

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0195b.8
(83 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
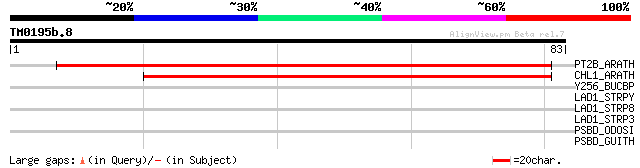
Score E
Sequences producing significant alignments: (bits) Value
PT2B_ARATH (P46032) Peptide transporter PTR2-B (Histidine transp... 104 4e-23
CHL1_ARATH (Q05085) Nitrate/chlorate transporter 72 3e-13
Y256_BUCBP (Q89AL2) Hypothetical UPF0259 protein bbp256 32 0.22
LAD1_STRPY (P63703) Tagatose 1,6-diphosphate aldolase 1 (EC 4.1.... 28 4.2
LAD1_STRP8 (P63704) Tagatose 1,6-diphosphate aldolase 1 (EC 4.1.... 28 4.2
LAD1_STRP3 (Q8K654) Tagatose 1,6-diphosphate aldolase 1 (EC 4.1.... 28 4.2
PSBD_ODOSI (P49478) Photosystem II D2 protein (Photosystem Q(A) ... 27 9.3
PSBD_GUITH (O78427) Photosystem II D2 protein (Photosystem Q(A) ... 27 9.3
>PT2B_ARATH (P46032) Peptide transporter PTR2-B (Histidine
transporting protein)
Length = 585
Score = 104 bits (260), Expect = 4e-23
Identities = 48/74 (64%), Positives = 61/74 (81%)
Query: 8 LVSDSESRSGDYANPWRLFTVTQVEELKILIRVLPIWATEIVFSAVYALLSTLLVEQGTM 67
++S+ ES+SGDY+N WRL TVTQVEELKILIR+ PIWA+ I+FSAVYA +ST+ V+QG
Sbjct: 318 VISEEESKSGDYSNSWRLCTVTQVEELKILIRMFPIWASGIIFSAVYAQMSTMFVQQGRA 377
Query: 68 MNRSFGSFNIPPAS 81
MN GSF +PPA+
Sbjct: 378 MNCKIGSFQLPPAA 391
>CHL1_ARATH (Q05085) Nitrate/chlorate transporter
Length = 590
Score = 72.0 bits (175), Expect = 3e-13
Identities = 35/61 (57%), Positives = 45/61 (73%)
Query: 21 NPWRLFTVTQVEELKILIRVLPIWATEIVFSAVYALLSTLLVEQGTMMNRSFGSFNIPPA 80
N W L T+T VEE+K ++R+LPIWAT I+F V+A L+TL V Q ++RS GSF IPPA
Sbjct: 323 NKWTLSTLTDVEEVKQIVRMLPIWATCILFWTVHAQLTTLSVAQSETLDRSIGSFEIPPA 382
Query: 81 S 81
S
Sbjct: 383 S 383
>Y256_BUCBP (Q89AL2) Hypothetical UPF0259 protein bbp256
Length = 250
Score = 32.3 bits (72), Expect = 0.22
Identities = 18/53 (33%), Positives = 31/53 (57%), Gaps = 3/53 (5%)
Query: 25 LFTVTQVEELKILIRVLPIWATEIVFSAVYALLSTLLVEQGTMMNRSFGSFNI 77
+FT T + +L L+ +P + T I+FS + LL+E+ T++N + S NI
Sbjct: 132 IFTTTIITQLGFLLYFIPGFTTIILFSLSPII---LLIEEKTILNSIYASINI 181
>LAD1_STRPY (P63703) Tagatose 1,6-diphosphate aldolase 1 (EC
4.1.2.40) (Tagatose-bisphosphate aldolase 1)
(D-tagatose-1,6-bisphosphate aldolase 1)
Length = 325
Score = 28.1 bits (61), Expect = 4.2
Identities = 15/34 (44%), Positives = 22/34 (64%), Gaps = 1/34 (2%)
Query: 27 TVTQVEELKILI-RVLPIWATEIVFSAVYALLST 59
TVTQ+E LK+L+ L +A+ I+ Y LL+T
Sbjct: 44 TVTQIETLKVLVSEELTPYASSILLDPEYGLLAT 77
>LAD1_STRP8 (P63704) Tagatose 1,6-diphosphate aldolase 1 (EC
4.1.2.40) (Tagatose-bisphosphate aldolase 1)
(D-tagatose-1,6-bisphosphate aldolase 1)
Length = 325
Score = 28.1 bits (61), Expect = 4.2
Identities = 15/34 (44%), Positives = 22/34 (64%), Gaps = 1/34 (2%)
Query: 27 TVTQVEELKILI-RVLPIWATEIVFSAVYALLST 59
TVTQ+E LK+L+ L +A+ I+ Y LL+T
Sbjct: 44 TVTQIETLKVLVSEELTPYASSILLDPEYGLLAT 77
>LAD1_STRP3 (Q8K654) Tagatose 1,6-diphosphate aldolase 1 (EC
4.1.2.40) (Tagatose-bisphosphate aldolase 1)
(D-tagatose-1,6-bisphosphate aldolase 1)
Length = 325
Score = 28.1 bits (61), Expect = 4.2
Identities = 15/34 (44%), Positives = 22/34 (64%), Gaps = 1/34 (2%)
Query: 27 TVTQVEELKILI-RVLPIWATEIVFSAVYALLST 59
TVTQ+E LK+L+ L +A+ I+ Y LL+T
Sbjct: 44 TVTQIETLKVLVSEELTPYASSILLDPEYGLLAT 77
>PSBD_ODOSI (P49478) Photosystem II D2 protein (Photosystem Q(A)
protein) (PSII D2 protein)
Length = 351
Score = 26.9 bits (58), Expect = 9.3
Identities = 13/37 (35%), Positives = 19/37 (51%)
Query: 9 VSDSESRSGDYANPWRLFTVTQVEELKILIRVLPIWA 45
V ++ GD AN +R FT TQ EE ++ W+
Sbjct: 217 VENTLFEDGDAANTFRAFTPTQSEETYSMVTANRFWS 253
>PSBD_GUITH (O78427) Photosystem II D2 protein (Photosystem Q(A)
protein) (PSII D2 protein)
Length = 351
Score = 26.9 bits (58), Expect = 9.3
Identities = 13/37 (35%), Positives = 19/37 (51%)
Query: 9 VSDSESRSGDYANPWRLFTVTQVEELKILIRVLPIWA 45
V ++ GD AN +R FT TQ EE ++ W+
Sbjct: 217 VQNTIFEDGDAANTFRAFTPTQSEETYSMVTANRFWS 253
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,764,282
Number of Sequences: 164201
Number of extensions: 257120
Number of successful extensions: 555
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 551
Number of HSP's gapped (non-prelim): 8
length of query: 83
length of database: 59,974,054
effective HSP length: 59
effective length of query: 24
effective length of database: 50,286,195
effective search space: 1206868680
effective search space used: 1206868680
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0195b.8


