

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0197.5
(317 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
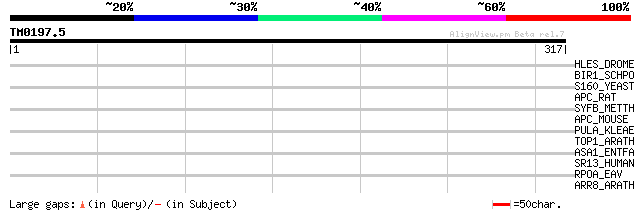
Score E
Sequences producing significant alignments: (bits) Value
HLES_DROME (Q02308) Hairless protein 35 0.25
BIR1_SCHPO (O14064) Bir1 protein (Chromosome segregation protein... 34 0.55
S160_YEAST (P06105) SCP160 protein (Protein HX) 33 1.2
APC_RAT (P70478) Adenomatous polyposis coli protein (APC protein) 33 1.2
SYFB_METTH (O26864) Phenylalanyl-tRNA synthetase beta chain (EC ... 32 2.7
APC_MOUSE (Q61315) Adenomatous polyposis coli protein (APC prote... 32 2.7
PULA_KLEAE (P07811) Pullulanase precursor (EC 3.2.1.41) (Alpha-d... 31 3.5
TOP1_ARATH (P30181) DNA topoisomerase I (EC 5.99.1.2) 31 4.6
ASA1_ENTFA (P17953) Aggregation substance precursor 30 6.1
SR13_HUMAN (Q9Y3M8) StAR-related lipid transfer protein 13 (StAR... 30 7.9
RPOA_EAV (P19811) POL polyprotein (ORF1A/1B) [Contains: RNA-dire... 30 7.9
ARR8_ARATH (O80365) Two-component response regulator ARR8 (Respo... 30 7.9
>HLES_DROME (Q02308) Hairless protein
Length = 1077
Score = 35.0 bits (79), Expect = 0.25
Identities = 23/97 (23%), Positives = 49/97 (49%), Gaps = 3/97 (3%)
Query: 64 LEMEGLVVFSNGVSTAR--KPETVVGESSSERDQYRPESDWNRSSVSRKANIAREVSSPV 121
+E+E + V + + KP+T+ GE +ER + P+ + S S++A+ ++V P
Sbjct: 391 VEIENVAVADTTTNEIKIEKPDTIKGEDDAERLEKEPKKAVSDDSESKEASPGQQV-EPQ 449
Query: 122 AVEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKG 158
++ + + + EE +L R+++ V G+ G
Sbjct: 450 PKDETVDVEMKMNTSEDEEPMTELPRITNAVNGDLNG 486
>BIR1_SCHPO (O14064) Bir1 protein (Chromosome segregation protein
cut17)
Length = 997
Score = 33.9 bits (76), Expect = 0.55
Identities = 25/67 (37%), Positives = 32/67 (47%), Gaps = 9/67 (13%)
Query: 76 VSTARKPETVVGESSSE--RDQYRPESDWNRSSVSRKANIAREVSSPVAVEQVAGSKPQA 133
V KP+T + E E R ES R S+ R R+VSSPV+ E ++
Sbjct: 642 VDFIEKPKTEISEVLPEEKRKAICDESQTVRVSIDRGVTKTRDVSSPVSDE-------KS 694
Query: 134 ENANHEE 140
EN NHEE
Sbjct: 695 ENVNHEE 701
>S160_YEAST (P06105) SCP160 protein (Protein HX)
Length = 1222
Score = 32.7 bits (73), Expect = 1.2
Identities = 35/150 (23%), Positives = 62/150 (41%), Gaps = 11/150 (7%)
Query: 37 EIGEDMRFMLAELTIEAMIDGWLFIVLLEMEGLVVFSNGVSTARKPETVVGESSS-ERDQ 95
E G +M F+ +A G + E+E N A++ E++V E+S +
Sbjct: 811 EYGVEMDFLQKSTDPKAQETGEV-----ELEITGSRQNIKDAAKRVESIVAEASDFVTEV 865
Query: 96 YRPESDWNRSSVSRKANIAREVSSPVAVEQVAGSKPQAENANHEEKPLDLGRMSHLVCGE 155
+ + +++S V +I RE+ S E++ NA+ E K + + V
Sbjct: 866 LKIDHKYHKSIVGSGGHILREIISKAGGEEIRNKSVDIPNADSENKDITVQGPQKFV--- 922
Query: 156 SKGATEEAMSVTAEARH-VVKNADVGLMRK 184
K EE + +A + V K D+ RK
Sbjct: 923 -KKVVEEINKIVKDAENSVTKTIDIPAERK 951
>APC_RAT (P70478) Adenomatous polyposis coli protein (APC protein)
Length = 2842
Score = 32.7 bits (73), Expect = 1.2
Identities = 27/122 (22%), Positives = 50/122 (40%), Gaps = 11/122 (9%)
Query: 74 NGVSTARK---------PETVVGESSS--ERDQYRPESDWNRSSVSRKANIAREVSSPVA 122
NG+ST K P T +SS + P ++ ++S++ +++ SS
Sbjct: 2334 NGISTPNKLSQLPRTSSPSTASTKSSGSGKMSYTSPGRQLSQQNLSKQTGLSKNASSIPR 2393
Query: 123 VEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKGATEEAMSVTAEARHVVKNADVGLM 182
E + Q N+N K ++L RMS S+ E ++ ++ + + L
Sbjct: 2394 SESASKGLNQMNNSNGSNKKVELSRMSSTKSSGSESDRSERPALVRQSTFIKEAPSPTLR 2453
Query: 183 RK 184
RK
Sbjct: 2454 RK 2455
>SYFB_METTH (O26864) Phenylalanyl-tRNA synthetase beta chain (EC
6.1.1.20) (Phenylalanine--tRNA ligase beta chain)
(PheRS)
Length = 549
Score = 31.6 bits (70), Expect = 2.7
Identities = 14/31 (45%), Positives = 20/31 (64%)
Query: 218 RSEQVCGDTQKAAKEDSWGASDASLQQVMVG 248
R E D A+ED+W +DAS+++VMVG
Sbjct: 345 RIEPELPDIATIAEEDTWSRADASIREVMVG 375
>APC_MOUSE (Q61315) Adenomatous polyposis coli protein (APC protein)
(mAPC)
Length = 2845
Score = 31.6 bits (70), Expect = 2.7
Identities = 26/122 (21%), Positives = 51/122 (41%), Gaps = 11/122 (9%)
Query: 74 NGVSTARK---------PETVVGESSS--ERDQYRPESDWNRSSVSRKANIAREVSSPVA 122
NG+S K P T +SS + P ++ +++++A++++ SS
Sbjct: 2334 NGISPPNKLSQLPRTSSPSTASTKSSGSGKMSYTSPGRQLSQQNLTKQASLSKNASSIPR 2393
Query: 123 VEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKGATEEAMSVTAEARHVVKNADVGLM 182
E + Q N N K ++L RMS S+ + E ++ ++ + + L
Sbjct: 2394 SESASKGLNQMSNGNGSNKKVELSRMSSTKSSGSESDSSERPALVRQSTFIKEAPSPTLR 2453
Query: 183 RK 184
RK
Sbjct: 2454 RK 2455
>PULA_KLEAE (P07811) Pullulanase precursor (EC 3.2.1.41)
(Alpha-dextrin endo-1,6-alpha-glucosidase) (Pullulan
6-glucanohydrolase)
Length = 1096
Score = 31.2 bits (69), Expect = 3.5
Identities = 23/66 (34%), Positives = 29/66 (43%), Gaps = 15/66 (22%)
Query: 258 DQARVSQVVDARNTN-----------IQDGFRSGIAVDDDVSQVCVTPQEGSKRRRGKPK 306
D A V + VD RNT I DG ++G Q C + G +RR GKP
Sbjct: 983 DGATVMKRVDFRNTGADQQTGLLVMTIDDGMQAG----RQSGQPCRRHRGGDQRRAGKPD 1038
Query: 307 GAAIRR 312
A +RR
Sbjct: 1039 AAGLRR 1044
>TOP1_ARATH (P30181) DNA topoisomerase I (EC 5.99.1.2)
Length = 916
Score = 30.8 bits (68), Expect = 4.6
Identities = 25/80 (31%), Positives = 37/80 (46%), Gaps = 5/80 (6%)
Query: 82 PETVVGESSSERDQYRPESDWNRSSVSRK-ANIAREVSSPVAVEQVAGSKPQAENANHEE 140
P TV S ++DQ + + S R ++I P + QV S PQ E N+ +
Sbjct: 99 PSTVKDRSQLQKDQSECKIEHEDSEDDRPLSSILSGNKGPTSSRQV--SSPQPEKKNNGD 156
Query: 141 KPLDLGRMSHLVCGESKGAT 160
+PLD R S ++ ES T
Sbjct: 157 RPLD--RASRIIKDESDDET 174
>ASA1_ENTFA (P17953) Aggregation substance precursor
Length = 1296
Score = 30.4 bits (67), Expect = 6.1
Identities = 26/105 (24%), Positives = 50/105 (46%), Gaps = 5/105 (4%)
Query: 61 IVLLEMEGLVVFSNGVSTARKPETVVGESSSERDQYRPE--SDWNRSSVSRKANIAREVS 118
I+ + + G+V + A + +T G ++ + D P+ S +++V+ +A + ++ +
Sbjct: 25 ILFIGVLGVVGLATDDVQAAELDTQPGTTTVQPDNPDPQVGSTTPKTAVTEEATVQKDTT 84
Query: 119 S-PVAVEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKGATEE 162
S P VE+VA K AE ++ P D G K A E+
Sbjct: 85 SQPTKVEEVASEKNGAEQSS--ATPNDTTNAQQPTVGAEKSAQEQ 127
>SR13_HUMAN (Q9Y3M8) StAR-related lipid transfer protein 13
(StARD13) (START domain-containing protein 13) (46H23.2)
Length = 995
Score = 30.0 bits (66), Expect = 7.9
Identities = 22/86 (25%), Positives = 38/86 (43%), Gaps = 8/86 (9%)
Query: 233 DSWGAS---DASLQQVMVGEHEVKDF-SPDQARVSQVVDARNTNIQDGFRSGIAVDDDVS 288
D W + +VGE + F SP+Q +D ++ +G + V+ DV+
Sbjct: 383 DDWSKDVLPELQTHDTLVGEPGLSTFPSPNQI----TLDFEGNSVSEGRTTPSDVERDVT 438
Query: 289 QVCVTPQEGSKRRRGKPKGAAIRRPN 314
+ + G + RR GA++ RPN
Sbjct: 439 SLNESEPPGVRDRRDSGVGASLTRPN 464
>RPOA_EAV (P19811) POL polyprotein (ORF1A/1B) [Contains: RNA-directed
RNA polymerase (EC 2.7.7.48); Helicase; Protease (EC
3.4.21.-)]
Length = 3175
Score = 30.0 bits (66), Expect = 7.9
Identities = 29/107 (27%), Positives = 46/107 (42%), Gaps = 5/107 (4%)
Query: 209 VVQINNVWLRSEQVCGDTQKAAKEDSWGASDASLQQVMVGEHEVKDFSPDQA---RVSQV 265
V Q N+ S QV D ++ + + A A Q+V G+ V D+ + V
Sbjct: 1493 VAQYRNILNASLQVDRDAARSRRLMAKLADFAVEQEVTAGDRVVVIDGLDRMAHFKDDLV 1552
Query: 266 VDARNTNIQDGFRSGIA--VDDDVSQVCVTPQEGSKRRRGKPKGAAI 310
+ T + G R I V ++ + V P +RR+G PKGA +
Sbjct: 1553 LVPLTTKVVGGSRCTICDVVKEEANDTPVKPMPSRRRRKGLPKGAQL 1599
>ARR8_ARATH (O80365) Two-component response regulator ARR8 (Response
reactor 3)
Length = 225
Score = 30.0 bits (66), Expect = 7.9
Identities = 11/36 (30%), Positives = 20/36 (55%)
Query: 110 KANIAREVSSPVAVEQVAGSKPQAENANHEEKPLDL 145
K + +E PVA+E++ SKP+ E E +++
Sbjct: 146 KTKLKKESEKPVAIEEIVVSKPEIEEEEEESSVIEI 181
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.131 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,052,238
Number of Sequences: 164201
Number of extensions: 1295354
Number of successful extensions: 3296
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 3292
Number of HSP's gapped (non-prelim): 13
length of query: 317
length of database: 59,974,054
effective HSP length: 110
effective length of query: 207
effective length of database: 41,911,944
effective search space: 8675772408
effective search space used: 8675772408
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0197.5


