRQMLEVILAIGNFMNK 811
Query: 625 GTYRGGAQAFRLDTLLKLSDVKGT-DGKTTLLHFVV 659
G RGGA FR+ +L K++D K + D +LLH+++
Sbjct: 812 G-QRGGAYGFRVASLNKIADTKSSIDRNISLLHYLI 846
Score = 35.0 bits (79), Expect = 0.89
Identities = 29/98 (29%), Positives = 35/98 (35%), Gaps = 15/98 (15%)
Query: 164 NELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSP 223
+ELS P + P PG L T + T + P P P P P P P P +P
Sbjct: 512 SELSTGPVSSPPPPGGPL---TLSSSMTTNDLPPPPPPLPFACCPPPPPPPLPPGGPPTP 568
Query: 224 PR------------QHPPPSPEKLQHVTYAPDPSPPDK 249
P Q P PS + P PS P K
Sbjct: 569 PGAPPCLGMGLPLPQDPYPSSDVPLRKKRVPQPSHPLK 606
Score = 33.9 bits (76), Expect = 2.0
Identities = 28/94 (29%), Positives = 37/94 (38%), Gaps = 6/94 (6%)
Query: 165 ELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPP 224
+LS G V SP P T + T + P P P P P P P +PP PP
Sbjct: 510 QLSELSTGPVSSPPPPGGPLTLSSSMTTNDLPPPPPPLPFACCPPPPPPP---LPPGGPP 566
Query: 225 RQHPPPSPEKL-QHVTYAPDPSPPDKGDRKQKAI 257
PP +P L + DP P ++K +
Sbjct: 567 T--PPGAPPCLGMGLPLPQDPYPSSDVPLRKKRV 598
>DAM2_MOUSE (Q80U19) Disheveled associated activator of
morphogenesis 2
Length = 1068
Score = 133 bits (334), Expect = 2e-30
Identities = 102/340 (30%), Positives = 157/340 (46%), Gaps = 34/340 (10%)
Query: 348 SSEGVGQVPEPPNGK------------PPPPPPGP----PPPPPPRKPPAPAPRPP-PPP 390
SS G P PP G PPPPPP P PPPP P PP P PP PP
Sbjct: 514 SSTGPVSSPPPPGGPLTLSSSRTTNDLPPPPPPLPFDSCPPPPAPPLPPGGPPIPPGAPP 573
Query: 391 RAGQPPPAPPKPVKVGNQGANQGGTSDGDAPKPK--LKPFFWDKVAAKPDQQMVWHEISA 448
PP P +N+ P+P LK F W K+ + VW+EI
Sbjct: 574 CFSSGPPPSHDPFS-----SNEAPLRKKRIPQPSHPLKSFNWVKLNEERVSGTVWNEIDD 628
Query: 449 GSF--VFNEEKMESLFGCANQNRNERKKDSPSV---DTSVQYIQIIDPKKAQNLSILLRA 503
+ + E E +F +++ + + + V+ + +ID ++AQN ILL
Sbjct: 629 SQVFRILDLEDFEKMFSAYQRHQQKELGSTEDIYITSRKVKELSVIDGRRAQNCIILLSK 688
Query: 504 LNVTTDEVIDALR---EGNEIPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFLK 560
L ++ DE+ A+ E ++ ++++ LLK P + L E+ ++ A+RFL
Sbjct: 689 LKLSNDEIRQAILRMDEQEDLAKDMLEQLLKFIPEKSDIDLLEEHKHEIERMARADRFLY 748
Query: 561 VLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGN 620
+ I +RL+ L F +E + K + +AS +L S+ ++LE VL GN
Sbjct: 749 EMSRIDHYQQRLQALFFKKKFQERLAEAKPKVEAILLASRELTLSQRLKQMLEVVLAIGN 808
Query: 621 RMNDGTYRGGAQAFRLDTLLKLSDVKGT-DGKTTLLHFVV 659
MN G RGGA FR+ +L K++D K + D +LLH+++
Sbjct: 809 FMNKG-QRGGAYGFRVASLNKIADTKSSIDRNISLLHYLI 847
Score = 35.8 bits (81), Expect = 0.52
Identities = 31/102 (30%), Positives = 36/102 (34%), Gaps = 21/102 (20%)
Query: 163 HNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTS 222
HNE S P + P PG L T + T + P P P P +S P P P +PP
Sbjct: 511 HNESSTGPVSSPPPPGGPL---TLSSSRTTNDLPPPPPPLPFDSCP---PPPAPPLPPGG 564
Query: 223 PP---------------RQHPPPSPEKLQHVTYAPDPSPPDK 249
PP P S E P PS P K
Sbjct: 565 PPIPPGAPPCFSSGPPPSHDPFSSNEAPLRKKRIPQPSHPLK 606
>DIA_DROME (P48608) Diaphanous protein
Length = 1091
Score = 132 bits (333), Expect = 3e-30
Identities = 123/537 (22%), Positives = 218/537 (39%), Gaps = 80/537 (14%)
Query: 356 PEPPNG---KPPPPPP-------GPPPPPPPRKPP---APAPRPPPPPRAGQPPPAPPKP 402
P PP G PPPPPP GPPPPPPP P P P PPPP G PPP P
Sbjct: 515 PPPPGGGGAPPPPPPPMPGRAGGGPPPPPPPPMPGRAGGPPPPPPPPGMGGPPPPPMPGM 574
Query: 403 VKVGN--------QGANQGGTSDGDAPKPK------LKPFFWDKVA-AKPDQQMVWHEIS 447
++ G G G PK K +K W + AK + W +
Sbjct: 575 MRPGGGPPPPPMMMGPMVPVLPHGLKPKKKWDVKNPMKRANWKAIVPAKMSDKAFWVKCQ 634
Query: 448 AGSFVFNEEKMESLFGCANQNRNERKKDSPSVDTSVQY----IQIIDPKKAQNLSILLRA 503
++ E +++ + +KD+ T++ ++++D K AQNL+I+L
Sbjct: 635 EDKLAQDDFLAELAVKFSSKPVKKEQKDAVDKPTTLTKKNVDLRVLDSKTAQNLAIMLGG 694
Query: 504 L--NVTTDEV-IDALREGNEI-PVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFL 559
+++ +++ I LR +I ++Q L++ P E +L+ + L P E+F
Sbjct: 695 SLKHLSYEQIKICLLRCDTDILSSNILQQLIQYLPPPEHLKRLQEIKAKGEPLPPIEQFA 754
Query: 560 KVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTG 619
+ +I RL L F + IK A +++R S+ F K+LE +L G
Sbjct: 755 ATIGEIKRLSPRLHNLNFKLTYADMVQDIKPDIVAGTAACEEIRNSKKFSKILELILLLG 814
Query: 620 NRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERESRS 679
N MN G+ A F + L KLS+ K D K TLLH++ + +
Sbjct: 815 NYMNSGSKNEAAFGFEISYLTKLSNTKDADNKQTLLHYLADLVEK--------------- 859
Query: 680 VVSSVGTDQDVDESPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLGH 739
+ P+A + ++L V +A+ ++ DA+ A+ ++
Sbjct: 860 ------------KFPDA-------------LNFYDDLSHVNKASRVNMDAIQKAMRQMNS 894
Query: 740 SLAKTQEFMNTDLKSLDEESEFQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADYFHG 799
++ + + + ++ +F +F E+ R++V L + + ++ K ++Y+
Sbjct: 895 AVKNLETDLQNNKVPQCDDDKFSEVMGKFAEECRQQVDVLGKMQLQMEKLYKDLSEYYAF 954
Query: 800 NAGRDEGLRLFLIVRDFLIILDKVCKEVRDKTLRAVKSSQRKEALSVPSSPDTRHQQ 856
+ + F ++ F + + +R + ++K L R QQ
Sbjct: 955 DPSKYTMEEFFADIKTF----KDAFQAAHNDNVRVREELEKKRRLQEAREQSAREQQ 1007
Score = 36.2 bits (82), Expect = 0.40
Identities = 23/89 (25%), Positives = 30/89 (32%)
Query: 156 NLISDPPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPT 215
N ++ P N+L P P +P P P P + P P P P P
Sbjct: 496 NGVAAPSPNKLPKVNIPMPPPPPGGGGAPPPPPPPMPGRAGGGPPPPPPPPMPGRAGGPP 555
Query: 216 AMVPPTSPPRQHPPPSPEKLQHVTYAPDP 244
PP PPP P ++ P P
Sbjct: 556 PPPPPPGMGGPPPPPMPGMMRPGGGPPPP 584
Score = 35.8 bits (81), Expect = 0.52
Identities = 22/68 (32%), Positives = 26/68 (37%), Gaps = 5/68 (7%)
Query: 183 SPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAP 242
+P+PN P N P P P +P P P PP PPP P + P
Sbjct: 500 APSPNKLPK-VNIPMPPPPPGGGGAPPPPPPPMPGRAGGGPPPPPPPPMPGRAG----GP 554
Query: 243 DPSPPDKG 250
P PP G
Sbjct: 555 PPPPPPPG 562
Score = 33.1 bits (74), Expect = 3.4
Identities = 24/72 (33%), Positives = 24/72 (33%), Gaps = 11/72 (15%)
Query: 161 PPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPP 220
PP P AG P P P P P A P P P P P P P M P
Sbjct: 525 PPPPPPMPGRAGGGPPP------PPPPPMPGRAGGPPPPPPPPGMGGPPPPPMPGMMRPG 578
Query: 221 TSPPRQHPPPSP 232
PPP P
Sbjct: 579 GG-----PPPPP 585
Score = 33.1 bits (74), Expect = 3.4
Identities = 24/81 (29%), Positives = 29/81 (35%), Gaps = 8/81 (9%)
Query: 167 SPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQ 226
+PSP +P + P P G P P P P P + P P PP P R
Sbjct: 500 APSP-NKLPKVNIPMPPPPPGGGGAP---PPPPPPMPGRAGGGPPPPP----PPPMPGRA 551
Query: 227 HPPPSPEKLQHVTYAPDPSPP 247
PP P + P P P
Sbjct: 552 GGPPPPPPPPGMGGPPPPPMP 572
>FM14_MOUSE (Q05859) Formin 1 isoform IV (Limb deformity protein)
Length = 1206
Score = 122 bits (307), Expect = 3e-27
Identities = 124/531 (23%), Positives = 209/531 (39%), Gaps = 80/531 (15%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPP-------PRAGQPPPAPPKP------ 402
P P PPPPPP PP P PP P P P PPPP P +G PPP PP P
Sbjct: 679 PVCPVSPPPPPPPPPPTPVPPSDGPPPPPPPPPPLPNVLALPNSGGPPPPPPPPPPPPGL 738
Query: 403 -------VKVG-NQGANQGGTSDGDAPKPKLKPFFWDKVAAKPDQQ----MVWHEISAGS 450
+ G + ++Q P +KP +W ++ Q +W +
Sbjct: 739 APPPPPGLSFGLSSSSSQYPRKPAIEPSCPMKPLYWTRIQINDKSQDAAPTLWDSLEE-P 797
Query: 451 FVFNEEKMESLFG--CANQNRNERKKDSPSVDTSVQYIQIIDPKKAQNLSILLRALNVTT 508
+ + + E LF Q + + + + I+++D K++Q + IL+ +L++
Sbjct: 798 HIRDTSEFEYLFSKDTTQQKKKPLSEAYEKKNKVKKIIKLLDGKRSQTVGILISSLHLEM 857
Query: 509 DEVIDAL--REGNEIPVELIQTLLKMAPTTEEELKLRLF-----TGELSQLGPAERFLKV 561
++ A+ + + + +E + L + +E K+R + +L L E+FL
Sbjct: 858 KDIQQAIFTVDDSVVDLETLAALYENRAQEDELTKIRKYYETSKEEDLKLLDKPEQFLHE 917
Query: 562 LVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNR 621
L IP +R + ++F + E +S+ + AS L + +L +L GN
Sbjct: 918 LAQIPNFAERAQCIIFRAVFSEGITSLHRKVEIVTRASKGLLHMKSVKDILALILAFGNY 977
Query: 622 MNDGT-YRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERESRSV 680
MN G RG A + L+ L KL DVK D L+ +VV+ +R A
Sbjct: 978 MNGGNRTRGQADGYSLEILPKLKDVKSRDNGMNLVDYVVKYYLRYYDQEA---------- 1027
Query: 681 VSSVGTDQDVDESPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLGHS 740
GTD+ V PE +D A+ + + L + KL
Sbjct: 1028 ----GTDKSVFPLPEP--------------------QDFFLASQVKFEDLLKDLRKLKRQ 1063
Query: 741 LAKTQEFMNTDLKSLDEE--SEFQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADYF- 797
L +++ M K E F+ E F +KA++E + + ++T YF
Sbjct: 1064 LEASEQQMKLVCKESPREYLQPFKDKLEEFFKKAKKEHKMEESHLENAQKSFETTVGYFG 1123
Query: 798 -HGNAGRDEGLRLFLIV------RDFLIILDKVCKEVRDKTLRAVKSSQRK 841
G E ++ + DF I + K + + L+ ++S K
Sbjct: 1124 MKPKTGEKEVTPSYVFMVWFEFCSDFKTIWKRESKNISKERLKMAQASVSK 1174
Score = 47.0 bits (110), Expect = 2e-04
Identities = 31/87 (35%), Positives = 34/87 (38%), Gaps = 2/87 (2%)
Query: 161 PPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPP 220
PP P P P P P P P P P P SP P P P TP P + PP
Sbjct: 647 PPTPPPLPPPLIPPPPPLPPGLGPLPPAPPIPPVCPVSPPPPPP--PPPPTPVPPSDGPP 704
Query: 221 TSPPRQHPPPSPEKLQHVTYAPDPSPP 247
PP P P+ L + P P PP
Sbjct: 705 PPPPPPPPLPNVLALPNSGGPPPPPPP 731
Score = 42.7 bits (99), Expect = 0.004
Identities = 33/97 (34%), Positives = 35/97 (36%), Gaps = 16/97 (16%)
Query: 157 LISDPPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTA 216
LI PP P P G P P P P P P PTP P + P P P
Sbjct: 657 LIPPPP-----PLPPGLGPLPPAPPIPPVCPVSPPPPPPPPPPTPVPPSDGPPPPPPPPP 711
Query: 217 MVP-----PTS--PPRQHPPPSPEKLQHVTYAPDPSP 246
+P P S PP PPP P AP P P
Sbjct: 712 PLPNVLALPNSGGPPPPPPPPPPPP----GLAPPPPP 744
Score = 42.4 bits (98), Expect = 0.006
Identities = 44/143 (30%), Positives = 57/143 (39%), Gaps = 10/143 (6%)
Query: 112 KVLDCIRRSTPPAPPVSQKKTGSRYLLTKHSRSTLGLESGHHGRNLISDPPHNELSPSPA 171
K D ++T + V +K T S L+ S+ + + + R S ++SP PA
Sbjct: 589 KPCDAESKATRSSQIVPKKLTISLTQLSP-SKDSKDIHAPFQTREGTSSSSQQKISP-PA 646
Query: 172 GTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTS-PPRQH--- 227
P P P P P P P P P P P S P P PPT PP
Sbjct: 647 PPTPPPLPPPLIPPPPPLP-PGLGPLPPAPPIPPVCPVSPPPPPPPPPPTPVPPSDGPPP 705
Query: 228 PPPSPEKLQHVTYAPD---PSPP 247
PPP P L +V P+ P PP
Sbjct: 706 PPPPPPPLPNVLALPNSGGPPPP 728
Score = 38.9 bits (89), Expect = 0.062
Identities = 25/75 (33%), Positives = 30/75 (39%), Gaps = 6/75 (8%)
Query: 161 PPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPP 220
PP +SP P P P P P+ GP P P P P P + ++ P PP
Sbjct: 678 PPVCPVSPPPPPPPPPPTPV----PPSDGPPPP--PPPPPPLPNVLALPNSGGPPPPPPP 731
Query: 221 TSPPRQHPPPSPEKL 235
PP PP P L
Sbjct: 732 PPPPPGLAPPPPPGL 746
Score = 35.8 bits (81), Expect = 0.52
Identities = 26/79 (32%), Positives = 27/79 (33%), Gaps = 9/79 (11%)
Query: 172 GTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPS 231
GT S + P P PTP P P P P P P A P P PPP
Sbjct: 633 GTSSSSQQKISPPAP---PTPPPLPPPLIPPPPPLPPGLGPLPPAPPIPPVCPVSPPPPP 689
Query: 232 PEKLQHVTYAPDPSPPDKG 250
P P P PP G
Sbjct: 690 PPP------PPTPVPPSDG 702
>FMN1_MOUSE (Q05860) Formin 1 isoforms I/II/III (Limb deformity
protein)
Length = 1468
Score = 120 bits (300), Expect = 2e-26
Identities = 128/553 (23%), Positives = 216/553 (38%), Gaps = 88/553 (15%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPP-------PRAGQPPPAPPKP------ 402
P P PPPPPP PP P PP P P P PPPP P +G PPP PP P
Sbjct: 905 PVCPVSPPPPPPPPPPTPVPPSDGPPPPPPPPPPLPNVLALPNSGGPPPPPPPPPPPPGL 964
Query: 403 -------VKVG-NQGANQGGTSDGDAPKPKLKPFFWDKVAAKPDQQ----MVWHEISAGS 450
+ G + ++Q P +KP +W ++ Q +W +
Sbjct: 965 APPPPPGLSFGLSSSSSQYPRKPAIEPSCPMKPLYWTRIQINDKSQDAAPTLWDSLEE-P 1023
Query: 451 FVFNEEKMESLFG--CANQNRNERKKDSPSVDTSVQYIQIIDPKKAQNLSILLRALNVTT 508
+ + + E LF Q + + + + I+++D K++Q + IL+ +L++
Sbjct: 1024 HIRDTSEFEYLFSKDTTQQKKKPLSEAYEKKNKVKKIIKLLDGKRSQTVGILISSLHLEM 1083
Query: 509 DEVIDAL--REGNEIPVELIQTLLKMAPTTEEELKLRLF-----TGELSQLGPAERFLKV 561
++ A+ + + + +E + L + +E K+R + +L L E+FL
Sbjct: 1084 KDIQQAIFTVDDSVVDLETLAALYENRAQEDELTKIRKYYETSKEEDLKLLDKPEQFLHE 1143
Query: 562 LVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNR 621
L IP +R + ++F + E +S+ + AS L + +L +L GN
Sbjct: 1144 LAQIPNFAERAQCIIFRAVFSEGITSLHRKVEIVTRASKGLLHMKSVKDILALILAFGNY 1203
Query: 622 MNDGT-YRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIR----------SEGIR- 669
MN G RG A + L+ L KL DVK D L+ +VV+ +R R
Sbjct: 1204 MNGGNRTRGQADGYSLEILPKLKDVKSRDNGMNLVDYVVKYYLRYYDQCKHHDQEASCRG 1263
Query: 670 ----------AVRTERES-RSVVSSVGTDQDVDESPEASEEHYRSLGLQVVSSLSNELED 718
AV +R+S + GTD+ V PE +D
Sbjct: 1264 KDLFSLYFHIAVHPQRKSGLELKQEAGTDKSVFPLPEP--------------------QD 1303
Query: 719 VKRAALIDGDALSSAVSKLGHSLAKTQEFMNTDLKSLDEE--SEFQRCTERFVEKAREEV 776
A+ + + L + KL L +++ M K E F+ E F +KA++E
Sbjct: 1304 FFLASQVKFEDLLKDLRKLKRQLEASEQQMKLVCKESPREYLQPFKDKLEEFFKKAKKEH 1363
Query: 777 TWLQEEEKRIIAQVKSTADYF--HGNAGRDEGLRLFLIV------RDFLIILDKVCKEVR 828
+ + ++T YF G E ++ + DF I + K +
Sbjct: 1364 KMEESHLENAQKSFETTVGYFGMKPKTGEKEVTPSYVFMVWFEFCSDFKTIWKRESKNIS 1423
Query: 829 DKTLRAVKSSQRK 841
+ L+ ++S K
Sbjct: 1424 KERLKMAQASVSK 1436
Score = 47.0 bits (110), Expect = 2e-04
Identities = 31/87 (35%), Positives = 34/87 (38%), Gaps = 2/87 (2%)
Query: 161 PPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPP 220
PP P P P P P P P P P P SP P P P TP P + PP
Sbjct: 873 PPTPPPLPPPLIPPPPPLPPGLGPLPPAPPIPPVCPVSPPPPPP--PPPPTPVPPSDGPP 930
Query: 221 TSPPRQHPPPSPEKLQHVTYAPDPSPP 247
PP P P+ L + P P PP
Sbjct: 931 PPPPPPPPLPNVLALPNSGGPPPPPPP 957
Score = 42.7 bits (99), Expect = 0.004
Identities = 33/97 (34%), Positives = 35/97 (36%), Gaps = 16/97 (16%)
Query: 157 LISDPPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTA 216
LI PP P P G P P P P P P PTP P + P P P
Sbjct: 883 LIPPPP-----PLPPGLGPLPPAPPIPPVCPVSPPPPPPPPPPTPVPPSDGPPPPPPPPP 937
Query: 217 MVP-----PTS--PPRQHPPPSPEKLQHVTYAPDPSP 246
+P P S PP PPP P AP P P
Sbjct: 938 PLPNVLALPNSGGPPPPPPPPPPPP----GLAPPPPP 970
Score = 38.9 bits (89), Expect = 0.062
Identities = 25/75 (33%), Positives = 30/75 (39%), Gaps = 6/75 (8%)
Query: 161 PPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPP 220
PP +SP P P P P P+ GP P P P P P + ++ P PP
Sbjct: 904 PPVCPVSPPPPPPPPPPTPV----PPSDGPPPP--PPPPPPLPNVLALPNSGGPPPPPPP 957
Query: 221 TSPPRQHPPPSPEKL 235
PP PP P L
Sbjct: 958 PPPPPGLAPPPPPGL 972
Score = 38.5 bits (88), Expect = 0.080
Identities = 43/143 (30%), Positives = 56/143 (39%), Gaps = 10/143 (6%)
Query: 112 KVLDCIRRSTPPAPPVSQKKTGSRYLLTKHSRSTLGLESGHHGRNLISDPPHNELSPSPA 171
K D ++T + V +K T S L+ S+ + + + R S ++SP PA
Sbjct: 815 KPCDAESKATRSSQIVPKKLTISLTQLSP-SKDSKDIHAPFQTREGTSSSSQQKISP-PA 872
Query: 172 GTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTS--PPR--QH 227
P P P P P P P P P P P S P P PPT P
Sbjct: 873 PPTPPPLPPPLIPPPPPLP-PGLGPLPPAPPIPPVCPVSPPPPPPPPPPTPVPPSDGPPP 931
Query: 228 PPPSPEKLQHVTYAPD---PSPP 247
PPP P L +V P+ P PP
Sbjct: 932 PPPPPPPLPNVLALPNSGGPPPP 954
Score = 35.8 bits (81), Expect = 0.52
Identities = 26/79 (32%), Positives = 27/79 (33%), Gaps = 9/79 (11%)
Query: 172 GTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPS 231
GT S + P P PTP P P P P P P A P P PPP
Sbjct: 859 GTSSSSQQKISPPAP---PTPPPLPPPLIPPPPPLPPGLGPLPPAPPIPPVCPVSPPPPP 915
Query: 232 PEKLQHVTYAPDPSPPDKG 250
P P P PP G
Sbjct: 916 PPP------PPTPVPPSDG 928
>FMN2_MOUSE (Q9JL04) Formin 2
Length = 1567
Score = 119 bits (299), Expect = 3e-26
Identities = 106/376 (28%), Positives = 164/376 (43%), Gaps = 63/376 (16%)
Query: 351 GVGQVPEPP---NGKPPPPP-PGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP---- 402
GVG P PP G PPPPP PG PPPP P P PPP P G PPPAPP P
Sbjct: 1017 GVGIPPPPPLPGMGIPPPPPLPGSGIPPPPALPGVAIPPPPPLPGMGVPPPAPPPPGAGI 1076
Query: 403 --------------VKVGNQ-----------------------GANQGGTSDGDAPKP-- 423
+VG+ G NQ + +P
Sbjct: 1077 PPPPLLPGSGPPHSSQVGSSTLPAAPQGCGFLFPPLPTGLFGLGMNQDRVARKQPIEPCR 1136
Query: 424 KLKPFFWDKVA--AKPDQ--QMVWHEISAGSFVFNEEKMESLFG-CANQNRNERKKDSPS 478
+KP +W ++ +K D ++W +I S +E E LF A + R + D+ S
Sbjct: 1137 PMKPLYWTRIQLHSKRDSSPSLIWEKIEEPSIDCHE--FEELFSKTAVKERKKPISDTIS 1194
Query: 479 VDTSVQYIQIIDPKKAQNLSILLRALNVTTDEVIDAL--REGNEIPVELIQTLLKMAPTT 536
+ Q ++++ K++Q + IL+ +L++ ++ A+ + + + +E +Q L + +
Sbjct: 1195 KTKAKQVVKLLSNKRSQAVGILMSSLHLDMKDIQHAVVNLDNSVVDLETLQALYENRAQS 1254
Query: 537 EEELKLRLFT------GELSQLGPAERFLKVLVDIPFAFKRLETLLFMFILREEASSIKE 590
+E K+ + L E+FL L IP +R+ +LF E SI+
Sbjct: 1255 DELEKIEKHSRSSKDKENAKSLDKPEQFLYELSLIPNFSERVFCILFQSTFSESICSIRR 1314
Query: 591 SFATLEVASDKLRKSRLFLKLLEAVLKTGNRMNDGT-YRGGAQAFRLDTLLKLSDVKGTD 649
L+ + L+ +++L VL GN MN G RG A F LD L KL DVK +D
Sbjct: 1315 KLELLQKLCETLKNGPGVMQVLGLVLAFGNYMNAGNKTRGQADGFGLDILPKLKDVKSSD 1374
Query: 650 GKTTLLHFVVQEIIRS 665
+LL ++V +R+
Sbjct: 1375 NSRSLLSYIVSYYLRN 1390
Score = 55.1 bits (131), Expect = 8e-07
Identities = 31/56 (55%), Positives = 31/56 (55%), Gaps = 5/56 (8%)
Query: 351 GVGQVPEPPN---GKPPPPP-PGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
GVG P PP G PPPPP PG PPPP P P PPP P G PPP PP P
Sbjct: 973 GVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPP-PPLP 1027
Score = 55.1 bits (131), Expect = 8e-07
Identities = 31/56 (55%), Positives = 31/56 (55%), Gaps = 5/56 (8%)
Query: 351 GVGQVPEPPN---GKPPPPP-PGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
GVG P PP G PPPPP PG PPPP P P PPP P G PPP PP P
Sbjct: 951 GVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPP-PPLP 1005
Score = 54.7 bits (130), Expect = 1e-06
Identities = 31/56 (55%), Positives = 31/56 (55%), Gaps = 5/56 (8%)
Query: 351 GVGQVPEPPN---GKPPPPP-PGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
GVG P PP G PPPPP PG PPPP P P PPP P G PPP PP P
Sbjct: 984 GVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGMGIPPP-PPLP 1038
Score = 54.3 bits (129), Expect = 1e-06
Identities = 30/56 (53%), Positives = 31/56 (54%), Gaps = 5/56 (8%)
Query: 351 GVGQVPEPP---NGKPPPPP-PGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
G+G P PP G PPPPP PG PPPP P P PPP P G PPP PP P
Sbjct: 929 GMGIPPPPPLPGMGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPP-PPLP 983
Score = 53.1 bits (126), Expect = 3e-06
Identities = 30/59 (50%), Positives = 31/59 (51%), Gaps = 8/59 (13%)
Query: 351 GVGQVPEPPN------GKPPPPP-PGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
G+G P PP G PPPPP PG PPPP P P PPP P G PPP PP P
Sbjct: 904 GLGVPPPPPAPPLPGMGIPPPPPLPGMGIPPPPPLPGMGIPPPPPLPGVGIPPP-PPLP 961
Score = 52.8 bits (125), Expect = 4e-06
Identities = 28/62 (45%), Positives = 34/62 (54%), Gaps = 7/62 (11%)
Query: 344 VEATSSEGVGQVPEPPNGKPPPPPPGPPPPPPPRKPPAPA---PRPPPPPRAGQPPPAPP 400
++ ++ +G +P PP PPP PG PPPP PP P P PPP P G PPP PP
Sbjct: 882 LQFSNLQGPEMLPAPPQ---PPPLPGLGVPPPPPAPPLPGMGIPPPPPLPGMGIPPP-PP 937
Query: 401 KP 402
P
Sbjct: 938 LP 939
Score = 40.4 bits (93), Expect = 0.021
Identities = 27/90 (30%), Positives = 29/90 (32%)
Query: 161 PPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPP 220
PP P P +P P P P P P P P P P P P +PP
Sbjct: 908 PPPPPAPPLPGMGIPPPPPLPGMGIPPPPPLPGMGIPPPPPLPGVGIPPPPPLPGVGIPP 967
Query: 221 TSPPRQHPPPSPEKLQHVTYAPDPSPPDKG 250
P P P L V P P P G
Sbjct: 968 PPPLPGVGIPPPPPLPGVGIPPPPPLPGVG 997
Score = 40.0 bits (92), Expect = 0.028
Identities = 32/104 (30%), Positives = 36/104 (33%), Gaps = 5/104 (4%)
Query: 150 SGHHGRNLISDPPHNELSPSPAGTVPSPGPALESP---TPNHGPTPANSPSSPTPTPENS 206
S G ++ PP + P P VP P PA P P P P P P P
Sbjct: 885 SNLQGPEMLPAPP--QPPPLPGLGVPPPPPAPPLPGMGIPPPPPLPGMGIPPPPPLPGMG 942
Query: 207 SPSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKG 250
P P P +PP P P P L V P P P G
Sbjct: 943 IPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVG 986
Score = 38.5 bits (88), Expect = 0.080
Identities = 25/83 (30%), Positives = 27/83 (32%)
Query: 168 PSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQH 227
P P +P P P P P P P P P P P P +PP P
Sbjct: 981 PLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGMGIPPPPPLPGS 1040
Query: 228 PPPSPEKLQHVTYAPDPSPPDKG 250
P P L V P P P G
Sbjct: 1041 GIPPPPALPGVAIPPPPPLPGMG 1063
Score = 37.7 bits (86), Expect = 0.14
Identities = 23/57 (40%), Positives = 25/57 (43%), Gaps = 17/57 (29%)
Query: 363 PPPP-------PPGPPPPPP---------PRKPPAPAPRPPPPPRAGQPPPAPPKPV 403
PPPP P P P P P PAP P+PPP P G PPP P P+
Sbjct: 861 PPPPCLSDITVPALPSPTAPALQFSNLQGPEMLPAP-PQPPPLPGLGVPPPPPAPPL 916
Score = 37.4 bits (85), Expect = 0.18
Identities = 25/83 (30%), Positives = 27/83 (32%)
Query: 168 PSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQH 227
P P +P P P P P P P P P P P P +PP P
Sbjct: 948 PLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGV 1007
Query: 228 PPPSPEKLQHVTYAPDPSPPDKG 250
P P L V P P P G
Sbjct: 1008 GIPPPPPLPGVGIPPPPPLPGMG 1030
Score = 37.4 bits (85), Expect = 0.18
Identities = 25/83 (30%), Positives = 27/83 (32%)
Query: 168 PSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQH 227
P P +P P P P P P P P P P P P +PP P
Sbjct: 926 PLPGMGIPPPPPLPGMGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGV 985
Query: 228 PPPSPEKLQHVTYAPDPSPPDKG 250
P P L V P P P G
Sbjct: 986 GIPPPPPLPGVGIPPPPPLPGVG 1008
Score = 37.0 bits (84), Expect = 0.23
Identities = 25/83 (30%), Positives = 27/83 (32%)
Query: 168 PSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQH 227
P P +P P P P P P P P P P P P +PP P
Sbjct: 937 PLPGMGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGV 996
Query: 228 PPPSPEKLQHVTYAPDPSPPDKG 250
P P L V P P P G
Sbjct: 997 GIPPPPPLPGVGIPPPPPLPGVG 1019
Score = 36.6 bits (83), Expect = 0.31
Identities = 24/83 (28%), Positives = 27/83 (31%)
Query: 168 PSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQH 227
P P +P P P P P P P P P P P P +PP P
Sbjct: 959 PLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGVGIPPPPPLPGV 1018
Query: 228 PPPSPEKLQHVTYAPDPSPPDKG 250
P P L + P P P G
Sbjct: 1019 GIPPPPPLPGMGIPPPPPLPGSG 1041
Score = 35.4 bits (80), Expect = 0.68
Identities = 46/159 (28%), Positives = 53/159 (32%), Gaps = 32/159 (20%)
Query: 122 PPAPPV----SQKKTGSRYLLTKHSRSTLGLESGHHGRNLISDP----PHNELS---PSP 170
PP PP+ SQ + L T+ S S G N P P E S P
Sbjct: 797 PPPPPLLGSDSQGQPSQPSLHTESETSHEHSVSSSFGNNCNVPPAPPLPCTESSSFMPGL 856
Query: 171 AGTVPSPG-------PALESPTPN-------HGPTPANSPSSPTPTPENSSPSSTPSPT- 215
+P P PAL SPT GP +P P P P P P+P
Sbjct: 857 GMAIPPPPCLSDITVPALPSPTAPALQFSNLQGPEMLPAPPQPPPLPGLGVPPPPPAPPL 916
Query: 216 ----AMVPPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKG 250
PP P PPP P L + P P P G
Sbjct: 917 PGMGIPPPPPLPGMGIPPPPP--LPGMGIPPPPPLPGVG 953
Score = 35.4 bits (80), Expect = 0.68
Identities = 23/86 (26%), Positives = 27/86 (30%), Gaps = 6/86 (6%)
Query: 168 PSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSP---- 223
P P +P P P P P P P P P + P P +PP P
Sbjct: 1003 PLPGVGIPPPPPLPGVGIPPPPPLPGMGIPPPPPLPGSGIPPPPALPGVAIPPPPPLPGM 1062
Query: 224 --PRQHPPPSPEKLQHVTYAPDPSPP 247
P PPP + P PP
Sbjct: 1063 GVPPPAPPPPGAGIPPPPLLPGSGPP 1088
>FMN_CHICK (Q05858) Formin (Limb deformity protein)
Length = 1213
Score = 117 bits (292), Expect = 2e-25
Identities = 118/506 (23%), Positives = 203/506 (39%), Gaps = 105/506 (20%)
Query: 339 NNTSVVEATSSEGVGQVPEPPNGKPPPPPPGPPP----------PPPPRKPP-------- 380
++ SV+ + + P PP PPPPPP PPP PPPP P
Sbjct: 638 SSESVLSCQPKQMLPPSPPPPPPPPPPPPPPPPPFSDSSLPGLVPPPPPLPTGPTSVTPH 697
Query: 381 -------------------APAPRPPPP------------PRAGQPPPAPPKPVKV---G 406
APAP PPP P +G PPP PP +
Sbjct: 698 FAFGPPLPPQLSEGCRDFQAPAPPAPPPLPGLGPPVPPPLPGSGLPPPPPPPGPGLFFNS 757
Query: 407 NQGANQGGTSDGDAPKPKLKPFFWDKVAAKPDQQ----MVWHEISAGSFVFNEEKMESLF 462
++QG P +KP +W ++ + ++ +W + + + + E LF
Sbjct: 758 TLSSSQGPRKPAIEPSRPMKPLYWTRIQLQGSRKTAIPTLWESLEEPD-ILDTTEFEYLF 816
Query: 463 GCANQNRNERKKDSPSVDTSV---QYIQIIDPKKAQNLSILLRALNVTTDEVIDALR--E 517
+ + +RK S + + + I+++D K++Q + IL+ +L++ ++ A+ +
Sbjct: 817 S-KDTTQEKRKPLSETYEKKTKAKKIIKLLDGKRSQTVGILISSLHLEMKDIQQAILCVD 875
Query: 518 GNEIPVELIQTLLKMAPTTEEELKLRLF-----TGELSQLGPAERFLKVLVDIPFAFKRL 572
+ + +E ++ L + +E K+ + EL L E+FL L IP +R
Sbjct: 876 DSVVDLETLEALYENRAQKDELEKIEQYYQTSKEEELKLLDKPEQFLYELSQIPNFTERA 935
Query: 573 ETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRMNDGT-YRGGA 631
+ ++F + E +S+ + S L ++L +L GN MN G RG A
Sbjct: 936 QCIIFQSVFSEGITSVHRKVDIITRVSKALLNMTSVKEILGLILAFGNYMNGGNRTRGQA 995
Query: 632 QAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERESRSVVSSVGTDQDVD 691
F L+ L KL DVK D + L+ +VV +R ++E+ GTD+ +
Sbjct: 996 DGFGLEILPKLKDVKSRDNRINLVDYVVIYYLR-------HCDKEA-------GTDKSIF 1041
Query: 692 ESPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLGHSLAKTQEFMNTD 751
PE +D +A+ + + L + KL L +++ M
Sbjct: 1042 PLPEP--------------------QDFFQASQVKFEDLIKDLRKLKRDLEASEKQMKLV 1081
Query: 752 LKSLDEE--SEFQRCTERFVEKAREE 775
+ EE F+ E F +KA+EE
Sbjct: 1082 CRESSEEHLQPFKEKLEEFFQKAKEE 1107
Score = 35.8 bits (81), Expect = 0.52
Identities = 60/243 (24%), Positives = 86/243 (34%), Gaps = 61/243 (25%)
Query: 48 FHVHGK------QVENVGKHCRKELVEKSNDDVKDFDLHLLERGSDLRCIHLL--ENEIP 99
FH+ G+ Q+E H + EL K N R + R I + ++ +P
Sbjct: 531 FHIRGEHAVSTAQLEETIAHLKNELDNKLN-----------RRNEEARDIGVSTEDDNLP 579
Query: 100 KTTKSLSPVMRKKVLDCIR--RST--PPAPPVSQKKTGSRYLLTKHSRSTLGLESGHHGR 155
KT +++ CI+ R T P+ ++ ++ + K + S+L G
Sbjct: 580 KTYRNV----------CIQTDRETFIKPSEEENRAVKNNQIVPKKLNISSLTHSISTQGE 629
Query: 156 NLIS-DPPHNE--LSPSPAGTVPSPGPALESPTPNHGPTPANSPSS------PTPTPENS 206
N S D P +E LS P +P P P P P P S P P P +
Sbjct: 630 NKDSYDVPSSESVLSCQPKQMLPPSPPPPPPPPPPPPPPPPPFSDSSLPGLVPPPPPLPT 689
Query: 207 SPSSTPSPTAMVPP--------------TSPPRQHP-----PPSPEKLQHVTYAPDPSPP 247
P+S A PP +PP P PP P L P P PP
Sbjct: 690 GPTSVTPHFAFGPPLPPQLSEGCRDFQAPAPPAPPPLPGLGPPVPPPLPGSGLPPPPPPP 749
Query: 248 DKG 250
G
Sbjct: 750 GPG 752
Score = 33.1 bits (74), Expect = 3.4
Identities = 31/103 (30%), Positives = 38/103 (36%), Gaps = 22/103 (21%)
Query: 161 PPHNELSPSPAGTVPSPGPALESPT---PNHG---PTP-----------ANSPSSPTPTP 203
PP S G VP P P PT P+ P P A +P +P P P
Sbjct: 668 PPPPFSDSSLPGLVPPPPPLPTGPTSVTPHFAFGPPLPPQLSEGCRDFQAPAPPAPPPLP 727
Query: 204 ENSSPSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSP 246
P P P + +PP PP PPP P + T + P
Sbjct: 728 GLGPPVPPPLPGSGLPP--PP---PPPGPGLFFNSTLSSSQGP 765
>FHOS_HUMAN (Q9Y613) FH1/FH2 domains-containing protein (Formin
homolog overexpressed in spleen) (FHOS) (Formin homology
2 domain containing 1)
Length = 1164
Score = 90.9 bits (224), Expect = 1e-17
Identities = 129/516 (25%), Positives = 201/516 (38%), Gaps = 92/516 (17%)
Query: 338 DNNTSVVEATSSEGVGQVPEPPNGKP-----PPPPPGPPPPPPPRKPPAPAPRPPPPPRA 392
D + ++ S E +P P P PPPPP PPP PP P PPPPP
Sbjct: 552 DEDQDMLNVESVEAGKDIPAPSPPLPLLSGVPPPPPLPPP------PPIKGPFPPPPPL- 604
Query: 393 GQPPPAPPKPVKVGNQGANQGGTSDGDAPKPKLKPFFWDKVAAKPDQQMVWHEISAGSF- 451
P A P P V + A K K FW V H +SA F
Sbjct: 605 ---PLAAPLPHSVPDSSAL--------PTKRKTVKLFWRDVKLAGG-----HGVSASRFG 648
Query: 452 ------------VFNEEKMESLFGCANQNRNERKKDSPSVDTSVQYIQIIDPKKAQNLSI 499
+ ++E LF + KK T ++DPK+ ++I
Sbjct: 649 PCATLWASLDPVSVDTARLEHLFESRAKEVLPSKKAGEGRRTMT---TVLDPKRTNAINI 705
Query: 500 LLRALNVTTDEVIDALREGNEIPV--ELIQTLLKMAPTTEEELKL---RLFTGELSQLGP 554
L L + AL +E V + I+ LL M PT EE K+ +L ++ LGP
Sbjct: 706 GLTTL-PPVHVIKAALLNFDEFAVSKDGIEKLLTMMPTEEERQKIEGAQLANPDI-PLGP 763
Query: 555 AERFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEA 614
AE FL L I RL+ F I E L+V ++L ++ F +L
Sbjct: 764 AENFLMTLASIGGLAARLQLWAFKLDYDSMEREIAEPLFDLKVGMEQLVQNATFRCILAT 823
Query: 615 VLKTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRA-VRT 673
+L GN +N G F L L K+SDVK T + +LLH + ++++ + + +
Sbjct: 824 LLAVGNFLNGSQSSG----FELSYLEKVSDVKDTVRRQSLLHHLCSLVLQTRPESSDLYS 879
Query: 674 ERESRSVVSSVGTDQ------DVDESPEASEEHYRSLGLQVVSSLSNELEDVKRAAL--- 724
E + + + V +Q ++ A+EE RSL +EL RA L
Sbjct: 880 EIPALTRCAKVDFEQLTENLGQLERRSRAAEESLRSLA-------KHELAPALRARLTHF 932
Query: 725 IDGDALSSAVSKLGH------------------SLAKTQEFMNTDLKSLDEESEFQRCTE 766
+D A A+ ++ H A+ M + E++ C E
Sbjct: 933 LDQCARRVAMLRIVHRRVCNRFHAFLLYLGYTPQAAREVRIMQFCHTLREFALEYRTCRE 992
Query: 767 RFVEKAREEVTWLQEEEKRIIAQVKSTADYFHGNAG 802
R +++ +++ T+ + + R ++ + + F G AG
Sbjct: 993 RVLQQQQKQATYRERNKTR--GRMITETEKFSGVAG 1026
>FMNL_HUMAN (O95466) Formin-like protein (Protein C17orf1)
Length = 463
Score = 84.3 bits (207), Expect = 1e-15
Identities = 107/447 (23%), Positives = 178/447 (38%), Gaps = 83/447 (18%)
Query: 433 VAAKPDQQMVWHEISAGSFV-FNEEK-MESLFGCANQNRNERKKDSPSVDTSV------- 483
VA KP Q I+ F N+EK ++ L + + + K PS+D S
Sbjct: 7 VALKPSQ------ITGTVFTELNDEKVLQELDMSDFEEQFKTKSQGPSLDLSALKSKAAQ 60
Query: 484 ---QYIQIIDPKKAQNLSILLRALNVTTDEVIDALR--EGNEIPVELIQTLLKMAPTTEE 538
+I+ +A+NL+I LR N+ + + A+ + + ++ ++ L++ PT E
Sbjct: 61 KAPSKATLIEANRAKNLAITLRKGNLGAERICQAIEAYDLQALGLDFLELLMRFLPTEYE 120
Query: 539 ELKLRLFTGE---LSQLGPAERFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATL 595
+ F E + +L +RF+ IP +R+ TL F+ + A + +
Sbjct: 121 RSLITRFEREQRPMEELSEEDRFMLCFSRIPRLPERMTTLTFLGNFPDTAQLLMPQLNAI 180
Query: 596 EVASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLL 655
AS ++ S ++LE VL GN MN RG A FRL +L L ++K TD K TLL
Sbjct: 181 IAASMSIKSSDKLRQILEIVLAFGNYMNSSK-RGAAYGFRLQSLDALLEMKSTDRKQTLL 239
Query: 656 HFVVQEIIRSEGIRAVRTERESRSVVSSVGTDQDVDESPEASEEHYRSLGLQVVSSLSNE 715
H++V+ I ++ P+ + H ++
Sbjct: 240 HYLVKVI---------------------------AEKYPQLTGFH-------------SD 259
Query: 716 LEDVKRAALIDGDALSSAVSKLGHSLAKTQ-EFMNTDLKSLDEESEFQRCTERFVEKARE 774
L + +A + D++ + V L L TQ EF+ D EF R ++K
Sbjct: 260 LHFLDKAGSVSLDSVLADVRSLQRGLELTQREFVRQD--DCMVLKEFLRANSPTMDK--- 314
Query: 775 EVTWLQEEEKRIIAQVKSTADYFHGNAGRDEGLRLFLIVRDFLIILDKVCKEVRDKTLRA 834
L + K +S +YF N F + F+ K +EV
Sbjct: 315 ----LLADSKTAQEAFESVVEYFGENPKTTSPGLFFSLFSRFIKAYKKAEQEV------- 363
Query: 835 VKSSQRKEALSVPSSPDTRHQQSPPAP 861
+KEA + + DT + PPAP
Sbjct: 364 --EQWKKEAAAQEAGADTPGKGEPPAP 388
>YPRO_OWEFU (P21260) Hypothetical proline-rich protein (Fragment)
Length = 141
Score = 79.3 bits (194), Expect = 4e-14
Identities = 31/47 (65%), Positives = 31/47 (65%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
P PP PPPPPP PPPPPPP PP P P PPPPP PPP PP P
Sbjct: 9 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 55
Score = 79.3 bits (194), Expect = 4e-14
Identities = 31/47 (65%), Positives = 31/47 (65%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
P PP PPPPPP PPPPPPP PP P P PPPPP PPP PP P
Sbjct: 12 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 58
Score = 79.3 bits (194), Expect = 4e-14
Identities = 31/47 (65%), Positives = 31/47 (65%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
P PP PPPPPP PPPPPPP PP P P PPPPP PPP PP P
Sbjct: 11 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 57
Score = 79.3 bits (194), Expect = 4e-14
Identities = 31/47 (65%), Positives = 31/47 (65%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
P PP PPPPPP PPPPPPP PP P P PPPPP PPP PP P
Sbjct: 10 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 56
Score = 79.0 bits (193), Expect = 5e-14
Identities = 30/50 (60%), Positives = 33/50 (66%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKPVKV 405
P PP PPPPPP PPPPPPP PP P P PPPPP PPP PP+ ++
Sbjct: 14 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPRRARI 63
Score = 42.7 bits (99), Expect = 0.004
Identities = 20/55 (36%), Positives = 22/55 (39%)
Query: 180 ALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPSPEK 234
+L P P P P P P P P P P P PP PP PPP P +
Sbjct: 6 SLTPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPRR 60
Score = 42.4 bits (98), Expect = 0.006
Identities = 21/52 (40%), Positives = 21/52 (40%)
Query: 181 LESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPSP 232
L S TP P P P P P P P P P PP PP PPP P
Sbjct: 4 LHSLTPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 55
Score = 41.6 bits (96), Expect = 0.010
Identities = 19/54 (35%), Positives = 21/54 (38%)
Query: 184 PTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPSPEKLQH 237
P P P P P P P P P P P PP PP PPP ++ H
Sbjct: 12 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPRRARIHH 65
Score = 39.3 bits (90), Expect = 0.047
Identities = 19/52 (36%), Positives = 20/52 (37%)
Query: 175 PSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQ 226
P P P P P P P P P P P P P P PP PPR+
Sbjct: 9 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPRR 60
Score = 38.1 bits (87), Expect = 0.11
Identities = 19/55 (34%), Positives = 21/55 (37%)
Query: 198 SPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKGDR 252
+P P P P P P PP PP PPP P P P PP + R
Sbjct: 8 TPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPRRAR 62
Score = 37.7 bits (86), Expect = 0.14
Identities = 19/54 (35%), Positives = 21/54 (38%)
Query: 194 NSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSPP 247
+S + P P P P P P PP PP PPP P P P PP
Sbjct: 5 HSLTPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 58
>SSGP_VOLCA (P21997) Sulfated surface glycoprotein 185 precursor
(SSG 185)
Length = 485
Score = 75.1 bits (183), Expect = 8e-13
Identities = 29/47 (61%), Positives = 30/47 (63%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
P PP+ PPPPPP PPPPPPP P P P PPPPP PPP P P
Sbjct: 253 PPPPSPSPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSP 299
Score = 75.1 bits (183), Expect = 8e-13
Identities = 29/47 (61%), Positives = 30/47 (63%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
P PP+ PP P P PPPPPPP PP P P PPPPP PPP PP P
Sbjct: 248 PRPPSPPPPSPSPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPP 294
Score = 74.3 bits (181), Expect = 1e-12
Identities = 29/47 (61%), Positives = 29/47 (61%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
P PP P PPPP PPPPPPP PP P PPPPP PPP PP P
Sbjct: 251 PSPPPPSPSPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSP 297
Score = 74.3 bits (181), Expect = 1e-12
Identities = 29/47 (61%), Positives = 30/47 (63%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
P PP PPPPPP PPP PPP PP P P PPPPP + PP PP P
Sbjct: 260 PPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSPPRKPPSP 306
Score = 73.2 bits (178), Expect = 3e-12
Identities = 28/46 (60%), Positives = 30/46 (64%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPK 401
P P PPPPPP PPPPPPP PP P P PPPPP P P+PP+
Sbjct: 256 PSPSPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSPPR 301
Score = 72.4 bits (176), Expect = 5e-12
Identities = 29/49 (59%), Positives = 30/49 (61%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKPVK 404
P P PPPPPP PPPPPPP P P P PPPPP PPP+P P K
Sbjct: 254 PPPSPSPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSPPRK 302
Score = 70.5 bits (171), Expect = 2e-11
Identities = 28/48 (58%), Positives = 30/48 (62%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKPV 403
P PP PPPPPP PPPPPP PP P P PPP P + PP+P PV
Sbjct: 263 PPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSPPRKPPSPSPPV 310
Score = 70.1 bits (170), Expect = 2e-11
Identities = 29/64 (45%), Positives = 33/64 (51%)
Query: 339 NNTSVVEATSSEGVGQVPEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPA 398
NN+ + + + P PP PP PP P P PPP PP P P PPPPP PPP
Sbjct: 225 NNSPLPPSPQPTASSRPPSPPPSPRPPSPPPPSPSPPPPPPPPPPPPPPPPPSPPPPPPP 284
Query: 399 PPKP 402
PP P
Sbjct: 285 PPPP 288
Score = 68.6 bits (166), Expect = 7e-11
Identities = 29/51 (56%), Positives = 30/51 (57%), Gaps = 1/51 (1%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPP-PPPRAGQPPPAPPKPVKV 405
P PP P PPPP PPPPPPP PP P+P PP PP P P PP P V
Sbjct: 269 PPPPPPPPSPPPPPPPPPPPPPPPPPPSPSPPRKPPSPSPPVPPPPSPPSV 319
Score = 67.8 bits (164), Expect = 1e-10
Identities = 28/48 (58%), Positives = 29/48 (60%), Gaps = 1/48 (2%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQP-PPAPPKP 402
P PP PPP PP PPPPPPP PP P P P PP + P PP PP P
Sbjct: 267 PPPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSPPRKPPSPSPPVPPPP 314
Score = 67.4 bits (163), Expect = 2e-10
Identities = 27/47 (57%), Positives = 28/47 (59%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
P PP PPPPP PPPPPPP PP P P P P P P P+PP P
Sbjct: 265 PPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSPPRKPPSPSPPVP 311
Score = 64.3 bits (155), Expect = 1e-09
Identities = 27/45 (60%), Positives = 29/45 (64%), Gaps = 3/45 (6%)
Query: 356 PEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPP 400
P PP PPPPPP PP P PPRKPP+P+P PPPP PP P
Sbjct: 280 PPPPPPPPPPPPPPPPSPSPPRKPPSPSPPVPPPP---SPPSVLP 321
Score = 61.6 bits (148), Expect = 9e-09
Identities = 40/112 (35%), Positives = 47/112 (41%), Gaps = 10/112 (8%)
Query: 156 NLISDPPHNE-LSPSPAGTV----PSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSS 210
N I P+N L PSP T PSP P+ P+P P SPS P P P P
Sbjct: 218 NPIGPAPNNSPLPPSPQPTASSRPPSPPPSPRPPSP-----PPPSPSPPPPPPPPPPPPP 272
Query: 211 TPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKGDRKQKAIILAVT 262
P P+ PP PP PPP P P PSPP +++ A T
Sbjct: 273 PPPPSPPPPPPPPPPPPPPPPPPSPSPPRKPPSPSPPVPPPPSPPSVLPAAT 324
Score = 57.0 bits (136), Expect = 2e-07
Identities = 40/112 (35%), Positives = 45/112 (39%), Gaps = 7/112 (6%)
Query: 161 PPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTA--MV 218
PP + PSP PSP P P P P P + P P P P P PSP+
Sbjct: 244 PPPSPRPPSPPPPSPSPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSPPRKP 303
Query: 219 PPTSPPRQHPPPSPEKLQHVTYAP-----DPSPPDKGDRKQKAIILAVTVSG 265
P SPP PP P L T P SP R A + AVT+SG
Sbjct: 304 PSPSPPVPPPPSPPSVLPAATGFPFCECVSRSPSSYPWRVTVANVSAVTISG 355
Score = 50.1 bits (118), Expect = 3e-05
Identities = 31/75 (41%), Positives = 33/75 (43%), Gaps = 20/75 (26%)
Query: 356 PEPPNGKPPPPPPGPPPPPP----PRKP--------PAPAPRPPPPPRAGQP-------- 395
P PP PPPPPP PPPPPP PRKP P P+P P G P
Sbjct: 276 PSPPPPPPPPPPPPPPPPPPSPSPPRKPPSPSPPVPPPPSPPSVLPAATGFPFCECVSRS 335
Query: 396 PPAPPKPVKVGNQGA 410
P + P V V N A
Sbjct: 336 PSSYPWRVTVANVSA 350
Score = 48.5 bits (114), Expect = 8e-05
Identities = 24/56 (42%), Positives = 26/56 (45%), Gaps = 4/56 (7%)
Query: 348 SSEGVGQVPEPPNGKPPPPPPGPPPPP-PPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
S V + PN P PP P P PP PP+P P PPPP P P PP P
Sbjct: 213 SGPNVNPIGPAPNNSPLPPSPQPTASSRPPSPPPSPRPPSPPPP---SPSPPPPPP 265
Score = 43.9 bits (102), Expect = 0.002
Identities = 31/70 (44%), Positives = 33/70 (46%), Gaps = 18/70 (25%)
Query: 181 LESPTPNH-GPTPANSPSSPTPTPENSS--PSSTPSPTAMVPPTSPPRQHPPPSPEKLQH 237
L P N GP P NSP P+P P SS PS PSP PP+ PP P PSP
Sbjct: 212 LSGPNVNPIGPAPNNSPLPPSPQPTASSRPPSPPPSPR---PPSPPP---PSPSP----- 260
Query: 238 VTYAPDPSPP 247
P P PP
Sbjct: 261 ----PPPPPP 266
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.312 0.132 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 121,977,001
Number of Sequences: 164201
Number of extensions: 6692597
Number of successful extensions: 150076
Number of sequences better than 10.0: 2553
Number of HSP's better than 10.0 without gapping: 1413
Number of HSP's successfully gapped in prelim test: 1198
Number of HSP's that attempted gapping in prelim test: 43658
Number of HSP's gapped (non-prelim): 35226
length of query: 896
length of database: 59,974,054
effective HSP length: 119
effective length of query: 777
effective length of database: 40,434,135
effective search space: 31417322895
effective search space used: 31417322895
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0203.14


