

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0201.10
(137 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
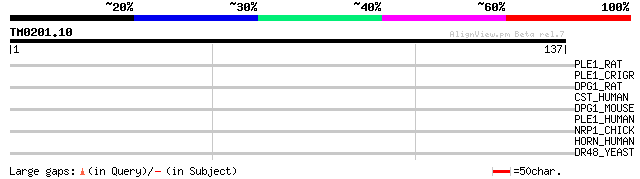
Score E
Sequences producing significant alignments: (bits) Value
PLE1_RAT (P30427) Plectin 1 (PLTN) (PCN) 31 0.91
PLE1_CRIGR (Q9JI55) Plectin 1 (PLTN) (PCN) (300-kDa intermediate... 30 1.6
DPG1_RAT (Q9QYV8) DNA polymerase gamma subunit 1 (EC 2.7.7.7) (M... 29 3.5
CST_HUMAN (Q86V15) Probable transcription factor CST (Castor rel... 29 3.5
DPG1_MOUSE (P54099) DNA polymerase gamma subunit 1 (EC 2.7.7.7) ... 28 4.5
PLE1_HUMAN (Q15149) Plectin 1 (PLTN) (PCN) (Hemidesmosomal prote... 28 5.9
NRP1_CHICK (P79795) Neuropilin-1 precursor (A5 protein) 28 5.9
HORN_HUMAN (Q86YZ3) Hornerin 28 7.7
DR48_YEAST (P18899) DDR48 stress protein (DNA damage-responsive ... 28 7.7
>PLE1_RAT (P30427) Plectin 1 (PLTN) (PCN)
Length = 4687
Score = 30.8 bits (68), Expect = 0.91
Identities = 14/53 (26%), Positives = 28/53 (52%), Gaps = 3/53 (5%)
Query: 69 NRNNNRAGAASYDKKGSFHYDSLSYALNFDDGEEDVYSYGGFSTRFASVPAST 121
+R +R G+ + ++GSF +++ F YS G+ R+AS P+++
Sbjct: 4629 SRTGSRTGSRAGSRRGSFDATGSGFSMTFSSSS---YSSSGYGRRYASGPSAS 4678
>PLE1_CRIGR (Q9JI55) Plectin 1 (PLTN) (PCN) (300-kDa intermediate
filament-associated protein) (IFAP300) (Fragment)
Length = 4473
Score = 30.0 bits (66), Expect = 1.6
Identities = 14/53 (26%), Positives = 27/53 (50%), Gaps = 3/53 (5%)
Query: 69 NRNNNRAGAASYDKKGSFHYDSLSYALNFDDGEEDVYSYGGFSTRFASVPAST 121
+R +R G+ + ++GSF +++ F YS G+ R+AS P ++
Sbjct: 4415 SRTGSRTGSRAGSRRGSFDATGSGFSMTFSSSS---YSSSGYGRRYASGPPAS 4464
>DPG1_RAT (Q9QYV8) DNA polymerase gamma subunit 1 (EC 2.7.7.7)
(Mitochondrial DNA polymerase catalytic subunit)
(PolG-alpha)
Length = 1216
Score = 28.9 bits (63), Expect = 3.5
Identities = 14/44 (31%), Positives = 22/44 (49%), Gaps = 7/44 (15%)
Query: 5 LFAKQACICYLIPCSSSSETSLSWWERVRSPENKEWWWAHGWNK 48
L A++ + YL +S SE L P+ ++W WA GW +
Sbjct: 124 LLAQKQSLPYLEAAASLSEAQLP-------PQPRKWVWAEGWTR 160
>CST_HUMAN (Q86V15) Probable transcription factor CST (Castor
related protein)
Length = 1044
Score = 28.9 bits (63), Expect = 3.5
Identities = 21/79 (26%), Positives = 30/79 (37%), Gaps = 12/79 (15%)
Query: 61 WKTFIRRFNRNNNRAGAASYDKKGSFH---------YDSLSYAL---NFDDGEEDVYSYG 108
WK ++RRF + G + K FH + +L A+ NF E + G
Sbjct: 884 WKVYLRRFGTKDFCDGQCDFLHKAHFHCVVEECGALFSTLDGAIKHANFHFRTEGGAAKG 943
Query: 109 GFSTRFASVPASTKPTFTP 127
F + A TKP P
Sbjct: 944 NTEAAFPASAAETKPPMAP 962
>DPG1_MOUSE (P54099) DNA polymerase gamma subunit 1 (EC 2.7.7.7)
(Mitochondrial DNA polymerase catalytic subunit)
(PolG-alpha)
Length = 1239
Score = 28.5 bits (62), Expect = 4.5
Identities = 15/44 (34%), Positives = 20/44 (45%), Gaps = 7/44 (15%)
Query: 5 LFAKQACICYLIPCSSSSETSLSWWERVRSPENKEWWWAHGWNK 48
L A++ + YL +S E L PE K W WA GW +
Sbjct: 124 LLAQKQSLPYLEAAASLLEAQLP-------PEPKSWAWAEGWTR 160
>PLE1_HUMAN (Q15149) Plectin 1 (PLTN) (PCN) (Hemidesmosomal protein 1)
(HD1)
Length = 4684
Score = 28.1 bits (61), Expect = 5.9
Identities = 13/48 (27%), Positives = 24/48 (49%), Gaps = 3/48 (6%)
Query: 69 NRNNNRAGAASYDKKGSFHYDSLSYALNFDDGEEDVYSYGGFSTRFAS 116
+R +R G+ + ++GSF +++ F YS G+ R+AS
Sbjct: 4626 SRTGSRTGSRAGSRRGSFDATGSGFSMTFSSSS---YSSSGYGRRYAS 4670
>NRP1_CHICK (P79795) Neuropilin-1 precursor (A5 protein)
Length = 914
Score = 28.1 bits (61), Expect = 5.9
Identities = 24/86 (27%), Positives = 37/86 (42%), Gaps = 14/86 (16%)
Query: 21 SSETSLSWWE---RVRSPENKEWWWAHGWNKVREWSEIIVGPKWKTFIRRFNRNNNRAGA 77
SS+ S W R+ PEN W G + VREW ++ +G + RF GA
Sbjct: 292 SSQYSAIWSSERSRLNYPENG---WTPGEDSVREWIQVDLG------LLRFVSGIGTQGA 342
Query: 78 ASYDKKGSFHYDSLSYALNFDDGEED 103
S + K ++ +Y ++ ED
Sbjct: 343 ISKETKKEYYLK--TYRVDVSSNGED 366
>HORN_HUMAN (Q86YZ3) Hornerin
Length = 2850
Score = 27.7 bits (60), Expect = 7.7
Identities = 20/66 (30%), Positives = 30/66 (45%), Gaps = 10/66 (15%)
Query: 77 AASYDKKGSFHYDSLSYALNFDDGEEDVYSY---------GGFSTRFASVPASTKPTFTP 127
++SY + GS S Y+ + +D YSY G + F S P+ST P +
Sbjct: 2782 SSSYGQHGSGSGQSSGYSQHGSGSGQDGYSYCKGGSNHDGGSSGSYFLSFPSSTSP-YEY 2840
Query: 128 VMVRRC 133
V +RC
Sbjct: 2841 VQEQRC 2846
>DR48_YEAST (P18899) DDR48 stress protein (DNA damage-responsive
protein 48) (DDRP 48) (YP 75) (Flocculent specific
protein)
Length = 429
Score = 27.7 bits (60), Expect = 7.7
Identities = 17/59 (28%), Positives = 30/59 (50%), Gaps = 2/59 (3%)
Query: 69 NRNNNRAGAASYDKKGSFHYDSLSYALNFDD--GEEDVYSYGGFSTRFASVPASTKPTF 125
+ NN+ G+ + D GS + + SY N DD G + SYG + + +S +S ++
Sbjct: 156 SNNNDSYGSNNDDSYGSSNKNKSSYGSNNDDSYGSNNDDSYGSSNKKKSSYGSSNNDSY 214
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.133 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,645,488
Number of Sequences: 164201
Number of extensions: 724144
Number of successful extensions: 1414
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 1409
Number of HSP's gapped (non-prelim): 13
length of query: 137
length of database: 59,974,054
effective HSP length: 98
effective length of query: 39
effective length of database: 43,882,356
effective search space: 1711411884
effective search space used: 1711411884
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0201.10


