

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0195a.11
(377 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
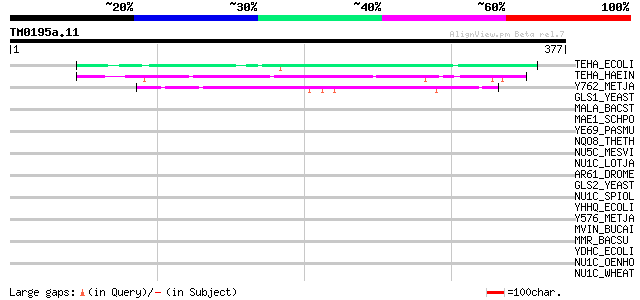
TEHA_ECOLI (P25396) Tellurite resistance protein tehA 74 5e-13
TEHA_HAEIN (P44741) Tellurite resistance protein tehA homolog 62 2e-09
Y762_METJA (Q58172) Hypothetical protein MJ0762 47 1e-04
GLS1_YEAST (P38631) 1,3-beta-glucan synthase component GLS1 (EC ... 37 0.082
MALA_BACST (Q45632) Maltose permease 35 0.41
MAE1_SCHPO (P50537) Malic acid transport protein (Malate permease) 35 0.41
YE69_PASMU (Q9CKY2) Hypothetical protein PM1469 34 0.70
NQO8_THETH (Q60019) NADH-quinone oxidoreductase chain 8 (EC 1.6.... 33 0.91
NU5C_MESVI (Q9MUK8) NAD(P)H-quinone oxidoreductase chain 5, chlo... 33 1.2
NU1C_LOTJA (Q9BBN9) NAD(P)H-quinone oxidoreductase chain 1, chlo... 33 1.2
AR61_DROME (Q9VES1) ARL-6 interacting protein-1 homolog 33 1.5
GLS2_YEAST (P40989) 1,3-beta-glucan synthase component GLS2 (EC ... 32 2.0
NU1C_SPIOL (Q9M3I6) NAD(P)H-quinone oxidoreductase chain 1, chlo... 32 2.6
YHHQ_ECOLI (P37619) Hypothetical protein yhhQ 32 3.5
Y576_METJA (Q57996) Hypothetical protein MJ0576 32 3.5
MVIN_BUCAI (P57415) Virulence factor mviN homolog 31 4.5
MMR_BACSU (Q00538) Methylenomycin A resistance protein (MMR pept... 31 4.5
YDHC_ECOLI (P37597) Hypothetical transport protein ydhC 31 5.9
NU1C_OENHO (Q9MTH7) NAD(P)H-quinone oxidoreductase chain 1, chlo... 31 5.9
NU1C_WHEAT (Q95H43) NAD(P)H-quinone oxidoreductase chain 1, chlo... 30 7.7
>TEHA_ECOLI (P25396) Tellurite resistance protein tehA
Length = 330
Score = 74.3 bits (181), Expect = 5e-13
Identities = 84/320 (26%), Positives = 126/320 (39%), Gaps = 29/320 (9%)
Query: 46 LHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSL 105
L AGYF I L G W+ S + V H + + LA+ LL+
Sbjct: 9 LPAGYFGIVLGTIGMGFAWRYA-------SQVWQVSHWLGDGLVI----LAMIIWGLLTS 57
Query: 106 LYLLRCLFHFNMVKAEFLHHVGVNYL-FAPWISWFLLLQSAPFVAPKTATYLVLWWVFAV 164
++ R + + V AE H V +++ P + + + P+ P +F+
Sbjct: 58 AFIARLIRFPHSVLAEVRHPVLSSFVSLFPATTMLVAIGFVPWFRPLAVC------LFSF 111
Query: 165 PVVVLDVKIYGQWFTKGK---RFLSTAANPTSQL-SVIGNLVGAQAAAHMGWKESAVCLF 220
VVV Y W T G A P L +V N + A A +G+ ++ +
Sbjct: 112 GVVVQ--LAYAAWQTAGLWRGSHPEEATTPGLYLPTVANNFISAMACGALGYTDAGLVFL 169
Query: 221 SLGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGG-FDTLSKM 279
G+ +L L + QRL LP LR + A VA AW S+ GG DTL+KM
Sbjct: 170 GAGVFSWLSLEPVILQRLRSSGELPTALRTSLGIQLAPALVACSAWLSVNGGEGDTLAKM 229
Query: 280 LFFLSLF-LFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVL 338
LF L L L P + FN ++W++SF V+ LA H L
Sbjct: 230 LFGYGLLQLLFMLRLMPWYLSQP---FNASFWSFSFGVSALATTGLHLGSGSDNGFFHTL 286
Query: 339 MLVLLALSVLVSLALTVFTF 358
+ L + + L + TF
Sbjct: 287 AVPLFIFTNFIIAILLIRTF 306
>TEHA_HAEIN (P44741) Tellurite resistance protein tehA homolog
Length = 328
Score = 62.4 bits (150), Expect = 2e-09
Identities = 77/318 (24%), Positives = 138/318 (43%), Gaps = 35/318 (11%)
Query: 46 LHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFL--VLWSLALFTLALL 103
L GYF I L L +L W H+ ++ P++ + VL +A L
Sbjct: 23 LPTGYFGIPLGLAALSLAWF-------------HLENLFPAARMVSDVLGIVASAVWILF 69
Query: 104 SLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWWVFA 163
L+Y + ++F V+AE+ H V + F I +L VL W+
Sbjct: 70 ILMYAYKLRYYFEEVRAEY--HSPVRFSFIALIPITTMLVGDILYRWNPLIAEVLIWIGT 127
Query: 164 VPVVVLD-VKIYGQWFTKGKRFLSTAANPTSQL-SVIGNLVGAQAAAHMGWKESAVCLFS 221
+ ++ +++ W +G F + +P+ L +V N A + A +G+ + F
Sbjct: 128 IGQLLFSTLRVSELW--QGGVFEQKSTHPSFYLPAVAANFTSASSLALLGYHDLGYLFFG 185
Query: 222 LGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGG-FDTLSKML 280
GM+ +++ L Q L + P R + A V A+ SI G DTL+K+L
Sbjct: 186 AGMIAWIIFEPVLLQHLRISSLEPQF-RATMGIVLAPAFVCVSAYLSINHGEVDTLAKIL 244
Query: 281 F---FLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDY--AEEVKGT-- 333
+ FL LF + L P + + + N+ WA+SF + +A ++T + ++G
Sbjct: 245 WGYGFLQLFFLLRLF--PWIVEKGL---NIGLWAFSFGLASMANSATAFYHGNVLQGVSI 299
Query: 334 ISHVLMLVLLALSVLVSL 351
+ V V++ L VL+++
Sbjct: 300 FAFVFSNVMIGLLVLMTI 317
>Y762_METJA (Q58172) Hypothetical protein MJ0762
Length = 342
Score = 46.6 bits (109), Expect = 1e-04
Identities = 59/267 (22%), Positives = 115/267 (42%), Gaps = 26/267 (9%)
Query: 87 SAFLVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAP 146
S L +++ LF + L+ L++LR + + + AE H V + F+P ++ +L+
Sbjct: 40 SFLLFYFNILLFFVFLM--LWILRWVKYPKNMIAELKHPVLSS--FSPTVAVAMLVLGID 95
Query: 147 FVAPKTATYL-VLWWVFAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLV--- 202
F+ K +L ++WVF + L I + K + NP + +G +V
Sbjct: 96 FILIKNNLFLGKIFWVFGAIGMFLFSLIVPFYMFKSESIKLDHVNPGWYIPPVGLIVIPI 155
Query: 203 -GAQAAAHMG--WKESAVCL----FSLGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLF 255
G+ H+ W E V + + G YL L + R + LP + P ++
Sbjct: 156 AGSLIMPHLTGVWHELTVLINYFGWGAGFFLYLALLAVVIYRFILHHPLPSAMAPTVWIN 215
Query: 256 FAAPGVASLAWGSIVGGFDTLS-KMLFFLSLFLF---------MSLVCRPTLFRRSMKRF 305
G +A ++V ++ K F++ F+F M+++ ++ +
Sbjct: 216 LGPIGAGIVALINMVNNSPFITIKEPFYIFSFIFWGFGLWWSLMAIIMTLYYVKKLKLPY 275
Query: 306 NVAWWAYSFPVTVLAMASTDYAEEVKG 332
++WWA+ FP+ V +AST ++ G
Sbjct: 276 AMSWWAFIFPLGVY-IASTHLVYKIFG 301
>GLS1_YEAST (P38631) 1,3-beta-glucan synthase component GLS1 (EC
2.4.1.34) (1,3-beta-D-glucan-UDP glucosyltransferase)
(CND1 protein) (CWN53 protein) (FKS1 protein)
(Papulacandin B sensitivity protein 1)
Length = 1876
Score = 37.0 bits (84), Expect = 0.082
Identities = 45/210 (21%), Positives = 79/210 (37%), Gaps = 27/210 (12%)
Query: 94 SLALFTLALLSLLYL----LRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVA 149
SL +F L L++L L + C++ N K + L +G Y F P + W V
Sbjct: 1307 SLQMFMLTLVNLSSLAHESIMCIYDRNKPKTDVLVPIGC-YNFQPAVDW---------VR 1356
Query: 150 PKTATYLVLWWVFAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVI-GNLVGAQAAA 208
T + +++W+ VP+VV ++ G W +RF + + V G + + +
Sbjct: 1357 RYTLSIFIVFWIAFVPIVVQELIERGLW-KATQRFFCHLLSLSPMFEVFAGQIYSSALLS 1415
Query: 209 HMGWKESAVCLFSLGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGS 268
+ + G + F LY R +G ++ A + L +G+
Sbjct: 1416 DLAIGGARYISTGRGFATSRIPFSILYSRFAGS-----------AIYMGARSMLMLLFGT 1464
Query: 269 IVGGFDTLSKMLFFLSLFLFMSLVCRPTLF 298
+ L LS +F V P F
Sbjct: 1465 VAHWQAPLLWFWASLSSLIFAPFVFNPHQF 1494
>MALA_BACST (Q45632) Maltose permease
Length = 394
Score = 34.7 bits (78), Expect = 0.41
Identities = 25/86 (29%), Positives = 44/86 (51%), Gaps = 1/86 (1%)
Query: 23 SREPPSFIVARRLLASLNSVLTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVH 82
+R+P ++A +LA+L V+ L AG +R+ + L ++ I P D LRHV
Sbjct: 74 TRKPVGLLLAALVLAALFGVMYAL-AGSYRLFVVLTVLLSAMQSAIVPLSDSLALRHVHE 132
Query: 83 MVPSSAFLVLWSLALFTLALLSLLYL 108
+ + LW F +A+L++ +L
Sbjct: 133 QGGNYGAIRLWGSLGFAMAVLAVGWL 158
>MAE1_SCHPO (P50537) Malic acid transport protein (Malate permease)
Length = 438
Score = 34.7 bits (78), Expect = 0.41
Identities = 60/320 (18%), Positives = 117/320 (35%), Gaps = 41/320 (12%)
Query: 91 VLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQS-APFVA 149
+++ L +F +L L R + + + +K + HH+ ++ +S + A +
Sbjct: 68 IVYILQIFLFSLFGSCMLFRFIKYPSTIKDSWNHHLEKLFIATCLLSISTFIDMLAIYAY 127
Query: 150 PKTATYLV-----LWWVFAVPVVVLDVKIYGQWFTKGKRFLSTAAN-------PTSQLSV 197
P T ++V L++++ + V + F + TA+ P V
Sbjct: 128 PDTGEWMVWVIRILYYIYVAVSFIYCVMAFFTIFNNHVYTIETASPAWILPIFPPMICGV 187
Query: 198 IGNLVGAQAAAHM--GWKESAVCLFSLGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLF 255
I V + AH + LG YL+LF R RP F+F
Sbjct: 188 IAGAVNSTQPAHQLKNMVIFGILFQGLGFWVYLLLFAVNVLRFFTVGLAKPQDRPGMFMF 247
Query: 256 FAAPGVASLAWGSIVGGF----------DTLSKMLFFLSLFLFMSL-------VCRPTL- 297
P + LA +I G S+ L F+S F+ + + C +
Sbjct: 248 VGPPAFSGLALINIARGAMGSRPYIFVGANSSEYLGFVSTFMAIFIWGLAAWCYCLAMVS 307
Query: 298 -----FRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHV---LMLVLLALSVLV 349
F R+ +F W+A+ FP + + + + + ++ V+L + ++
Sbjct: 308 FLAGFFTRAPLKFACGWFAFIFPNVGFVNCTIEIGKMIDSKAFQMFGHIIGVILCIQWIL 367
Query: 350 SLALTVFTFINSKMLLPDDD 369
+ L V F+ + + P D
Sbjct: 368 LMYLMVRAFLVNDLCYPGKD 387
>YE69_PASMU (Q9CKY2) Hypothetical protein PM1469
Length = 579
Score = 33.9 bits (76), Expect = 0.70
Identities = 27/93 (29%), Positives = 44/93 (47%), Gaps = 9/93 (9%)
Query: 92 LWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFV--- 148
++S + L+ LYL + LF+F + K GV Y+ P+I+ L Q +
Sbjct: 206 IFSPVVILYTLIVYLYLAKILFYFELPKG------GVAYIIMPYIALGLCCQGLRLLLID 259
Query: 149 APKTATYLVLWWVFAVPVVVLDVKIYGQWFTKG 181
A T Y V ++ P+V+L V I+ + T G
Sbjct: 260 AKWTGFYRVFAYLSIAPLVLLWVGIHTRITTYG 292
>NQO8_THETH (Q60019) NADH-quinone oxidoreductase chain 8 (EC
1.6.99.5) (NADH dehydrogenase I, chain 8) (NDH-1, chain
8)
Length = 365
Score = 33.5 bits (75), Expect = 0.91
Identities = 33/133 (24%), Positives = 52/133 (38%), Gaps = 27/133 (20%)
Query: 225 VHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLS 284
+H++ + GG +PVL P ++F K+ FFL
Sbjct: 250 IHFITASALIPTLFLGGWTMPVLEVPYLWMFL---------------------KIAFFLF 288
Query: 285 LFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLA 344
F+++ R T FR + W + FP+ +L T A V + +L L A
Sbjct: 289 FFIWI----RATWFRLRYDQLLRFGWGFLFPLALLWFLVT--ALVVALDLPRTYLLYLSA 342
Query: 345 LSVLVSLALTVFT 357
LS LV L ++T
Sbjct: 343 LSFLVLLGAVLYT 355
>NU5C_MESVI (Q9MUK8) NAD(P)H-quinone oxidoreductase chain 5,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
5) (NADH-plastoquinone oxidoreductase chain 5)
Length = 652
Score = 33.1 bits (74), Expect = 1.2
Identities = 21/79 (26%), Positives = 38/79 (47%), Gaps = 4/79 (5%)
Query: 90 LVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVA 149
++ W + +ALL + +L CL F+ + A ++F ++ LL+ FV
Sbjct: 35 ILRWRYSFLIIALLGISLILSCLILFSQINAT----PSYQWIFQWIVTNNFLLEIGYFVD 90
Query: 150 PKTATYLVLWWVFAVPVVV 168
P TA LV+ A+ V++
Sbjct: 91 PLTAVMLVIVTTVAILVLI 109
>NU1C_LOTJA (Q9BBN9) NAD(P)H-quinone oxidoreductase chain 1,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
1) (NADH-plastoquinone oxidoreductase chain 1)
Length = 363
Score = 33.1 bits (74), Expect = 1.2
Identities = 30/90 (33%), Positives = 45/90 (49%), Gaps = 10/90 (11%)
Query: 230 LFVTLYQRLSGGN-RLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLF 288
LFVT+ L G N +P + FF GV +G+ +G F TL+K LFLF
Sbjct: 269 LFVTVLY-LGGSNISIPYISLFEFFEINKEYGV----FGTTIGIFITLAKTY----LFLF 319
Query: 289 MSLVCRPTLFRRSMKRFNVAWWAYSFPVTV 318
+S++ R TL R M + W + P+++
Sbjct: 320 VSIITRWTLPRLRMDQLLNLGWKFLLPISL 349
>AR61_DROME (Q9VES1) ARL-6 interacting protein-1 homolog
Length = 197
Score = 32.7 bits (73), Expect = 1.5
Identities = 22/60 (36%), Positives = 30/60 (49%)
Query: 71 THDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNY 130
T +K VV V S +LVLW L L + LLSLL ++ L ++ L GVN+
Sbjct: 34 TWEKQYYAGVVFGVISCLYLVLWYLDLSLITLLSLLGVISILLNYAFPMVSRLIFGGVNW 93
>GLS2_YEAST (P40989) 1,3-beta-glucan synthase component GLS2 (EC
2.4.1.34) (1,3-beta-D-glucan-UDP glucosyltransferase)
Length = 1895
Score = 32.3 bits (72), Expect = 2.0
Identities = 24/88 (27%), Positives = 42/88 (47%), Gaps = 14/88 (15%)
Query: 94 SLALFTLALLSLLYL----LRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVA 149
SL +F L L++L L + C++ + + L+ +G Y F P I W V
Sbjct: 1326 SLQMFMLTLVNLHALAHESILCVYDRDKPITDVLYPIGC-YNFHPAIDW---------VR 1375
Query: 150 PKTATYLVLWWVFAVPVVVLDVKIYGQW 177
T + +++W+ VP+VV ++ G W
Sbjct: 1376 RYTLSIFIVFWIAFVPIVVQELIERGLW 1403
>NU1C_SPIOL (Q9M3I6) NAD(P)H-quinone oxidoreductase chain 1,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
1) (NADH-plastoquinone oxidoreductase chain 1)
Length = 365
Score = 32.0 bits (71), Expect = 2.6
Identities = 29/90 (32%), Positives = 43/90 (47%), Gaps = 10/90 (11%)
Query: 230 LFVTLYQRLSGGN-RLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLF 288
LFVT+ L G N +P + FF GV +G+ +G F TL+K LFLF
Sbjct: 271 LFVTVLY-LGGWNLSIPYIFISEFFEINKIDGV----FGTTIGIFITLAKTF----LFLF 321
Query: 289 MSLVCRPTLFRRSMKRFNVAWWAYSFPVTV 318
+ + R TL R M + W + P+++
Sbjct: 322 IPITTRWTLPRLRMDQLLNLGWKFLLPISL 351
>YHHQ_ECOLI (P37619) Hypothetical protein yhhQ
Length = 221
Score = 31.6 bits (70), Expect = 3.5
Identities = 23/87 (26%), Positives = 44/87 (50%), Gaps = 6/87 (6%)
Query: 278 KMLFFLSLFLFMSLVCRPTLFRR--SMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGT-I 334
K LF+LSLF + + L + S+ F+ W A+SFP LA TD + G +
Sbjct: 11 KALFWLSLFHLLVITSSNYLVQLPVSILGFHTTWGAFSFPFIFLA---TDLTVRIFGAPL 67
Query: 335 SHVLMLVLLALSVLVSLALTVFTFINS 361
+ ++ ++ ++L+S ++ ++ S
Sbjct: 68 ARRIIFAVMIPALLISYVISSLFYMGS 94
>Y576_METJA (Q57996) Hypothetical protein MJ0576
Length = 347
Score = 31.6 bits (70), Expect = 3.5
Identities = 51/256 (19%), Positives = 103/256 (39%), Gaps = 27/256 (10%)
Query: 90 LVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVA 149
L +++ LF + L+ ++LR + + A+ H V F P I+ L+ A F+
Sbjct: 50 LFYFNVLLFFIFLVP--WVLRWIMFKDNALADLKHPV--LSAFYPTIAVSCLVLGADFIN 105
Query: 150 PKTATYL--VLWWVFAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLV----G 203
+ V W + A+ + + + + F K + NP + +G +V G
Sbjct: 106 IGHNMFWGGVFWTLGAIGMFLFSLIVPFYMF-KSESIKLDHVNPGWYIPPVGLIVIPIAG 164
Query: 204 AQAAAHMG--WKESAVCL----FSLGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFA 257
+ H+ W E V + + G YL L + R + LP + P ++
Sbjct: 165 SLIMPHLTGVWHELTVLINYFGWGAGFFLYLALLAVVIYRFILHHPLPSAMAPTVWINLG 224
Query: 258 APGVASLAWGSIVGGFDTLS-KMLFFLSLFLF---------MSLVCRPTLFRRSMKRFNV 307
G +A ++V ++ K F++ F+F M+++ ++ + +
Sbjct: 225 PIGAGIVALINMVNNSPFITIKEPFYIFSFIFWGFGLWWSLMAIIMTLYYVKKLKLPYAM 284
Query: 308 AWWAYSFPVTVLAMAS 323
+WWA+ FP+ +S
Sbjct: 285 SWWAFIFPLGAYVASS 300
>MVIN_BUCAI (P57415) Virulence factor mviN homolog
Length = 511
Score = 31.2 bits (69), Expect = 4.5
Identities = 26/96 (27%), Positives = 41/96 (42%), Gaps = 8/96 (8%)
Query: 190 NPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYLVL--FVTLYQRLSGGNRLPVL 247
NP +L + NL+ + S++C L +Y + F ++ +S ++
Sbjct: 119 NPPEKLILSTNLLRIMFPYILLISLSSLCSSILNSWNYFSIPAFSPIFLNIS------II 172
Query: 248 LRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFL 283
VFF F P + LAW I+GG L L FL
Sbjct: 173 FFSVFFSSFFCPSIIVLAWSVIIGGLVQLLYQLPFL 208
>MMR_BACSU (Q00538) Methylenomycin A resistance protein (MMR
peptide)
Length = 466
Score = 31.2 bits (69), Expect = 4.5
Identities = 40/164 (24%), Positives = 68/164 (41%), Gaps = 28/164 (17%)
Query: 138 WFLLLQSAPFVAPKTATYLV--LWWVFAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQL 195
W L+ +A + P LV W + +++V I R LS +S++
Sbjct: 147 WAALVSAASALGPFIGGVLVQLAGWQ---SIFLINVPIGAAALISAYRILSRVPGKSSRV 203
Query: 196 SVIGNLVGAQAAAHM----------GWKESAVCLFSLGMVHYLVLFVTLYQRLSGGNRLP 245
++IG+L+G A + GW+ + + V VLF L + +S + P
Sbjct: 204 NIIGHLLGMMALGFLSYALIQGPSAGWRSPVILVAFTAAVLAFVLF--LLREISA--KTP 259
Query: 246 VLLRPVF--FLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFL 287
+L ++ F AA + L ++ GG +F LSLFL
Sbjct: 260 ILPASLYKNGRFSAAQFIGFLLNFALFGG-------MFMLSLFL 296
>YDHC_ECOLI (P37597) Hypothetical transport protein ydhC
Length = 403
Score = 30.8 bits (68), Expect = 5.9
Identities = 57/240 (23%), Positives = 95/240 (38%), Gaps = 36/240 (15%)
Query: 153 ATYLVLWWVFAVPV----VVLDVKIYGQWFTKGKRFLSTAANPTSQLS-VIGNLVGAQAA 207
AT LVL +V AV V V+ + + ++ + A P LS + L+G+
Sbjct: 95 ATLLVLRFVQAVGVCAAAVIWQALVTDYYPSQKVNRIFAAIMPLVGLSPALAPLLGSWLL 154
Query: 208 AHMGWKESAVCLFSLGMVHYLVLF-----VTLYQRLSGGNRLPVLLRPVFF----LFFAA 258
H W+ LF++ +V L +F G LLR + L +AA
Sbjct: 155 VHFSWQAIFATLFAITVVLILPIFWLKPTTKARNNSQDGLTFTDLLRSKTYRGNVLIYAA 214
Query: 259 PGVASLAWGSIVGGFDTLSKMLFFLSL-----------FLFMSLVCRPTLFRRSMKRFNV 307
+ AW + G LS+M + ++ FL CR L + K+ +
Sbjct: 215 CSASFFAW--LTGSPFILSEMGYSPAVIGLSYVPQTIAFLIGGYGCRAALQKWQGKQL-L 271
Query: 308 AWWAYSFPVTVLAMASTDYAEEVKGTISHV-LMLVLLALSVLVSLALTVFTFINSKMLLP 366
W F V+V+A + G ISHV L+ +L+ V+ ++ + ++ L P
Sbjct: 272 PWLLVLFAVSVIATWAA-------GFISHVSLVEILIPFCVMAIANGAIYPIVVAQALRP 324
>NU1C_OENHO (Q9MTH7) NAD(P)H-quinone oxidoreductase chain 1,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
1) (NADH-plastoquinone oxidoreductase chain 1)
Length = 363
Score = 30.8 bits (68), Expect = 5.9
Identities = 18/53 (33%), Positives = 29/53 (53%), Gaps = 4/53 (7%)
Query: 266 WGSIVGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTV 318
+G+ +G F TL+K LFLF+S+ R TL R M + W + P+++
Sbjct: 301 FGTTIGIFTTLAKTY----LFLFISITTRWTLPRLRMDQLLNLGWKFLLPISL 349
>NU1C_WHEAT (Q95H43) NAD(P)H-quinone oxidoreductase chain 1,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
1) (NADH-plastoquinone oxidoreductase chain 1)
Length = 362
Score = 30.4 bits (67), Expect = 7.7
Identities = 34/119 (28%), Positives = 50/119 (41%), Gaps = 10/119 (8%)
Query: 201 LVGAQAAAHMGWKESAVCLFSLGMVHYLVLFVTLYQRLSGGN-RLPVLLRPVFFLFFAAP 259
LV + G K L S + LFVT+ L G N +P + FF A
Sbjct: 239 LVAGYQTEYSGIKYGLFYLVSYLNLLVSSLFVTVLY-LGGWNLSIPYISFFDFFQMNKAV 297
Query: 260 GVASLAWGSIVGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTV 318
G+ + G + TL+K LFLF+S+ R TL R M + W + P+++
Sbjct: 298 GILEMTMGIFI----TLTKAY----LFLFISITIRWTLPRMRMDQLLNLGWKFLLPISL 348
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.330 0.141 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,781,853
Number of Sequences: 164201
Number of extensions: 1555922
Number of successful extensions: 4760
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 4750
Number of HSP's gapped (non-prelim): 27
length of query: 377
length of database: 59,974,054
effective HSP length: 112
effective length of query: 265
effective length of database: 41,583,542
effective search space: 11019638630
effective search space used: 11019638630
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0195a.11


