

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0193.7
(848 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
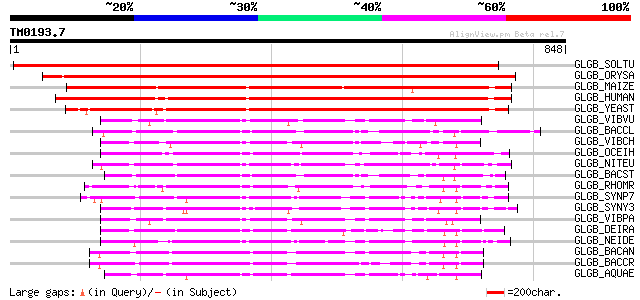
Score E
Sequences producing significant alignments: (bits) Value
GLGB_SOLTU (P30924) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 1238 0.0
GLGB_ORYSA (Q01401) 1,4-alpha-glucan branching enzyme, chloropla... 1202 0.0
GLGB_MAIZE (Q08047) 1,4-alpha-glucan branching enzyme IIB, chlor... 824 0.0
GLGB_HUMAN (Q04446) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 761 0.0
GLGB_YEAST (P32775) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 659 0.0
GLGB_VIBVU (Q8D4P0) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 207 7e-53
GLGB_BACCL (P30537) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 206 2e-52
GLGB_VIBCH (Q9KNE8) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 206 3e-52
GLGB_OCEIH (Q8CZE8) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 205 5e-52
GLGB_NITEU (Q81ZU6) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 201 7e-51
GLGB_BACST (P30538) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 201 9e-51
GLGB_RHOMR (Q93HU3) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 197 1e-49
GLGB_SYNP7 (P16954) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 196 2e-49
GLGB_SYNY3 (P52981) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 192 2e-48
GLGB_VIBPA (Q87FR0) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 191 5e-48
GLGB_DEIRA (Q9RTB7) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 191 5e-48
GLGB_NEIDE (Q9RQI5) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 191 9e-48
GLGB_BACAN (Q81K82) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 189 2e-47
GLGB_BACCR (Q816G6) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 187 1e-46
GLGB_AQUAE (O66936) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 185 5e-46
>GLGB_SOLTU (P30924) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Starch branching enzyme) (Q-enzyme)
Length = 861
Score = 1238 bits (3203), Expect = 0.0
Identities = 583/745 (78%), Positives = 654/745 (87%), Gaps = 3/745 (0%)
Query: 6 SLQSFNIASTAHNSRNKQDLAKQNSVELVLGYRNPKGCNRFSFGSRRSIHERVSTGFKGV 65
S SF+ ++ SRNK Q+S L G + + SR ER+
Sbjct: 15 SFPSFSPKVSSGASRNKICFPSQHSTGLKFGSQERSWDISSTPKSRVRKDERMKHSSAIS 74
Query: 66 AVITDNKSAMSATEEDL--ENIGILHIDPAIKPFKDHFKCRLKRYIDQKKLIEEYEGGLE 123
AV+TD+ S M+ EED+ ENIG+L++DP ++P+ DHF+ R+KRY+DQK LIE+YEG LE
Sbjct: 75 AVLTDDNSTMAPLEEDVKTENIGLLNLDPTLEPYLDHFRHRMKRYVDQKMLIEKYEGPLE 134
Query: 124 EFAQGYLKFGFNREEGGIVYREWAPAAQEAQIIGDFNEWNGSNHPMEKNQFGVWSIKIPD 183
EFAQGYLKFGFNRE+G IVYREWAPAAQE ++IGDFN WNGSNH MEK+QFGVWSI+IPD
Sbjct: 135 EFAQGYLKFGFNREDGCIVYREWAPAAQEDEVIGDFNGWNGSNHMMEKDQFGVWSIRIPD 194
Query: 184 VAGNPAIPHNSRVKFRFRHG-GVWADRIPAWIKYATVDPTKFAAPYDGVYWDPPLSERYQ 242
V P IPHNSRVKFRF+HG GVW DRIPAWIKYAT D TKFAAPYDGVYWDPP SERY
Sbjct: 195 VDSKPVIPHNSRVKFRFKHGNGVWVDRIPAWIKYATADATKFAAPYDGVYWDPPPSERYH 254
Query: 243 FKYPRPPKPKAPRIYEAHVGMSSSEPRINSYKEFADDILPRIRANNYNTVQLMAVMEHSY 302
FKYPRPPKP+APRIYEAHVGMSSSEPR+NSY+EFADD+LPRI+ANNYNTVQLMA+MEHSY
Sbjct: 255 FKYPRPPKPRAPRIYEAHVGMSSSEPRVNSYREFADDVLPRIKANNYNTVQLMAIMEHSY 314
Query: 303 YASFGYHVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDV 362
Y SFGYHVTNFFAVSSR G PEDLKYLIDKAHSLGL VL+DVVHSHASNNVTDGLNGFD+
Sbjct: 315 YGSFGYHVTNFFAVSSRYGNPEDLKYLIDKAHSLGLQVLVDVVHSHASNNVTDGLNGFDI 374
Query: 363 GQVSQESYFHTGDRGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSM 422
GQ SQESYFH G+RGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEE+ FDGFRFDG+TSM
Sbjct: 375 GQGSQESYFHAGERGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEYNFDGFRFDGITSM 434
Query: 423 LYHHHGVNIAFSGDYNEYFSEATDVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGR 482
LY HHG+N+ F+G+YNEYFSEATDVDAVVYLMLAN+LIH I PDATVIAEDVSGMPGLGR
Sbjct: 435 LYVHHGINMGFTGNYNEYFSEATDVDAVVYLMLANNLIHKIFPDATVIAEDVSGMPGLGR 494
Query: 483 PISEVGIGFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRYSEKCVSYAESHDQ 542
P+SE GIGFDYRLAMAIPDKWIDYLKNK D +WSMKE++ SLTNRRY+EKC++YAESHDQ
Sbjct: 495 PVSEGGIGFDYRLAMAIPDKWIDYLKNKNDEDWSMKEVTSSLTNRRYTEKCIAYAESHDQ 554
Query: 543 SIVGDKTFSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMSLGGEGYLNFMGNE 602
SIVGDKT +FLLMD+E+YSGMSCL DASP ++RGIALHKMIHF TM+LGGEGYLNFMGNE
Sbjct: 555 SIVGDKTIAFLLMDKEMYSGMSCLTDASPVVDRGIALHKMIHFFTMALGGEGYLNFMGNE 614
Query: 603 FGHPEWIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSFLASTKQ 662
FGHPEWIDFPREGN WSY+KCRRQW+L D++HLRYKFMNAFD+AMN LD+KFSFLAS KQ
Sbjct: 615 FGHPEWIDFPREGNNWSYDKCRRQWNLADSEHLRYKFMNAFDRAMNSLDEKFSFLASGKQ 674
Query: 663 IVSSTNEEDKVIVFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREFGGHGR 722
IVSS ++++KV+VFERGDLVFVFNFHP+ TYEGYKVGCDLPGKYRVALDSDA EFGGHGR
Sbjct: 675 IVSSMDDDNKVVVFERGDLVFVFNFHPKNTYEGYKVGCDLPGKYRVALDSDAWEFGGHGR 734
Query: 723 VGHNVDHFTAPEGIPGVPESNFNNR 747
GH+VDHFT+PEGIPGVPE+NFN R
Sbjct: 735 TGHDVDHFTSPEGIPGVPETNFNGR 759
>GLGB_ORYSA (Q01401) 1,4-alpha-glucan branching enzyme, chloroplast
precursor (EC 2.4.1.18) (Starch branching enzyme)
(Q-enzyme)
Length = 820
Score = 1202 bits (3110), Expect = 0.0
Identities = 554/723 (76%), Positives = 639/723 (87%), Gaps = 2/723 (0%)
Query: 51 RRSIHERVSTGFKGVAVITDNKSAMSATEEDLENIGILHIDPAIKPFKDHFKCRLKRYID 110
RRS +V T F A NK+ ++ EE ++++ I +DP ++ FKDHF R+KRY+D
Sbjct: 43 RRSWPGKVKTNFSVPATARKNKTMVTVVEE-VDHLPIYDLDPKLEEFKDHFNYRIKRYLD 101
Query: 111 QKKLIEEYEGGLEEFAQGYLKFGFNREEGGIVYREWAPAAQEAQIIGDFNEWNGSNHPME 170
QK LIE++EGGLEEF++GYLKFG N +G +YREWAPAAQEAQ+IG+FN WNG+ H ME
Sbjct: 102 QKCLIEKHEGGLEEFSKGYLKFGINTVDGATIYREWAPAAQEAQLIGEFNNWNGAKHKME 161
Query: 171 KNQFGVWSIKIPDVAGNPAIPHNSRVKFRFRHGG-VWADRIPAWIKYATVDPTKFAAPYD 229
K++FG+WSIKI V G PAIPHNS+VKFRFRHGG W DRIPAWI+YAT D +KF APYD
Sbjct: 162 KDKFGIWSIKISHVNGKPAIPHNSKVKFRFRHGGGAWVDRIPAWIRYATFDASKFGAPYD 221
Query: 230 GVYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEPRINSYKEFADDILPRIRANNY 289
GV+WDPP ERY FK+PRPPKP APRIYEAHVGMS EP +++Y+EFAD++LPRIRANNY
Sbjct: 222 GVHWDPPACERYVFKHPRPPKPDAPRIYEAHVGMSGEEPEVSTYREFADNVLPRIRANNY 281
Query: 290 NTVQLMAVMEHSYYASFGYHVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHA 349
NTVQLMA+MEHSYYASFGYHVTNFFAVSSRSGTPEDLKYL+DKAHSLGL VLMDVVHSHA
Sbjct: 282 NTVQLMAIMEHSYYASFGYHVTNFFAVSSRSGTPEDLKYLVDKAHSLGLRVLMDVVHSHA 341
Query: 350 SNNVTDGLNGFDVGQVSQESYFHTGDRGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEF 409
SNNVTDGLNG+DVGQ + ESYFHTGDRGYHKLWDSRLFNYANWEVLRFLLSNLR+W++EF
Sbjct: 342 SNNVTDGLNGYDVGQNTHESYFHTGDRGYHKLWDSRLFNYANWEVLRFLLSNLRYWMDEF 401
Query: 410 KFDGFRFDGVTSMLYHHHGVNIAFSGDYNEYFSEATDVDAVVYLMLANSLIHNILPDATV 469
FDGFRFDGVTSMLYHHHG+N F+G+Y EYFS TDVDA+VY+MLAN L+H +LP+AT+
Sbjct: 402 MFDGFRFDGVTSMLYHHHGINKGFTGNYKEYFSLDTDVDAIVYMMLANHLMHKLLPEATI 461
Query: 470 IAEDVSGMPGLGRPISEVGIGFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRY 529
+AEDVSGMP L RP+ E G+GFD+RLAMAIPD+WIDYLKNK+D +WSM EI +LTNRRY
Sbjct: 462 VAEDVSGMPVLCRPVDEGGVGFDFRLAMAIPDRWIDYLKNKEDRKWSMSEIVQTLTNRRY 521
Query: 530 SEKCVSYAESHDQSIVGDKTFSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMS 589
+EKC++YAESHDQSIVGDKT +FLLMD+E+Y+GMS L ASPTI RGIAL KMIHFITM+
Sbjct: 522 TEKCIAYAESHDQSIVGDKTIAFLLMDKEMYTGMSDLQPASPTINRGIALQKMIHFITMA 581
Query: 590 LGGEGYLNFMGNEFGHPEWIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNL 649
LGG+GYLNFMGNEFGHPEWIDFPREGN WSY+KCRRQWSLVDTDHLRYK+MNAFD+AMN
Sbjct: 582 LGGDGYLNFMGNEFGHPEWIDFPREGNNWSYDKCRRQWSLVDTDHLRYKYMNAFDQAMNA 641
Query: 650 LDDKFSFLASTKQIVSSTNEEDKVIVFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVA 709
L+++FSFL+S+KQIVS NE+DKVIVFERGDLVFVFNFHP TY+GYKVGCDLPGKYRVA
Sbjct: 642 LEEEFSFLSSSKQIVSDMNEKDKVIVFERGDLVFVFNFHPNKTYKGYKVGCDLPGKYRVA 701
Query: 710 LDSDAREFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVVYYRVDES 769
LDSDA FGGHGRVGH+VDHFT+PEG+PGVPE+NFNNRPNSFK+LSPPRTCV YYRVDE
Sbjct: 702 LDSDALVFGGHGRVGHDVDHFTSPEGMPGVPETNFNNRPNSFKVLSPPRTCVAYYRVDED 761
Query: 770 QEE 772
+EE
Sbjct: 762 REE 764
>GLGB_MAIZE (Q08047) 1,4-alpha-glucan branching enzyme IIB,
chloroplast precursor (EC 2.4.1.18) (Starch branching
enzyme IIB) (Q-enzyme)
Length = 799
Score = 824 bits (2129), Expect = 0.0
Identities = 403/693 (58%), Positives = 494/693 (71%), Gaps = 26/693 (3%)
Query: 87 ILHIDPAIKPFKDHFKCRLKRYIDQKKLIEEYEGGLEEFAQGYLKFGFNREEGGIVYREW 146
I IDP ++ +K H + R Y + I+E+EGGLE F++ Y KFGFN GI YREW
Sbjct: 121 IFQIDPMLQGYKYHLEYRYSLYRRIRSDIDEHEGGLEAFSRSYEKFGFNASAEGITYREW 180
Query: 147 APAAQEAQIIGDFNEWNGSNHPMEKNQFGVWSIKIPDVA-GNPAIPHNSRVKFRFRHGGV 205
AP A A ++GD N W+ + M KN+FGVW I +P+ A G IPH SRVK R
Sbjct: 181 APGAFSAALVGDVNNWDPNADRMSKNEFGVWEIFLPNNADGTSPIPHGSRVKVRMDTPSG 240
Query: 206 WADRIPAWIKYATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSS 265
D IPAWIKY+ P + PYDG+Y+DPP +Y F++ +P +PK+ RIYE HVGMSS
Sbjct: 241 IKDSIPAWIKYSVQAPGEI--PYDGIYYDPPEEVKYVFRHAQPKRPKSLRIYETHVGMSS 298
Query: 266 SEPRINSYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGYHVTNFFAVSSRSGTPED 325
EP+IN+Y F D++LPRI+ YN VQ+MA+ EHSYY SFGYHVTNFFA SSR GTPED
Sbjct: 299 PEPKINTYVNFRDEVLPRIKKLGYNAVQIMAIQEHSYYGSFGYHVTNFFAPSSRFGTPED 358
Query: 326 LKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFHTGDRGYHKLWDSR 385
LK LID+AH LGL VLMDVVHSHAS+N DGLNGFD + YFH+G RG+H +WDSR
Sbjct: 359 LKSLIDRAHELGLLVLMDVVHSHASSNTLDGLNGFDG---TDTHYFHSGPRGHHWMWDSR 415
Query: 386 LFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYHHHGVNIAFSGDYNEYFSEAT 445
LFNY NWEVLRFLLSN RWWLEE+KFDGFRFDGVTSM+Y HHG+ + F+G++NEYF AT
Sbjct: 416 LFNYGNWEVLRFLLSNARWWLEEYKFDGFRFDGVTSMMYTHHGLQVTFTGNFNEYFGFAT 475
Query: 446 DVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPISEVGIGFDYRLAMAIPDKWID 505
DVDAVVYLML N LIH + P+A I EDVSGMP P+ + G+GFDYR+ MA+ DKWID
Sbjct: 476 DVDAVVYLMLVNDLIHGLYPEAVTIGEDVSGMPTFALPVHDGGVGFDYRMHMAVADKWID 535
Query: 506 YLKNKKDHEWSMKEISLSLTNRRYSEKCVSYAESHDQSIVGDKTFSFLLMDEEIYSGMSC 565
LK + D W M +I +LTNRR+ EKCV+YAESHDQ++VGDKT +F LMD+++Y M+
Sbjct: 536 LLK-QSDETWKMGDIVHTLTNRRWLEKCVTYAESHDQALVGDKTIAFWLMDKDMYDFMAL 594
Query: 566 LADASPTIERGIALHKMIHFITMSLGGEGYLNFMGNEFGHPEWIDFPR-----------E 614
++PTI+RGIALHKMI ITM LGGEGYLNFMGNEFGHPEWIDFPR
Sbjct: 595 DRPSTPTIDRGIALHKMIRLITMGLGGEGYLNFMGNEFGHPEWIDFPRGPQRLPSGKFIP 654
Query: 615 GNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSFLASTKQIVSSTNEEDKVI 674
GN SY+KCRR++ L D D+LRY M FD+AM L+ K+ F+ S Q +S +EEDKVI
Sbjct: 655 GNNNSYDKCRRRFDLGDADYLRYHGMQEFDQAMQHLEQKYEFMTSDHQYISRKHEEDKVI 714
Query: 675 VFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREFGGHGRVGHNVDHFTAPE 734
VFE+GDLVFVFNFH +Y Y++GC PG Y+V LDSDA FGG R+ H +HFTA
Sbjct: 715 VFEKGDLVFVFNFHCNNSYFDYRIGCRKPGVYKVVLDSDAGLFGGFSRIHHAAEHFTA-- 772
Query: 735 GIPGVPESNFNNRPNSFKILSPPRTCVVYYRVD 767
+ + +NRP SF + +P RTCVVY V+
Sbjct: 773 ------DCSHDNRPYSFSVYTPSRTCVVYAPVE 799
>GLGB_HUMAN (Q04446) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (Brancher enzyme)
Length = 702
Score = 761 bits (1965), Expect = 0.0
Identities = 374/702 (53%), Positives = 486/702 (68%), Gaps = 19/702 (2%)
Query: 70 DNKSAMSATEEDLENIG-ILHIDPAIKPFKDHFKCRLKRYIDQKKLIEEYEGGLEEFAQG 128
D ++A++A D+ + +L IDP +KP+ F+ R K++ K I E EGG+++F++G
Sbjct: 13 DYEAALNAALADVPELARLLEIDPYLKPYAVDFQRRYKQFSQILKNIGENEGGIDKFSRG 72
Query: 129 YLKFGFNR-EEGGIVYREWAPAAQEAQIIGDFNEWNGSNHPMEKNQFGVWSIKIPDVAGN 187
Y FG +R +GG+ +EWAP A+ + GDFN WN ++P +K +G W + IP
Sbjct: 73 YESFGVHRCADGGLYSKEWAPGAEGVFLTGDFNGWNPFSYPYKKLDYGKWELYIPPKQNK 132
Query: 188 PA-IPHNSRVKFRFRH-GGVWADRIPAWIKYATVDPTKFAAPYDGVYWDPPLSERYQFKY 245
+PH S++K G RI W KY + YD ++WDP S Y+FK+
Sbjct: 133 SVLVPHGSKLKVVITSKSGEILYRISPWAKYVVREGDN--VNYDWIHWDPEHS--YEFKH 188
Query: 246 PRPPKPKAPRIYEAHVGMSSSEPRINSYKEFADDILPRIRANNYNTVQLMAVMEHSYYAS 305
RP KP++ RIYE+HVG+SS E ++ SYK F ++LPRI+ YN +QLMA+MEH+YYAS
Sbjct: 189 SRPKKPRSLRIYESHVGISSHEGKVASYKHFTCNVLPRIKGLGYNCIQLMAIMEHAYYAS 248
Query: 306 FGYHVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQV 365
FGY +T+FFA SSR GTPE+L+ L+D AHS+G+ VL+DVVHSHAS N DGLN FD
Sbjct: 249 FGYQITSFFAASSRYGTPEELQELVDTAHSMGIIVLLDVVHSHASKNSADGLNMFDG--- 305
Query: 366 SQESYFHTGDRGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYH 425
+ YFH+G RG H LWDSRLF Y++WEVLRFLLSN+RWWLEE++FDGFRFDGVTSMLYH
Sbjct: 306 TDSCYFHSGPRGTHDLWDSRLFAYSSWEVLRFLLSNIRWWLEEYRFDGFRFDGVTSMLYH 365
Query: 426 HHGVNIAFSGDYNEYFSEATDVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPIS 485
HHGV FSGDY+EYF D DA+ YLMLAN L+H + PD+ IAEDVSGMP L PIS
Sbjct: 366 HHGVGQGFSGDYSEYFGLQVDEDALTYLMLANHLVHTLCPDSITIAEDVSGMPALCSPIS 425
Query: 486 EVGIGFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRYSEKCVSYAESHDQSIV 545
+ G GFDYRLAMAIPDKWI LK KD +W+M +I +LTNRRY EKC++YAESHDQ++V
Sbjct: 426 QGGGGFDYRLAMAIPDKWIQLLKEFKDEDWNMGDIVYTLTNRRYLEKCIAYAESHDQALV 485
Query: 546 GDKTFSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMSLGGEGYLNFMGNEFGH 605
GDK+ +F LMD E+Y+ MS L +P I+RGI LHKMI IT LGGEGYLNFMGNEFGH
Sbjct: 486 GDKSLAFWLMDAEMYTNMSVLTPFTPVIDRGIQLHKMIRLITHGLGGEGYLNFMGNEFGH 545
Query: 606 PEWIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSFLASTKQIVS 665
PEW+DFPR+GN SY RRQ+ L D D LRYKF+N FD+ MN L++++ +LA+ + VS
Sbjct: 546 PEWLDFPRKGNNESYHYARRQFHLTDDDLLRYKFLNNFDRDMNRLEERYGWLAAPQAYVS 605
Query: 666 STNEEDKVIVFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREFGGHGRVGH 725
+E +K+I FER L+F+FNFHP +Y Y+VG LPGK+++ LDSDA E+GGH R+ H
Sbjct: 606 EKHEGNKIIAFERAGLLFIFNFHPSKSYTDYRVGTALPGKFKIVLDSDAAEYGGHQRLDH 665
Query: 726 NVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVVYYRVD 767
+ D F+ N RP S + P R ++ VD
Sbjct: 666 STDFFS--------EAFEHNGRPYSLLVYIPSRVALILQNVD 699
>GLGB_YEAST (P32775) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme)
Length = 704
Score = 659 bits (1700), Expect = 0.0
Identities = 359/707 (50%), Positives = 467/707 (65%), Gaps = 47/707 (6%)
Query: 86 GILHIDPAIKPFKDHFKCRLKRYIDQKKLIE--------EYEGGLEEFAQ-GYLKFGF-- 134
G + DP +KPF D R RY+ K L + Y+ L +FA+ Y +G
Sbjct: 10 GAVEFDPWLKPFADVLSER--RYLADKWLYDITHATPDGSYQS-LSKFARDSYKSYGLHA 66
Query: 135 NREEGGIVYREWAPAAQEAQIIGDFNEWNGSNHPME-KNQFGVWSIKI-PDVAGNPAIPH 192
N E I Y+EWAP A+ A ++GDFN W+ ++H ++ K++FG ++I + P G+ AIPH
Sbjct: 67 NPETKEITYKEWAPNAERAFLVGDFNNWDTTSHELKNKDEFGNFTITLHPLPNGDFAIPH 126
Query: 193 NSRVKFRF-RHGGVWADRIPAWIKYATVDPTK-----FAAPYDGVYWDPPLSERYQFKYP 246
+S++K F G R+PAWI AT P+K F Y+G +W+P Y+F +P
Sbjct: 127 DSKIKVMFILPDGSKIFRLPAWITRAT-QPSKETSKQFGPAYEGRFWNP--ENPYKFVHP 183
Query: 247 RPPKPKAP---RIYEAHVGMSSSEPRINSYKEFADDILPRIRANNYNTVQLMAVMEHSYY 303
RP ++ RIYEAHVG+SS EP+I +YKEF + +LPRI+ Y+ +QLMA+MEH+YY
Sbjct: 184 RPKFSESVDSLRIYEAHVGISSPEPKITTYKEFTEKVLPRIKYLGYDAIQLMAIMEHAYY 243
Query: 304 ASFGYHVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVG 363
ASFGY VTNFFA SSR GTPE+LK LID AHS+G+ VL+DVVHSHAS NV DGLN FD
Sbjct: 244 ASFGYQVTNFFAASSRFGTPEELKELIDTAHSMGILVLLDVVHSHASKNVEDGLNMFDG- 302
Query: 364 QVSQESYFHT--GDRGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTS 421
S YFH+ RG H LWDSRLFNY +EV RFLL+NL ++++ ++FDGFRFDGVTS
Sbjct: 303 --SDHQYFHSISSGRGEHPLWDSRLFNYGKFEVQRFLLANLAFYVDVYQFDGFRFDGVTS 360
Query: 422 MLYHHHGVNI--AFSGDYNEYFSEA---TDVDAVVYLMLANSLIHNILPD-ATVIAEDVS 475
MLY HHGV +FSGDYNEY S D +A+ YLMLAN L+H +LP+ A +AEDVS
Sbjct: 361 MLYVHHGVGAGGSFSGDYNEYLSRDRSFVDHEALAYLMLANDLVHEMLPNLAVTVAEDVS 420
Query: 476 GMPGLGRPISEVGIGFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRYSEKCVS 535
G P L P S G GFDYRLAMA+PD WI +K KKD EW M I +LTNRRY EK V+
Sbjct: 421 GYPTLCLPRSIGGTGFDYRLAMALPDMWIKLIKEKKDDEWEMGSIVYTLTNRRYGEKVVA 480
Query: 536 YAESHDQSIVGDKTFSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMSLGGEGY 595
Y ESHDQ++VGDKT +F LMD +Y+ M+ L + S I+RGIALHKMI IT SLGGE Y
Sbjct: 481 YCESHDQALVGDKTLAFWLMDAAMYTDMTVLKEPSIVIDRGIALHKMIRLITHSLGGEAY 540
Query: 596 LNFMGNEFGHPEWIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFS 655
LNF GNEFGHPEW+DFP NG SY+ RRQ++L D LRY+ +N FD++M L + +
Sbjct: 541 LNFEGNEFGHPEWLDFPNVNNGDSYKYARRQFNLADDPLLRYQNLNEFDRSMQLCEKRHK 600
Query: 656 FLASTKQIVSSTNEEDKVIVFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAR 715
+L + + VS +E DK+IVFER +L+F+FNFHP +Y Y+VG + G Y + L+SD
Sbjct: 601 WLNTKQAYVSLKHEGDKMIVFERNNLLFIFNFHPTNSYSDYRVGVEKAGTYHIVLNSDRA 660
Query: 716 EFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVV 762
EFGGH R+ + + FT + +NNR N ++ P R +V
Sbjct: 661 EFGGHNRINESSEFFTT--------DLEWNNRKNFLQVYIPSRVALV 699
>GLGB_VIBVU (Q8D4P0) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 715
Score = 207 bits (528), Expect = 7e-53
Identities = 158/604 (26%), Positives = 272/604 (44%), Gaps = 72/604 (11%)
Query: 140 GIVYREWAPAAQEAQIIGDFNEWNGSNHPMEKNQFGVWSIKIPDVAGNPAIPHNSRVKFR 199
GI + +AP A ++G FN+W+G HPM++ +G+W + IPD+A V ++
Sbjct: 126 GIRFLVYAPHATAVSLVGSFNDWDGRRHPMQRLDYGIWGLFIPDLAEG--------VSYK 177
Query: 200 FRHGGVWADRIP----AWIKYATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKPKAPR 255
F G + +P W YA P+ + YD + ++Q + + +A
Sbjct: 178 FEMKGPKGEGLPHKADPWGFYAEQYPSFASVTYDHARYQWQ-DAQWQTRPVTEKRKEALS 236
Query: 256 IYEAHVGM--SSSEPRINSYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGYHVTNF 313
YE H G + + +Y+E A +++P + Y V+LM V EH +Y S+GY
Sbjct: 237 FYELHAGSWKRNEQGEFLNYRELAAELVPYLVDMGYTHVELMPVSEHPFYGSWGYQPVGL 296
Query: 314 FAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFHT 373
FA +SR G+P+D K+ +D H G+ V++D V +H ++ GL FD FH
Sbjct: 297 FAPTSRYGSPDDFKFFVDACHQAGIGVVLDWVPAHFPSD-DHGLANFD-----GTPLFHD 350
Query: 374 GD--RGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLY----HHH 427
D RG+H+ W+S +++ +V RFL+SN +W E+F DG R D V SMLY H
Sbjct: 351 PDPRRGWHQDWNSFIYDLGREQVRRFLVSNALYWFEQFHIDGIRVDAVASMLYLDYSRSH 410
Query: 428 GVNIAFSGDYNEYFSEATDVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPISEV 487
G I NE + DA+ L N ++ P+A IAE+ + PG+ P
Sbjct: 411 GQWIPNMDGGNENY------DAIATLKWMNEEVYKYFPNAMTIAEESTAFPGVSAPTFMG 464
Query: 488 GIGFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRYSEKCVSYAESHDQSIVGD 547
G+GF ++ M + Y+K + H R+Y +++ + S
Sbjct: 465 GLGFGFKWNMGWMHDSLSYIKEEPVH-------------RKYHHNTLTFPLVYAHS---- 507
Query: 548 KTFSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMSLGGEGYLNFMGNEFGHPE 607
+ + L +E+ G + + P E + +F M LNFMG E G
Sbjct: 508 ENYVLSLSHDEVVYGKGSIHNKMPGDEWQQTANLRAYFGYMYGQPGKKLNFMGAEIG--- 564
Query: 608 WIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMN-------LLDDKFSFLAST 660
+ W+++ + QW L+D R++ + A + +N L D+ A
Sbjct: 565 ------QTAEWNHDD-QLQWFLLDFP--RHQGVQALTRDLNHLYRNQAALHDQDCIPAGF 615
Query: 661 KQIVSSTNEEDKVI---VFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREF 717
+ + E+ + + E G+ + V + + +++G G+Y++ L++D ++
Sbjct: 616 EWRLQDAAEQSIIAHERISEAGERILVVSNFTPVPRDEFRLGVPNKGRYQLLLNTDDSKY 675
Query: 718 GGHG 721
G G
Sbjct: 676 AGSG 679
>GLGB_BACCL (P30537) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 666
Score = 206 bits (524), Expect = 2e-52
Identities = 203/707 (28%), Positives = 311/707 (43%), Gaps = 89/707 (12%)
Query: 127 QGYLKFGFNREEGGIV----YREWAPAAQEAQIIGDFNEWNGSNHPMEK-NQFGVWSIKI 181
Q Y FG + GG + WAP A+E +++G FN+WNG+N P+ K N GVW+I +
Sbjct: 21 QSYELFGAHVIRGGGAVGTRFCVWAPHAREVRLVGSFNDWNGTNSPLTKVNDEGVWTIVV 80
Query: 182 PDVAGNPAIPHNSRVKFRFRHGGVWADRIPAWIKYATVDPTKFAAPYD--GVYW-DPPLS 238
P+ H + + G V P + Y+ + P + YD G W D P
Sbjct: 81 PENLEG----HLYKYEIITPDGRVLLKADP-YAFYSELRPHTASIVYDLKGYEWNDSPWQ 135
Query: 239 ERYQFK--YPRPPKPKAPRIYEAHVGMSSSEP--RINSYKEFADDILPRIRANNYNTVQL 294
+ + K Y +P IYE H G +P R +Y+E AD+++P + + ++L
Sbjct: 136 RKKRRKRIYDQPMV-----IYELHFGSWKKKPDGRFYTYREMADELIPYVLERGFTHIEL 190
Query: 295 MAVMEHSYYASFGYHVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVT 354
+ ++EH S+GY T +++V+SR GTP D Y +D+ H GL V++D V H +
Sbjct: 191 LPLVEHPLDRSWGYQGTGYYSVTSRYGTPHDFMYFVDRCHQAGLGVIIDWVPGHFCKD-A 249
Query: 355 DGLNGFDVGQVSQESYFHTGDRGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGF 414
GL FD G + E Y + DR + +W + F+ EV FL+SN +WLE + DGF
Sbjct: 250 HGLYMFD-GAPTYE-YANEKDRENY-VWGTANFDLGKPEVRSFLISNALFWLEYYHVDGF 306
Query: 415 RFDGVTSMLYHHHGVNIAFSGDYNEYFSEATDVDAVVYLMLANSLIHNILPDATVIAEDV 474
R D V +MLY + + + Y AV +L N + P+ +IAED
Sbjct: 307 RVDAVANMLYWPNNDRL-YENPY-----------AVEFLRQLNEAVFAYDPNVWMIAEDS 354
Query: 475 SGMPGLGRPISEVGIGFDYRLAMAIPDKWIDYLKN-KKDHEWSMKEISLSLTNRRYSEKC 533
+ P + P + G+GF+Y+ M + + Y++ + +++ ++S SL YSE
Sbjct: 355 TDWPRVTAPTYDGGLGFNYKWNMGWMNDMLKYMETPPHERKYAHNQVSFSLL-YAYSENF 413
Query: 534 VSYAESHDQSIVGDKTFSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMSLGGE 593
+ SHD+ + G K S L + E A ++++ M+ G+
Sbjct: 414 I-LPFSHDEVVHGKK---------------SLLNKMPGSYEEKFAQLRLLYGYMMAHPGK 457
Query: 594 GYLNFMGNEFGHPEWIDFPREGNG--WSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLD 651
L FMG+EF + F E + + +E R+ V YK F L
Sbjct: 458 KLL-FMGSEFAQFDEWKFAEELDWVLFDFELHRKMDEYVKQLIACYKRYKPF---YELDH 513
Query: 652 DKFSFLASTKQIVSSTNEEDKVIVFER-----GD-LVFVFNFHPETTYEGYKVGCDLPGK 705
D F + + N E + F R GD LV V NF Y+ YKV L
Sbjct: 514 DPRGF-----EWIDVHNAEQSIFSFIRRGKKEGDVLVIVCNF-TNQAYDDYKVSVPLLAP 567
Query: 706 YRVALDSDAREFGGHGRV-GHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVVYY 764
YR L+SDA EFGG G V G + F+ P F+ +P ++ PP +
Sbjct: 568 YREVLNSDAAEFGGSGHVNGKRLPAFSEP----------FHGKPYHVRMTIPPFGISILR 617
Query: 765 RVDESQEENSISNLVGVQETSTAADIVANIPDGSSASKEREVSNFNW 811
V + E V A A++ D K RE S W
Sbjct: 618 PVQKRGERKQNEEEVHRHVIGRRARKPASLAD----EKHRETSRAVW 660
>GLGB_VIBCH (Q9KNE8) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 666
Score = 206 bits (523), Expect = 3e-52
Identities = 171/610 (28%), Positives = 275/610 (45%), Gaps = 88/610 (14%)
Query: 140 GIVYREWAPAAQEAQIIGDFNEWNGSNHPMEKNQFGVWSIKIPDVAGNPAIPHNSRVKFR 199
G+ + +AP A +IG FN W+G HPM++ +G+W I IP +P ++ KF
Sbjct: 75 GVRFLVYAPHAAACSLIGAFNHWDGRRHPMQRLDYGIWGIFIP------GLPEGTQYKFE 128
Query: 200 FR--HGGVWADRIPAWIKYATVDPTKFAAPYDGVYWDPPLSERYQFK----YPRPPKPK- 252
+ HG + W YA P+ + YD RYQ++ RP K
Sbjct: 129 LKGPHGEGLPHKADPWGFYAEQYPSFASVTYD--------HRRYQWQDTAWQQRPVTEKR 180
Query: 253 --APRIYEAHVGM--SSSEPRINSYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGY 308
A YE HVG +Y+E AD ++P + Y V+LM V EH +Y S+GY
Sbjct: 181 KQALSFYELHVGSWKRGENGEFLNYRELADQLVPYLVEMGYTHVELMPVAEHPFYGSWGY 240
Query: 309 HVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQE 368
FA +SR G+P+D KY +D H G+ V++D V +H ++ + GL FD
Sbjct: 241 QPVGLFAPTSRYGSPDDFKYFVDLCHQAGIGVVLDWVPAHFPSD-SHGLANFD-----GT 294
Query: 369 SYFHTGD--RGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYHH 426
FH D RG+H+ W+S +++ V RFL++N +W E F DG R D V SMLY
Sbjct: 295 PLFHDPDPRRGWHQDWNSYIYDLGREHVRRFLVANALYWFEMFHIDGIRVDAVASMLY-- 352
Query: 427 HGVNIAFSGDYNEYFSEA----TDVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGR 482
+ +S ++++ + DA+ N ++ P+A IAE+ + PG+
Sbjct: 353 ----LDYSRSHDQWIPNVDGGRENYDAIATFKWMNEEVYKHFPNAMTIAEESTAFPGVSA 408
Query: 483 PISEVGIGFDYRLAMAIPDKWIDYLKNKKDH-EWSMKEISLSLTNRRYSEKCVSYAESHD 541
P G+GF ++ M + Y+K H ++ ++ L +SE V + SHD
Sbjct: 409 PTFMGGLGFGFKWNMGWMHDSLSYIKEDPVHRKYHHNTLTFPLI-YAFSENYV-LSLSHD 466
Query: 542 QSIVGDKTFSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMSLGGEG-YLNFMG 600
+ + G ++ + + +E + A ++ G G LNFMG
Sbjct: 467 EVVYGKRSLMYKMPGDEWQQTANLRA-----------------YLGYMYGQPGKKLNFMG 509
Query: 601 NEFGHPEWIDFPREGN-GWSYEKCRR----QWSLVDTDHLRYKFMNAFDKAMNLLD-DKF 654
E G + ++ +G W + R Q + D +HL A++ LD D
Sbjct: 510 TELG--QTAEWDHDGQLQWFLTQFERHAGIQRLVRDLNHLYQA-----QTALHQLDCDPR 562
Query: 655 SFLASTKQIVSSTNEEDKVIVFERGD-----LVFVFNFHPETTYEGYKVGCDLPGKYRVA 709
F + N + VI ER D ++ + NF P E +++G GKYR+
Sbjct: 563 GF-----EWRLQDNADLSVIAHERMDEAGNRVLVITNFTPVPQQE-FRLGVPKTGKYRLL 616
Query: 710 LDSDAREFGG 719
L++DA+++ G
Sbjct: 617 LNTDAKQYNG 626
>GLGB_OCEIH (Q8CZE8) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 637
Score = 205 bits (521), Expect = 5e-52
Identities = 167/644 (25%), Positives = 292/644 (44%), Gaps = 74/644 (11%)
Query: 140 GIVYREWAPAAQEAQIIGDFNEWNGSNHPMEK-NQFGVWSIKIPDVAGNPAIPHN--SRV 196
G + WAP A + ++GDFN W ++HP+EK G+W I D+ + ++ S
Sbjct: 36 GYRFAVWAPNALKVCVVGDFNNWEENSHPLEKFTDEGLWGGFIADIPPATSYKYHICSSE 95
Query: 197 KFRFRHGGVWADRIPAWIKYATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKPKAPRI 256
+A + K A+V P + W + +R + P I
Sbjct: 96 DTSILKADPFATQAERRPKTASVIPAANGYQWSDDQW---IEQRNTYD----PYSSPISI 148
Query: 257 YEAHVGM--SSSEPRINSYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGYHVTNFF 314
YE H+G + + + SY+E A ++P +++ Y ++L+ + EH + S+GY +T +F
Sbjct: 149 YEVHLGTWKKTLKKQFLSYRELATQLIPYVKSLGYTHIELLPINEHPFDRSWGYQITGYF 208
Query: 315 AVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFHTG 374
AV+SR G P D KY ID+ H + V++D V H + GL FD + + +
Sbjct: 209 AVTSRYGNPSDFKYFIDQCHQHQIGVILDWVPGHFCKD-DFGLRQFDGAPLYE---YRDP 264
Query: 375 DRGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYHHHGVNIAFS 434
+ K W + F+Y EV FL+SN +WL+EF DG R D V SMLY +
Sbjct: 265 KKSEKKSWGTLAFDYGRPEVQSFLISNAIYWLKEFHIDGLRVDAVASMLYLNFDRYDEEE 324
Query: 435 GDYNEYFSEATDVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPISEVGIGFDYR 494
YN Y E +++A +L N ++ + +P A ++AED S +P + P+++ G+GF+Y+
Sbjct: 325 KIYNTYGGEE-NLEAFAFLRKLNKVVFSYIPGALMMAEDSSDLPLVTAPVNKGGLGFNYK 383
Query: 495 LAMAIPDKWIDYLKNKKDH-EWSMKEISLSLTNRRYSEKCVSYAESHDQSIVGDKTFSFL 553
M + + +++ + H +W ++ S +Y+E+ +
Sbjct: 384 WNMGWMNDLLSFMEKESIHRKWHHNRLTFSFM--------YTYSEN----------YLLP 425
Query: 554 LMDEEIYSGMSCLADASPTIE-RGIALHKMIHFITMSLGGEGYLNFMGNEFG-HPEWIDF 611
L +E+ G L D P + + A ++++ + G+ L FMG E + EW D
Sbjct: 426 LSHDEVVHGKKSLLDKMPGDQWQQFANLRLLYGYMYTHPGKK-LVFMGGELAQYAEWKDT 484
Query: 612 PREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDK------FSFLASTKQIVS 665
W L++ + +K + + K +N L + L+ + +
Sbjct: 485 EE-----------LDWHLLE--YPLHKGIYHYIKNLNELYQQHPELYELDHLSEGFEWID 531
Query: 666 STNEEDKVIVFERG------DLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREFGG 719
N + VI F R +L+ + NF P+ ++ YK+G G+Y+ +SD+ FGG
Sbjct: 532 PHNIDQSVIAFRRKANKPNQELIIICNFTPQVHFD-YKIGVPESGRYKEIFNSDSVRFGG 590
Query: 720 HGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVVY 763
G++ +HF+ PE G+ + KI PP V+
Sbjct: 591 SGQINEG-EHFSFPEKWHGLSQ--------HIKIKVPPLAISVF 625
>GLGB_NITEU (Q81ZU6) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 734
Score = 201 bits (511), Expect = 7e-51
Identities = 184/665 (27%), Positives = 288/665 (42%), Gaps = 79/665 (11%)
Query: 127 QGYLKFGFNREEG----GIVYREWAPAAQEAQIIGDFNEWNGSNHPMEKN-QFGVWSIKI 181
Q Y G +R G + WAP A+ ++GDFN W+G +PM + GVW + I
Sbjct: 124 QAYHMLGAHRVRNHGVTGTRFAVWAPNAERVSVVGDFNRWDGRVYPMMVHGHSGVWELFI 183
Query: 182 PDVAGNPAIPHNSRVKFRFRH---GGVWADRIPAWIKYATVDPTKFAAPYDGVY-W--DP 235
PD +P + K+ R+ G + P Y P + Y W D
Sbjct: 184 PD------LPEGAIYKYEIRNRISGEILLKTDPYATTYELRPNNAALTPIEQKYDWKDDD 237
Query: 236 PLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEP--RINSYKEFADDILPRIRANNYNTVQ 293
++ R + + P IYE HVG P R SY E AD ++P ++ Y+ V+
Sbjct: 238 WIARRKGWDWLHAPL----NIYELHVGSWKRHPDGRFYSYHELADHLIPYLQDMGYSHVE 293
Query: 294 LMAVMEHSYYASFGYHVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNV 353
L+ + EH S+GY T +FAV+SR G+PE +D+ H G+ V++D V +H +
Sbjct: 294 LLPISEHPLDESWGYQATGYFAVTSRYGSPEAFMSFVDRCHQAGIGVILDWVPAHFPQD- 352
Query: 354 TDGLNGFDVGQVSQESYFHTGDR-GYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFD 412
+ L FD Y H R GYH W + +FNY EV FLLS+ +WL F D
Sbjct: 353 SFSLARFD----GTALYEHEDPRLGYHHDWGTYIFNYGRNEVKSFLLSSAHYWLSAFHID 408
Query: 413 GFRFDGVTSMLYHHHGVNIAFSGDY--NEYFSEATDVDAVVYLMLANSLIHNILPDATVI 470
G R D V SMLY ++ G++ N Y +++A+ +L N+++H P A
Sbjct: 409 GLRVDAVASMLYLNYSRK---EGEWLPNRYGGH-ENLEAIEFLRALNTMVHGEFPGALTF 464
Query: 471 AEDVSGMPGLGRPISEVGIGFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRYS 530
AE+ + P + RP G+GF + M + + Y+++ H RRY
Sbjct: 465 AEESTSWPAVSRPAYLGGLGFSMKWNMGWMNDTLSYMQHDPVH-------------RRYH 511
Query: 531 EKCVSYAESHDQSIVGDKTFSFLLMDEEIYSGMSCLADASPTIE-RGIALHKMIHFITMS 589
+++ +Q + F L +E+ G + D P + A +++ M+
Sbjct: 512 HNELTF----NQLYAYTENFVLPLSHDEVVHGKKSMLDKMPGDGWQKFANLRLLFTYQMT 567
Query: 590 LGGEGYLNFMGNEFGHPEWIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNL 649
G+ +NFMGNE G E + + E+ + T L + ++N A++
Sbjct: 568 CPGK-KINFMGNELGQGHEWRVGHELDWYLLERDPHRGIQALTRDLNHLYLNT--PALHE 624
Query: 650 LDDKFSFLASTKQIVSSTNEEDKVIVFER----GDLVF-VFNFHPETTYEGYKVGCDLPG 704
LD F A + + E VI ++R G V V NF P GY+VG
Sbjct: 625 LD----FFAEGFSWIDCHDVEQSVISYQRHARDGSFVLVVLNFTP-ILRTGYRVGIPGSS 679
Query: 705 KYRVALDSDAREFGGH--GRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVV 762
Y+ +SD+ + G G G +P G P ++ +P+S I PP V+
Sbjct: 680 AYQEVFNSDSIYYDGSNAGNAGK-----ISPTGQP------WSGQPDSIIITLPPLAGVI 728
Query: 763 YYRVD 767
D
Sbjct: 729 LKAAD 733
>GLGB_BACST (P30538) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 639
Score = 201 bits (510), Expect = 9e-51
Identities = 176/631 (27%), Positives = 288/631 (44%), Gaps = 83/631 (13%)
Query: 146 WAPAAQEAQIIGDFNEWNGSNHPMEK--NQFGVWSIKIPDVAGNPAIPHNSRVKFRFRHG 203
WAP A+E +++G FNEWNG+N + K NQ GVW I IP+ H + + G
Sbjct: 44 WAPHAREVRLVGSFNEWNGTNFNLMKVSNQ-GVWMIFIPENLEG----HLYKYEITTNDG 98
Query: 204 GVWADRIPAWIKYATVDPTKFAAPYD--GVYWDPPLSERYQFKYPRPPKPKAPRIYEAHV 261
V P + Y+ + P + Y+ G W+ R + + +P IYE H
Sbjct: 99 NVLLKSDP-YAFYSELRPHTASIVYNIKGYQWNDQTWRRKKQRKRIYDQPLF--IYELHF 155
Query: 262 GM--SSSEPRINSYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGYHVTNFFAVSSR 319
G + +Y+E A++++P + + + ++L+ ++EH + S+GY +++ +SR
Sbjct: 156 GSWKKKEDGSFYTYQEMAEELIPYVLEHGFTHIELLPLVEHPFDRSWGYQGIGYYSATSR 215
Query: 320 SGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFHTGDRGYH 379
GTP DL Y ID+ H G+ V++D V H + + GL FD G + E Y + DR +
Sbjct: 216 YGTPHDLMYFIDRCHQAGIGVILDWVPGHFCKD-SHGLYMFD-GAPAYE-YANMQDRENY 272
Query: 380 KLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYHHHGVNIAFSGDYNE 439
+W + F+ EV FL+SN +W+E F DGFR D V +MLY + ++ + Y
Sbjct: 273 -VWGTANFDLGKPEVRSFLISNALFWMEYFHVDGFRVDAVANMLYWPNS-DVLYKNTY-- 328
Query: 440 YFSEATDVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPISEVGIGFDYRLAMAI 499
AV +L N + P+ +IAED + P + P + G+GF+Y+ M
Sbjct: 329 ---------AVEFLQKLNETVFAYDPNILMIAEDSTDWPRVTAPTYDGGLGFNYKWNMGW 379
Query: 500 PDKWIDYLKNKKDH-EWSMKEISLSLTNRRYSEKCVSYAESHDQSIVGDKTFSFLLMDEE 558
+ + Y++ +H ++ +++ SL YSE + SHD+ + G K
Sbjct: 380 MNDILTYMETPPEHRKYVHNKVTFSLL-YAYSENFI-LPFSHDEVVHGKK---------- 427
Query: 559 IYSGMSCLADASPTIERGIALHKMIHFITMSLGGEGYLNFMGNEFGH-PEWIDFPREGNG 617
S L+ T E A ++++ ++ G+ L FMG EFG EW D
Sbjct: 428 -----SLLSKMPGTYEEKFAQLRLLYGYLLTHPGKKLL-FMGGEFGQFDEWKDLE----- 476
Query: 618 WSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSFLASTK------QIVSSTNEED 671
+ W L D D ++ MN + K + ++ L + + N E
Sbjct: 477 ------QLDWMLFDFD--MHRNMNMYVKELLKCYKRYKPLYELDHSPDGFEWIDVHNAEQ 528
Query: 672 KVIVFER-----GDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREFGGHGRVGHN 726
+ F R DL+ V Y GYKVG L +YR ++SDA +FGG G +
Sbjct: 529 SIFSFIRRGKKEDDLLIVVCNFTNKVYHGYKVGVPLFTRYREVINSDAIQFGGFGNIN-- 586
Query: 727 VDHFTAPEGIPGVPESNFNNRPNSFKILSPP 757
P+ I + E F+ +P ++ PP
Sbjct: 587 ------PKPIAAM-EGPFHGKPYHIQMTIPP 610
>GLGB_RHOMR (Q93HU3) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 621
Score = 197 bits (500), Expect = 1e-49
Identities = 181/683 (26%), Positives = 296/683 (42%), Gaps = 111/683 (16%)
Query: 115 IEEYEGGLEEFAQGYLKFGFNREEGGIVYREWAPAAQEAQIIGDFNEWNGSNHPMEKNQF 174
I +E G F Y K G + ++ G + WAP A ++G FN+WN +P+E+
Sbjct: 9 IRRWESGT--FYDSYRKLGAHPDDEGTWFCVWAPHADGVSVLGAFNDWNPEANPLERYGG 66
Query: 175 GVWSIKIPDVAGNPAIPHNSRVKFRFRHGGVWADRIPAWIKYATVDPTKFAAPYDGVY-- 232
G+W+ +P A P ++ K+R RHG AD+ + +A PT +P +G+
Sbjct: 67 GLWAGYVPG-----ARPGHT-YKYRIRHGFYQADKTDPYA-FAMEPPT--GSPIEGLASI 117
Query: 233 -------WDPPLSERYQFKYPRPPKPKAP-RIYEAHVGM--SSSEPRINSYKEFADDILP 282
W + + + P P IYE H+G SY+E A+ +
Sbjct: 118 ITRLDYTWH---DDEWMRRRKGPASLYEPVSIYEVHLGSWRHKRPGESFSYREIAEPLAD 174
Query: 283 RIRANNYNTVQLMAVMEHSYYASFGYHVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLM 342
++ + V+L+ VMEH YY S+GY V ++A + R G+P+DL YLID H G+ V++
Sbjct: 175 YVQEMGFTHVELLPVMEHPYYGSWGYQVVGYYAPTFRYGSPQDLMYLIDYLHQRGIGVIL 234
Query: 343 DVVHSHASNNVTDGLNGFDVGQVSQESYFHTGDRGYHKLWDSRLFNYANWEVLRFLLSNL 402
D V SH + + GL FD + + + YH W + +F+Y V FL+SN
Sbjct: 235 DWVPSHFAAD-PQGLVFFDGTTLFE---YDDPKMRYHPDWGTYVFDYNKPGVRNFLISNA 290
Query: 403 RWWLEEFKFDGFRFDGVTSMLYHHHGVNIAFSGDYNE------YFSEATDVDAVVYLMLA 456
+WLE++ DG R D V SMLY DY+ F +++A+ ++
Sbjct: 291 LFWLEKYHVDGLRVDAVASMLYR----------DYSRKEWTPNIFGGRENLEAIDFIKKF 340
Query: 457 NSLIHNILPDATVIAEDVSGMPGLGRPISEVGIGFDYRLAMAIPDKWIDYLKNKKDH-EW 515
N ++ P+A IAE+ + PG+ P G+GF Y+ M +DY++ + ++
Sbjct: 341 NETVYLHFPEAMTIAEESTAWPGVSAPTYNNGLGFLYKWNMGWMHDTLDYIQRDPIYRKY 400
Query: 516 SMKEISLSLTNRRYSEKCVSYAESHDQSIVGDKTFSFLLMDEEIYSGMSCLADASPTIER 575
E++ SL YA S + L +E+ G L P +
Sbjct: 401 HHDELTFSLW----------YAFSEH--------YVLPLSHDEVVHGKGSLWGKMPGDDW 442
Query: 576 GIALHKMIHFITMSLGGEGYLNFMGNEFG-HPEWIDFPREGNGWSYEKCRRQWSLVDTDH 634
A + + F M L FMG EFG H EW + +W L+D
Sbjct: 443 QKAANLRLLFGHMWGHPGKKLLFMGGEFGQHHEW-----------NHDTQLEWHLLD--- 488
Query: 635 LRYKFMNAFDKAMNLLDDKFSFLASTK-----------QIVSSTNEEDKVIVFERGD--- 680
+ + + L + L T + + ++ + VI + R +
Sbjct: 489 ------QPYHRGIQLWVCDLNHLYRTNPALWHDGPEGFEWIDFSDRDQSVICYLRKNAGR 542
Query: 681 -LVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREFGGHGRVGHNVDHFTAPEGIPGV 739
L+FV NF P E Y+VG + G + L+SDA +GG G + +F E +P
Sbjct: 543 MLLFVLNFTP-VPREHYRVGVPIGGPWHEVLNSDAVAYGGSG-----MGNFGRVEAVP-- 594
Query: 740 PESNFNNRPNSFKILSPPRTCVV 762
+++ RP ++ PP ++
Sbjct: 595 --ESWHGRPFHLELTLPPLAALI 615
>GLGB_SYNP7 (P16954) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 773
Score = 196 bits (498), Expect = 2e-49
Identities = 192/709 (27%), Positives = 307/709 (43%), Gaps = 114/709 (16%)
Query: 108 YIDQKKLIEEYEGGLEEFAQG-----YLKFGFNREE----GGIVYREWAPAAQEAQIIGD 158
Y + L+ +Y+ + FA+G Y K G + E G+ + WAP+A+ I+GD
Sbjct: 107 YAFRSPLLTDYD--IHLFAEGNHHRIYEKLGAHPCELENVAGVNFAVWAPSARNVSILGD 164
Query: 159 FNEWNGSNHPMEKNQFGVWSIKIPDVAGNPAIPHNSRVKFRFRHGGVWADRIPAWIKYAT 218
FN W+G H M + G+W + IP++ A + + + G ++ P +
Sbjct: 165 FNSWDGRKHQMARRSNGIWELFIPELTVGAAYKY----EIKNYDGHIYEKSDPYGFQQEV 220
Query: 219 VDPT-KFAAPYDGVYW-DPPLSERYQFKYP-RPPKPKAPRIYEAHVGM---SSSEP---- 268
T A D W D ER + + P R P +YE H+G +SS+
Sbjct: 221 RPKTASIVADLDRYTWGDADWLERRRHQEPLRQPIS----VYEVHLGSWMHASSDAIATD 276
Query: 269 ------------------RINSYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGYHV 310
R +Y+E AD ++P + Y+ ++L+ + EH + S+GY V
Sbjct: 277 AQGKPLPPVPVADLKPGARFLTYRELADRLIPYVLDLGYSHIELLPIAEHPFDGSWGYQV 336
Query: 311 TNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESY 370
T ++A +SR G+PED Y +D+ H G+ V++D V H + GL FD Y
Sbjct: 337 TGYYAATSRYGSPEDFMYFVDRCHQNGIGVILDWVPGHFPKD-GHGLAFFD----GTHLY 391
Query: 371 FHTGDR-GYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYHHHGV 429
H R G H+ W + +FNY EV FL +N +W +++ DG R D V SMLY +
Sbjct: 392 EHADSRQGEHREWGTLVFNYGRHEVRNFLAANALFWFDKYHIDGIRVDAVASMLYLDYNR 451
Query: 430 NIAFSGDY--NEYFSEATDVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPISEV 487
G++ NEY +++A +L N LI + P A IAE+ + P + P
Sbjct: 452 K---EGEWIPNEY-GGRENIEAADFLRQVNHLIFSYFPGALSIAEESTSWPMVSWPTYVG 507
Query: 488 GIGFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRYSEKCVSYAESHDQSIVGD 547
G+GF+ + M +DY S+ R++ + V+++ + S
Sbjct: 508 GLGFNLKWNMGWMHDMLDY-------------FSMDPWFRQFHQNNVTFSIWYAFS---- 550
Query: 548 KTFSFLLMDEEIYSGMSCLADASPTIE--RGIALHKMIHFITMSLGGEGYLNFMGNEFGH 605
+ F L +E+ G S L P E + L ++ ++ G + FMG EFG
Sbjct: 551 ENFMLALSHDEVVHGKSNLIGKMPGDEWQKFANLRCLLGYMFTHPGKKTL--FMGMEFG- 607
Query: 606 PEWIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSFL-------A 658
+W E N W +W L+ + ++ + F K +N L L A
Sbjct: 608 -QW----AEWNVWG----DLEWHLLQYE--PHQGLKQFVKDLNHLYRNAPALYSEDCNQA 656
Query: 659 STKQIVSSTNEEDKVIVFERGD-----LVFVFNFHPETTYEGYKVGCDLPGKYRVALDSD 713
+ I S N V R LV V NF P+ + Y++G + G YR +SD
Sbjct: 657 GFEWIDCSDNRHSIVSFIRRAHESDRFLVVVCNFTPQ-PHAHYRIGVPVAGFYREIFNSD 715
Query: 714 AREFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVV 762
AR +GG +G+ +T E + +NRP S + PP T +V
Sbjct: 716 ARSYGG-SNMGNLGGKWT--------DEWSCHNRPYSLDLCLPPLTTLV 755
>GLGB_SYNY3 (P52981) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 770
Score = 192 bits (489), Expect = 2e-48
Identities = 176/680 (25%), Positives = 296/680 (42%), Gaps = 103/680 (15%)
Query: 140 GIVYREWAPAAQEAQIIGDFNEWNGSNHPMEKNQFGVWSIKIPDVAGNPAIPHNSRVKFR 199
G+ + WAP A+ I+GDFN W+G H M K VW + IP++ + + + + +
Sbjct: 141 GVYFAVWAPNARNVSILGDFNNWDGRLHQMRKRNNMVWELFIPELG----VGTSYKYEIK 196
Query: 200 FRHGGVWADRIPAWIKYATVDP--TKFAAPYDGVYWDPPLSERYQFKYPRPPKPKAPRIY 257
G ++ P Y V P A DG W + + + P K +Y
Sbjct: 197 NWEGHIYEKTDPYGF-YQEVRPKTASIVADLDGYQWHD--EDWLEARRTSDPLSKPVSVY 253
Query: 258 EAHVGM----SSSEP--------------------RINSYKEFADDILPRIRANNYNTVQ 293
E H+G + EP R +Y E D ++P ++ Y ++
Sbjct: 254 ELHLGSWLHTAYDEPVKTLHGEGVPVEVSEWNTGARFLTYYELVDKLIPYVKELGYTHIE 313
Query: 294 LMAVMEHSYYASFGYHVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNV 353
L+ + EH + S+GY VT ++A +SR G+PED Y +D+ H G+ V++D V H +
Sbjct: 314 LLPIAEHPFDGSWGYQVTGYYAPTSRFGSPEDFMYFVDQCHLNGIGVIIDWVPGHFPKD- 372
Query: 354 TDGLNGFDVGQVSQESYFHTGD--RGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKF 411
GL FD Y H GD +G HK W + +FNY EV FL++N +W +++
Sbjct: 373 GHGLAFFD----GTHLYEH-GDPRKGEHKEWGTLIFNYGRNEVRNFLVANALFWFDKYHI 427
Query: 412 DGFRFDGVTSMLY----HHHGVNIAFSGDYNEYFSEATDVDAVVYLMLANSLIHNILPDA 467
DG R D V SMLY G +A NEY +++A +L NS++++ P
Sbjct: 428 DGMRVDAVASMLYLDYCREEGEWVA-----NEY-GGRENLEAADFLRQVNSVVYSYFPGI 481
Query: 468 TVIAEDVSGMPGLGRPISEVGIGFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNR 527
IAE+ + P + P G+GF+ + M +DY S+ R
Sbjct: 482 LSIAEESTSWPMVSWPTYVGGLGFNLKWNMGWMHDMLDY-------------FSMDPWFR 528
Query: 528 RYSEKCVSYAESHDQSIVGDKTFSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFIT 587
++ + ++++ ++ S + + L +E+ G S + P E + F
Sbjct: 529 QFHQNSITFSMWYNHS----ENYMLALSHDEVVHGKSNMLGKMPGDEWQKYANVRALFTY 584
Query: 588 MSLGGEGYLNFMGNEFGHPEWIDFPREGNGWSYEKCRRQWSLVD-TDHLRYKFMNAFDKA 646
M FM EFG +W E N W +W L++ H + K F +
Sbjct: 585 MFTHPGKKTMFMSMEFG--QW----SEWNVWG----DLEWHLLNFPPHQQLK--QFFTEL 632
Query: 647 MNLLDDK-----FSFLASTKQIVSSTNEEDKVIVFERGD------LVFVFNFHPETTYEG 695
+L ++ F S Q + ++ V+ F R +V + NF P+ +
Sbjct: 633 NHLYKNEPALYSNDFDESGFQWIDCSDNRHSVVSFIRRAKNSAEFVVTICNFTPQ-PHSH 691
Query: 696 YKVGCDLPGKYRVALDSDAREFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILS 755
Y+VG +PG Y +SDAR++GG +G+ +T E +F+ +P S +
Sbjct: 692 YRVGVPVPGFYTELFNSDARQYGG-SNMGNLGGKWT--------EEWSFHEQPYSLDLCL 742
Query: 756 PPRTCVVYYRVDESQEENSI 775
PP + V+ ++ ++ EEN++
Sbjct: 743 PPLS-VLVLKLSQNAEENTV 761
>GLGB_VIBPA (Q87FR0) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 755
Score = 191 bits (486), Expect = 5e-48
Identities = 147/604 (24%), Positives = 271/604 (44%), Gaps = 76/604 (12%)
Query: 140 GIVYREWAPAAQEAQIIGDFNEWNGSNHPMEKNQFGVWSIKIPDVAGNPAIPHNSRVKFR 199
G+ + +AP A ++G FN+W+G HPM++ +G+W + IP + V+++
Sbjct: 165 GVRFLVYAPHASAVSLVGCFNQWDGRRHPMQRLDYGIWGLFIPGL--------EEGVQYK 216
Query: 200 FRHGGVWADRIP----AWIKYATVDPTKFAAPYDG--VYWDPPLSERYQFKYPRPPKPKA 253
F G + +P W Y+ P+ + YD W ++Q + + +A
Sbjct: 217 FELKGPNGEGLPHKQDPWGFYSEQYPSFASITYDHKRYQWQ---DAKWQNRAVTQKRDEA 273
Query: 254 PRIYEAHVGMSSSEPRIN--SYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGYHVT 311
YE H G + + + +Y+E A+ ++P + Y V+LM V EH +Y S+GY
Sbjct: 274 LSFYELHAGSWKRDGKGDFLNYRELAEQLVPYLVDMGYTHVELMPVSEHPFYGSWGYQPV 333
Query: 312 NFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYF 371
FA +SR G+P+D K+ +D H G+ V++D V +H ++ GL FD F
Sbjct: 334 GLFAPTSRYGSPDDFKFFVDACHQAGIGVVLDWVPAHFPSD-DHGLANFD-----GTPLF 387
Query: 372 HTGD--RGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYHHHGV 429
H D RG+H+ W+S +++ V RFL+SN +W E+F DG R D V SMLY
Sbjct: 388 HDPDPRRGWHQDWNSYIYDLGREHVRRFLVSNALYWFEQFHIDGIRVDAVASMLY----- 442
Query: 430 NIAFSGDYNEYFSEA----TDVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPIS 485
+ +S ++++ + DA+ L N ++ P+A IAE+ + PG+ P
Sbjct: 443 -LDYSRSHDQWVPNVDGGNENYDAIATLKWMNEEVYKHFPNAMTIAEESTAFPGVSAPTF 501
Query: 486 EVGIGFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRYSEKCVSYAESHDQSIV 545
G+GF ++ M + Y+K H R+Y +++ + S
Sbjct: 502 MGGLGFGFKWNMGWMHDSLSYVKEDPVH-------------RKYHHNTITFPLVYAHS-- 546
Query: 546 GDKTFSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMSLGGEGYLNFMGNEFGH 605
+ + L +E+ G + + P E + ++ M LNFMG E G
Sbjct: 547 --ENYVLSLSHDEVVYGKGSIHNKMPGDEWQQTANLRAYYGYMYGQPGKKLNFMGAEIG- 603
Query: 606 PEWIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSFLASTKQIVS 665
+ W+++ + QW L++ + R++ + + +N L + + + +
Sbjct: 604 --------QTAEWNHDD-QLQWFLLEYE--RHQGVQKLMRDLNHLYRNEAAMHDQDCVPA 652
Query: 666 S-----TNEEDKVI-----VFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAR 715
+E D I + + G+ + + +E +++G G+Y + L++D
Sbjct: 653 GFEWRLQDEADASILAHERISKEGERILIITNFTPVPHERFRLGVPNVGQYELLLNTDDS 712
Query: 716 EFGG 719
++GG
Sbjct: 713 KYGG 716
>GLGB_DEIRA (Q9RTB7) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 705
Score = 191 bits (486), Expect = 5e-48
Identities = 173/640 (27%), Positives = 284/640 (44%), Gaps = 77/640 (12%)
Query: 140 GIVYREWAPAAQEAQIIGDFNEWNGSNHPMEKNQFGVWSIKIPDVAGNPAIPHNSRVKFR 199
G+ + WAP AQ ++GDFN+WNG +HP+++ FG W +P A R KFR
Sbjct: 39 GVRFAVWAPNAQHVSVVGDFNDWNGFDHPLQRLDFGFWGAFVP------AAQPGQRYKFR 92
Query: 200 FRH-GGVWADRIPAWIKYATVDPTKFAAPY--DGVYWDPPLSERYQFKYPRPPKPKAPRI 256
GG D+ + + V P + + D + D E+ + P + I
Sbjct: 93 VTGAGGQTVDKTDPYGTFFEVRPNNASIIWQQDFEWTDAGWLEQRARRGQALEDPIS--I 150
Query: 257 YEAHVGMSS--SEPRINSYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGYHVTNFF 314
YE HVG + + +Y++ A + + Y V+L+ VMEH + S+GY VT ++
Sbjct: 151 YECHVGSWARRDDGWFLNYRDLAHRLGEYVTYMGYTHVELLGVMEHPFDGSWGYQVTGYY 210
Query: 315 AVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFHTG 374
A +SR G+PED KYL++ HSLG+ VL+D V H + L FD + + +
Sbjct: 211 APTSRLGSPEDFKYLVNHLHSLGIGVLLDWVPGHFPTD-EFALAHFDGSPLYE---YADP 266
Query: 375 DRGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYHHHGVNIAFS 434
+GYH W++ +F+Y EV+ FL+ + W++++ DG R D V SMLY ++ + +
Sbjct: 267 RKGYHYDWNTYIFDYGRNEVVMFLIGSALKWVQDYHVDGLRVDAVASMLY----LDFSRT 322
Query: 435 GDYNEYFSEATDVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPISEVGIGFDYR 494
+ +++A+ +L N + H++ P +IAE+ + PG+ P E G+GFDY+
Sbjct: 323 EWVPNIYGGRENLEAIAFLKRLNEVAHHMAPGCLMIAEESTSFPGVTTPTPE-GLGFDYK 381
Query: 495 LAMAIPDKWIDYLK-----NKKDHEWSMKEISLSLTNRRYSEKCVSYAESHDQSIVGDKT 549
AM + + Y + K DH L+ N + + A SHD+ +V K
Sbjct: 382 WAMGWMNDSLAYFEQDPIYRKYDHH------KLTFFNVYRTSENYVLAISHDE-VVHLKK 434
Query: 550 FSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMSLGGEG-YLNFMGNEFGHPEW 608
L + Y+ AD F+ M G L FMG +F
Sbjct: 435 PMVLKHPGDWYAQR---ADYRA-------------FLAMMWTTPGKKLLFMGQDFA---- 474
Query: 609 IDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSFLASTKQ-----I 663
+G+ W+++ + DH MN + L ++ + +
Sbjct: 475 -----QGSEWNHDAPLPWFHADQPDH--RGVMNLVRRLNELYRERPDWHTGDAREEGMAW 527
Query: 664 VSSTNEEDKVIVFERGDL------VFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREF 717
+S+ + ++ V + R D + + N P E Y +G G+YRV L +D EF
Sbjct: 528 ISADDTDNSVYAYARRDTYSGAWSLVIANLTP-VYREDYVIGVPQGGEYRVLLSTDDGEF 586
Query: 718 GGHGRVGHNVDHFTAPEGIPGVPES-NFNNRPNSFKILSP 756
GG G D E G + + N P+S +L P
Sbjct: 587 GGFGT--QQPDLSARDEAAHGQAHALHLNLPPSSVLVLEP 624
>GLGB_NEIDE (Q9RQI5) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 762
Score = 191 bits (484), Expect = 9e-48
Identities = 167/651 (25%), Positives = 286/651 (43%), Gaps = 90/651 (13%)
Query: 140 GIVYREWAPAAQEAQIIGDFNEWNGSNHPMEKNQF-GVWSIKIPDVAGNP-----AIPHN 193
G+ + WAP A+ +IG+FN W+ H M + G+W I IP V N + N
Sbjct: 149 GVRFAVWAPNARRVSVIGEFNGWDSRRHAMRPHTGNGLWDIFIPGVGLNALYKFSVLDAN 208
Query: 194 SRVKFRFRHGGVWADRIPAWIKYATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKPKA 253
++ + A+ P P K AP F+ R +A
Sbjct: 209 GNIREKADPYAFGAELRPTTASVVRGLPAKAEAP--------------AFRR-RANSVEA 253
Query: 254 P-RIYEAHVGMSSSEPRIN---SYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGYH 309
P IYE H+G P N +Y + AD+++ ++ + ++L+ + E+ + S+GY
Sbjct: 254 PISIYEVHLGSWRRNPENNYWLTYTQLADELVNYVKDMGFTHIELLPLSEYPFDGSWGYQ 313
Query: 310 VTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQES 369
T +A +SR G+P++LK LID AH+ G+ V++D V H + GLN FD
Sbjct: 314 ATGLYAPTSRFGSPDELKALIDAAHAAGISVILDWVAGHFPTD-DHGLNTFD----GTAL 368
Query: 370 YFHTGDR-GYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYHHHG 428
Y H R GYH+ W++ ++N+ EV FL N +W+E F FDG R D V SM+Y ++
Sbjct: 369 YEHADPREGYHQDWNTLIYNFGRNEVKNFLQGNALYWIERFGFDGIRVDAVASMIYRNYS 428
Query: 429 VNIAFSGDY--NEYFSEATDVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPISE 486
G++ N Y + +++A+ +L N+++ + P A AE+ + + R E
Sbjct: 429 RK---DGEWIPNRY-GGSENLEAIAFLRQTNAVLKSETPGAGSFAEESTSFADVTR---E 481
Query: 487 VGIGFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRYSEKCVSYAESHDQSIVG 546
G+ FD++ M + + Y++ H R+Y +++ + S
Sbjct: 482 AGLNFDFKWNMGWMNDTLRYMQEDPVH-------------RKYHHGKMTFGMMYQYS--- 525
Query: 547 DKTFSFLLMDEEIYSGMSCLADASP-TIERGIALHKMIHFITMSLGGEGYLNFMGNEFGH 605
+ F L +E+ G L P + A + + G+ L FMGNEF
Sbjct: 526 -ENFVLPLSHDEVVHGKRSLLGKMPGDCWQQFANLRAYYGFMYGFPGKKLL-FMGNEFA- 582
Query: 606 PEWIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSFLASTKQ--- 662
+G W+Y++ W L+D +K + + + +N + + L Q
Sbjct: 583 --------QGREWNYQE-GLDWHLLDEAGGWHKGVQDYVRDLNHIYTAHAPLYQLDQQPE 633
Query: 663 ---IVSSTNEEDKVIVFERGD-----LVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDA 714
+ + + ++ V VFER D ++ + NF P E Y+ G + PG+Y L+SD
Sbjct: 634 GFEWLVADDSDNSVFVFERRDRAGNRIIVISNFTP-VVREHYRFGVNAPGRYTEILNSDR 692
Query: 715 REFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVVYYR 765
++ G G + + D E +P + + S + PP V Y+
Sbjct: 693 TQYQGSG-IANGAD--ITAENVPS------HGKAQSLSLTLPPLATVYLYQ 734
>GLGB_BACAN (Q81K82) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 645
Score = 189 bits (481), Expect = 2e-47
Identities = 171/627 (27%), Positives = 282/627 (44%), Gaps = 84/627 (13%)
Query: 123 EEFAQGYLKFGFN----REEGGIVYREWAPAAQEAQIIGDFNEWNGSNHPM-EKNQFGVW 177
E++ + Y FG + E G+ + WAP A+ ++GDFNEW+ H M + + G+W
Sbjct: 17 EKYYESYNIFGAHIVTEDEMRGVRFTVWAPHAKAMSVVGDFNEWDYEQHKMLQVTEEGIW 76
Query: 178 SIKIPDVAGNPAIPHNSRVKFRFRHGGVWADRI---PAWIKYATVDPTKFAAPYD--GVY 232
S+ IP + R +++ + D I + YA V P + +D G
Sbjct: 77 SLFIPHI--------EEREIYKYAIETMAGDVIFKADPYAVYAEVRPNTASVVFDIKGYE 128
Query: 233 WDPPLSERYQFKYPRPPKPKAPRIYEAHVGM--SSSEPRINSYKEFADDILPRIRANNYN 290
W+ R + K + +A +YE H G + + SY+E A++++P + + +
Sbjct: 129 WNDKNWSRKKKK--KSVYKEAMTVYELHFGSWKKKEDGTLYSYREMAEELIPYVVEHQFT 186
Query: 291 TVQLMAVMEHSYYASFGYHVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHAS 350
+++M ++EH Y S+GY T ++A +SR GTP DL + +D+ H G+ V++D V H
Sbjct: 187 HIEIMPLVEHPYDRSWGYQGTGYYAATSRFGTPHDLMHFVDECHKYGIGVILDWVPGHFC 246
Query: 351 NNVTDGLNGFDVGQVSQESYFHTGDRGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFK 410
+ GL FD G + E + D + +W + F+ EV FL+SN +W+ F
Sbjct: 247 KDA-HGLYLFD-GTPTYE--YKDKDVQENPVWGTVNFDLGKREVRNFLISNALFWMRYFH 302
Query: 411 FDGFRFDGVTSMLYHHHGVNIAFSGDYNEYFSEATDVDAVVYLMLANSLIHNILPDATVI 470
DGFR D V +MLY +N+ E ++ AV +L N + D +
Sbjct: 303 IDGFRVDAVANMLY------------WNKEGQEQSNEHAVSFLRELNEAVFAEDEDFLMT 350
Query: 471 AEDVSGMPGLGRPISEVGIGFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRYS 530
AED + P + P E G+GF+Y+ M + + Y++ ++ R+Y
Sbjct: 351 AEDSTAWPLVTAPTYEGGLGFNYKWNMGWMNDVLKYMECAPEY-------------RKYI 397
Query: 531 EKCVSYAESHDQSIVGDKTFSFLLMDEEIYSGMSCLADASP-TIERGIALHKMIHFITMS 589
++++ + S + F L +E+ G L + P A ++++ +
Sbjct: 398 HDKMTFSLLYAYS----ENFILPLSHDEVVHGKKSLLNKMPGDYWDKFAQLRLLYGYFFT 453
Query: 590 LGGEGYLNFMGNEFGH-PEWIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMN 648
G+ L FMG EFG EW D W+L D + RY M+ + K +
Sbjct: 454 HPGKKLL-FMGGEFGQFDEWKDLED-----------LDWNLHDFEMHRY--MHDYFKELI 499
Query: 649 LLDDKFSFLASTK------QIVSSTNEEDKVIVFER-GD-----LVFVFNFHPETTYEGY 696
L + L Q + + N E + F R GD LV V NF + TYE Y
Sbjct: 500 ALYKRSKPLWQLDHSREGFQWIDANNNEQSIFSFIRQGDKQEDALVVVCNF-TKATYENY 558
Query: 697 KVGCDLPGKYRVALDSDAREFGGHGRV 723
KVG Y L+SDA ++GG G+V
Sbjct: 559 KVGVPDFEYYNEVLNSDAEQYGGSGQV 585
>GLGB_BACCR (Q816G6) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 645
Score = 187 bits (475), Expect = 1e-46
Identities = 171/623 (27%), Positives = 278/623 (44%), Gaps = 76/623 (12%)
Query: 123 EEFAQGYLKFGFN----REEGGIVYREWAPAAQEAQIIGDFNEWNGSNHPM-EKNQFGVW 177
E++ + Y FG + E G+ + WAP A+ ++GDFNEW+ H M + + G+W
Sbjct: 17 EKYYESYNIFGAHVVTEDEIQGVRFTVWAPHAKAMSVVGDFNEWDYEQHKMLQVTEEGIW 76
Query: 178 SIKIPDVAGNPAIPHNSRVKFRFRHGGVWADRIPAWIKYATVDPTKFAAPYD--GVYWDP 235
S+ IP + + G V P + YA V P + +D G W+
Sbjct: 77 SLFIPHIEEGEIYKY----AIETLAGDVILKADP-YAVYAEVRPNTASVVFDIKGYEWND 131
Query: 236 PLSERYQFKYPRPPKPKAPRIYEAHVGM--SSSEPRINSYKEFADDILPRIRANNYNTVQ 293
R + K + +A +YE H G + + SY+E ++++P + + + ++
Sbjct: 132 KNWNRKKKK--KSIYKEAMTVYELHFGSWKKKEDGTLYSYREMVEELIPYVVEHQFTHIE 189
Query: 294 LMAVMEHSYYASFGYHVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNV 353
+M ++EH Y S+GY T ++A +SR GTP DL + +D+ H G+ V++D V H +
Sbjct: 190 IMPLVEHPYDRSWGYQGTGYYAATSRFGTPHDLMHFVDECHKYGIGVILDWVPGHFCKDA 249
Query: 354 TDGLNGFDVGQVSQESYFHTGDRGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDG 413
GL FD G + E + D + +W + F+ EV FL+SN +W+ F DG
Sbjct: 250 -HGLYLFD-GTPTYE--YKDKDVQENPVWGTVNFDLGKREVRNFLISNALFWMRYFHIDG 305
Query: 414 FRFDGVTSMLYHHHGVNIAFSGDYNEYFSEATDVDAVVYLMLANSLIHNILPDATVIAED 473
FR D V +MLY +N+ E ++ AV +L N + D + AED
Sbjct: 306 FRVDAVANMLY------------WNKEGQEQSNEHAVSFLRELNEAVFAEDEDFLMTAED 353
Query: 474 VSGMPGLGRPISEVGIGFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRYSEKC 533
+ P + P E G+GF+Y+ M + + Y++ ++ + E YSE
Sbjct: 354 STAWPLVTTPTYEGGLGFNYKWNMGWMNDVLKYMECAPEYRKHIHEKMTFSLLYAYSENF 413
Query: 534 VSYAESHDQSIVGDKTFSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMSLGGE 593
+ SHD+ + G K+ M + + + L ++++ + G+
Sbjct: 414 I-LPLSHDEVVHGKKSL-LNKMPGDYWDKFAQL--------------RLLYGYFFTHPGK 457
Query: 594 GYLNFMGNEFGH-PEWIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDD 652
L FMG EFG EW D W+L D + RY M+ + K + L
Sbjct: 458 KLL-FMGGEFGQFDEWKDLED-----------LDWNLHDFEMHRY--MHDYFKELIALYK 503
Query: 653 KFSFLASTK------QIVSSTNEEDKVIVFER-GD-----LVFVFNFHPETTYEGYKVGC 700
+ L Q + + N E + F R GD LV V NF + TYE YKVG
Sbjct: 504 RSKPLWQLDHSPEGFQWIDANNNEQSIFSFIRQGDKQEDALVIVCNF-TKATYENYKVGV 562
Query: 701 DLPGKYRVALDSDAREFGGHGRV 723
Y L+SDA+++GG G+V
Sbjct: 563 PDFEYYNEILNSDAQQYGGSGQV 585
>GLGB_AQUAE (O66936) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 630
Score = 185 bits (469), Expect = 5e-46
Identities = 162/598 (27%), Positives = 266/598 (44%), Gaps = 71/598 (11%)
Query: 146 WAPAAQEAQIIGDFNEWNGSNHPMEKNQ--FGVWSIKIP-DVAGNPAIPHNSRVKFRFRH 202
WAP A +IGDFNEW+ + PM K + G+W + + D+ G S+ K+ ++
Sbjct: 45 WAPHADYVSLIGDFNEWDKGSTPMVKREDGSGIWEVLLEGDLTG-------SKYKYFIKN 97
Query: 203 GGVWADRIPAWIKYATVDPTKFAAPYDGVY-WDPPLSERYQFKYPRPPKPKAP-RIYEAH 260
G D+ + + P + + Y W+ Y K R +P IYE H
Sbjct: 98 GNYEVDKSDPFAFFCEQPPGNASVVWKLNYRWN---DSEYMKKRKRVNSHDSPISIYEVH 154
Query: 261 VGMSSSEP----RINSYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGYHVTNFFAV 316
VG P R SY+E A+ + ++ + V+ + VMEH +Y S+GY +T +FA
Sbjct: 155 VGSWRRVPEEGNRFLSYRELAEYLPYYVKEMGFTHVEFLPVMEHPFYGSWGYQITGYFAP 214
Query: 317 SSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFHTGDR 376
+SR GTP+D YLIDK H G+ V++D V SH + GL FD + + + +
Sbjct: 215 TSRYGTPQDFMYLIDKLHQEGIGVILDWVPSHFPTD-AHGLAYFDGTHLYE---YEDWRK 270
Query: 377 GYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYHHHGVNIAFSGD 436
+H W+S +F+Y EV FLLS+ +WL+++ DG R D V SMLY ++ +
Sbjct: 271 RWHPDWNSFVFDYGKPEVRSFLLSSAHFWLDKYHADGLRVDAVASMLY----LDYSRKEW 326
Query: 437 YNEYFSEATDVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPISEVGIGFDYRLA 496
+ +++A+ +L N ++ PD IAE+ + P + RP G+GF +
Sbjct: 327 VPNIYGGKENLEAIEFLRKFNESVYRNFPDVQTIAEESTAWPMVSRPTYVGGLGFGMKWN 386
Query: 497 MAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRYSEKCVSYAESHDQSIVGDKTFSFLLMD 556
M + + Y + R+Y + ++++ + S + F L
Sbjct: 387 MGWMNDTLFYFSKDPIY-------------RKYHHEVLTFSIWYAFS----ENFVLPLSH 429
Query: 557 EEIYSGMSCLADASP--TIERGIALHKMIHFITMSLGGEGYLNFMGNEFGHPEWIDFPRE 614
+E+ G L P ++ L + ++ G + L FMG EFG +
Sbjct: 430 DEVVHGKGSLIGKMPGDYWQKFANLRALFGYMWAHPGKK--LLFMGGEFG---------Q 478
Query: 615 GNGWSYEKCRRQWSLVDTDHLR-----YKFMNAFDKAMNLLDDKFSFLASTKQIVSSTNE 669
W +E W L++ R K +N + L + F + V +
Sbjct: 479 FKEWDHE-TSLDWHLLEYPSHRGIQRLVKDLNEVYRREKALHET-DFSPEGFEWVDFHDW 536
Query: 670 EDKVIVFERGD------LVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREFGGHG 721
E VI F R D ++ V NF P Y+ Y+VG G +R +++DA+E+ G G
Sbjct: 537 EKSVISFLRKDKSGKEIILVVCNFTPVPRYD-YRVGVPKGGYWREIMNTDAKEYWGSG 593
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.136 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 108,859,380
Number of Sequences: 164201
Number of extensions: 5069116
Number of successful extensions: 12167
Number of sequences better than 10.0: 180
Number of HSP's better than 10.0 without gapping: 137
Number of HSP's successfully gapped in prelim test: 43
Number of HSP's that attempted gapping in prelim test: 11677
Number of HSP's gapped (non-prelim): 258
length of query: 848
length of database: 59,974,054
effective HSP length: 119
effective length of query: 729
effective length of database: 40,434,135
effective search space: 29476484415
effective search space used: 29476484415
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0193.7


