

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0193.4
(1140 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
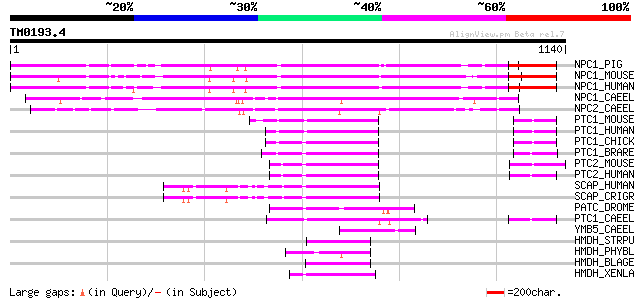
Score E
Sequences producing significant alignments: (bits) Value
NPC1_PIG (P56941) Niemann-Pick C1 protein precursor 562 e-159
NPC1_MOUSE (O35604) Niemann-Pick C1 protein precursor 555 e-157
NPC1_HUMAN (O15118) Niemann-Pick C1 protein precursor 555 e-157
NPC1_CAEEL (Q19127) Niemann-Pick C1 protein homolog 1 precursor 314 8e-85
NPC2_CAEEL (P34389) Niemann-Pick C1 protein homolog 2 precursor 283 1e-75
PTC1_MOUSE (Q61115) Patched protein homolog 1 (PTC1) (PTC) 127 2e-28
PTC1_HUMAN (Q13635) Patched protein homolog 1 (PTC1) (PTC) 124 1e-27
PTC1_CHICK (Q90693) Patched protein homolog 1 (PTC1) (PTC) 124 1e-27
PTC1_BRARE (Q98864) Patched protein homolog 1 (Patched 1) (PTC1) 121 1e-26
PTC2_MOUSE (O35595) Patched protein homolog 2 (PTC2) 118 1e-25
PTC2_HUMAN (Q9Y6C5) Patched protein homolog 2 (PTC2) 113 3e-24
SCAP_HUMAN (Q12770) Sterol regulatory element binding protein cl... 99 7e-20
SCAP_CRIGR (P97260) Sterol regulatory element binding protein cl... 96 4e-19
PATC_DROME (P18502) Patched protein (Hedgehog receptor) 94 3e-18
PTC1_CAEEL (Q09614) Patched protein homolog 1 87 3e-16
YMB5_CAEEL (Q03602) Hypothetical protein F54G8.5 in chromosome III 67 3e-10
HMDH_STRPU (P16393) 3-hydroxy-3-methylglutaryl-coenzyme A reduct... 55 1e-06
HMDH_PHYBL (Q12649) 3-hydroxy-3-methylglutaryl-coenzyme A reduct... 52 7e-06
HMDH_BLAGE (P54960) 3-hydroxy-3-methylglutaryl-coenzyme A reduct... 52 9e-06
HMDH_XENLA (P20715) 3-hydroxy-3-methylglutaryl-coenzyme A reduct... 52 1e-05
>NPC1_PIG (P56941) Niemann-Pick C1 protein precursor
Length = 1277
Score = 562 bits (1448), Expect = e-159
Identities = 370/1094 (33%), Positives = 570/1094 (51%), Gaps = 103/1094 (9%)
Query: 2 YDICGTRSDGKVLNCPF-GSPAVKPDDLLSSKIQSMCPTIT-GNV--CCTKAQFDTLQTQ 57
Y CG S K NC + G P P+D +Q +CP GNV CC Q TL+
Sbjct: 28 YGECGIASGDKRYNCRYSGPPKPLPEDGYDL-VQELCPGFFFGNVSLCCDVQQLRTLKDN 86
Query: 58 VQQAIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTS----VDKAGGNS--TVGGIDY 111
+Q + FL CP+C N +NLFCELTCSP QS F+NVT+ VD + V ++Y
Sbjct: 87 LQLPLQFLSRCPSCFYNLMNLFCELTCSPRQSQFLNVTATEDYVDPVTNQTKTNVKELEY 146
Query: 112 FVSDAFGEGLYESCKDVKFGSMNSRAIQFI---GAGAQNFKEWFAFIGRKAAPNSPGSPY 168
+V + F +Y +C+DV+ S N +A+ + A A N W ++ K ++ +P+
Sbjct: 147 YVGETFANAMYNACRDVEAPSSNEKALGLLCGREAQACNATNWIEYMFNK---DNGQAPF 203
Query: 169 AIMFRPNATKSSGMKPMNVSAYSCSDT----SLGCSCGDCPSSSVCSNSASTTINKANSC 224
I + + GM+PMN + C ++ + CSC DC S VC
Sbjct: 204 TITPIFSDLPTHGMEPMNNATKGCDESVDEVTGPCSCQDC--SIVCGPKPQPPPPPVPWR 261
Query: 225 SIKVGSLTVKCVDFILAVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYAR 284
+ + ++ V +A L + F W R+R P+ I+ V
Sbjct: 262 ILGLDAMYVIMWSSYMAFLIVFFGAFFAVWCY----RKRYFVSEYTPIDGNIAFSV---N 314
Query: 285 NQEKDENLPMQMIEDVPQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVLL 344
+ +K + + + + + + ++G+ RHP V+ +LA ++
Sbjct: 315 SSDKGQAFCCDPLG-----------AAFERGLRRLFAQWGAFCVRHPGCVVFFSLAFIVA 363
Query: 345 LCLGLIRFKVETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSPR 404
GL+ +V T P LW PGS+A +EK++FD+H PF+R+EQLI+ + + P
Sbjct: 364 CSSGLVFIRVTTDPVDLWSAPGSQARREKEYFDTHFGPFFRMEQLIIRATNNQSHIYHPY 423
Query: 405 IVSADN----------IRFLFEVQKKVDAIRANYSGLMVSLQDICMKPL---DKDCATQS 451
AD + + ++Q ++ I A+Y+ V+LQDIC+ PL +K+C S
Sbjct: 424 PAGADVPFGPPLSRDILHQVLDLQTAIENITASYNNETVTLQDICLAPLSPYNKNCTILS 483
Query: 452 VLQYFKMDPRNFDD---------SGAVEHLNYCFQQYSSA-------DQCMSAFKAPLDP 495
VL YF+ D + H YC + +S D C+ F P+ P
Sbjct: 484 VLNYFQNSHSVLDHQVGDFFFVYADYHTHFLYCVRAPASLNDASLLHDPCLGTFGGPVFP 543
Query: 496 STVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMAQSR 555
VLGG+ ++Y+ A+A ++T+PVNN ++ + +A AWE FI VK+ P
Sbjct: 544 WLVLGGYDDQNYNNATALVITFPVNNYYNDT-EKLQRAQAWESEFINFVKNYKNP----- 597
Query: 556 NLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKVLLG 615
NLT++F +E SIE+EL RES +D TIL+SY +MF YIS+ LG S + SK+ LG
Sbjct: 598 NLTISFMAERSIEDELNRESNSDLFTILISYAIMFLYISIALGHIKSCSRLLVDSKISLG 657
Query: 616 LSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQP--LE 673
++G+++V+ SV S+ IFS +GV TLI++EVIPFLVLAVGVDN+ ILV +R
Sbjct: 658 IAGILIVLSSVACSLGIFSYIGVPLTLIVIEVIPFLVLAVGVDNIFILVQTYQRDERLQG 717
Query: 674 LPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVT 733
L+ ++ L EV PS+ L+S SE +AF +G +PA FS+FA +AVL+DF+LQ+T
Sbjct: 718 ETLDQQLGRVLGEVAPSMFLSSFSETVAFFLGGLSVVPAVHTFSLFAGMAVLIDFLLQIT 777
Query: 734 AFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAPILSIW 793
FV+L+ LD +R E R+D C++ A+ G+ Q L R+ K +AP+L
Sbjct: 778 CFVSLLGLDIKRQEKNRLDVVCCVQ----GAEDGAGV-QASESCLFRFFKNSYAPLLLKD 832
Query: 794 GVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYFV 853
++ +VIA+FV SIA+ ++E GL+Q + +P DSY+ YF ++S YL GPP+YFV
Sbjct: 833 WMRPIVIAVFVGVLSFSIAVLNKVEIGLDQSLSMPDDSYVMDYFQSLSRYLHAGPPVYFV 892
Query: 854 V-KNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWIS 912
V + +NY+S N +C CN+DSL+ +I A+ + + I +SW+DD+ WI
Sbjct: 893 VEEGHNYTSLKGQ-NMVCGGLGCNNDSLVQQIFTAAQLDNYTRIGFAPSSWIDDYFDWIK 951
Query: 913 PEAFGCCRKF-TNGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQF 971
P++ CCR + + +C + S V C C R M+F
Sbjct: 952 PQS-SCCRVYNSTDQFC------------NASVVDPTCIRCRPLTSEGKQRPQGEDFMRF 998
Query: 972 RDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDYVNSM 1031
LP FLS P+ C KGGH AY+S+V++ G SG + A+ F TYHT L D++++M
Sbjct: 999 ---LPMFLSDNPNPKCGKGGHAAYSSAVNILGNGSG-VGATYFMTYHTVLQASADFIDAM 1054
Query: 1032 RAAREFSSKVSDSL 1045
+ AR +S ++ ++
Sbjct: 1055 QKARLIASNITRTM 1068
Score = 98.2 bits (243), Expect = 1e-19
Identities = 49/99 (49%), Positives = 73/99 (73%), Gaps = 1/99 (1%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVAS-GDKDQRVKEALGTMGASVFSGITLTKLVGVIVLY 1083
V+ V S + EF S ++ + +++ G + R +EAL MG+SVFSGITLTK G++VL
Sbjct: 1154 VNLVMSCGISVEFCSHITRAFTLSTKGSRVDRAEEALAHMGSSVFSGITLTKFGGIVVLA 1213
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGP 1122
F+++++F I+YF+MYL++VLLG HGL+FLPV+LS GP
Sbjct: 1214 FAKSQIFQIFYFRMYLAIVLLGATHGLIFLPVLLSYIGP 1252
>NPC1_MOUSE (O35604) Niemann-Pick C1 protein precursor
Length = 1278
Score = 555 bits (1431), Expect = e-157
Identities = 371/1103 (33%), Positives = 575/1103 (51%), Gaps = 109/1103 (9%)
Query: 2 YDICGTRSDGKVLNCPF-GSPAVKPDDLLSSKIQSMCPTI---TGNVCCTKAQFDTLQTQ 57
Y CG + K NC + G P P D +Q +CP + ++CC Q TL++
Sbjct: 29 YGECGIATGDKRYNCKYSGPPKPLPKDGYDL-VQELCPGLFFDNVSLCCDIQQLQTLKSN 87
Query: 58 VQQAIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVD-----KAGGNST-VGGIDY 111
+Q + FL CP+C N + LFCELTCSP+QS F+NVT+ + K N T V ++Y
Sbjct: 88 LQLPLQFLSRCPSCFYNLMTLFCELTCSPHQSQFLNVTATEDYFDPKTPENKTNVKELEY 147
Query: 112 FVSDAFGEGLYESCKDVKFGSMNSRAIQFI---GAGAQNFKEWFAFIGRKAAPNSPGSPY 168
+V +F +Y +C+DV+ S N +A+ + A A N W ++ K ++ +P+
Sbjct: 148 YVGQSFANAMYNACRDVEAPSSNEKALGLLCGRDARACNATNWIEYMFNK---DNGQAPF 204
Query: 169 AIMFRPNATKSSGMKPMNVSAYSCSDT----SLGCSCGDCPSSSVCSNSASTTINKANSC 224
I+ + GM+PM + C+++ + CSC DC S VC
Sbjct: 205 TIIPVFSDLSILGMEPMRNATKGCNESVDEVTGPCSCQDC--SIVCGPKPQPP---PPPM 259
Query: 225 SIKVGSLTVKCVDFILAVLYIILICVFLGWALY---HRIRERKMTYRTEPVSNVISGGVL 281
++ L V I+ V Y+ + VF G L HR R Y P+ + I+ V
Sbjct: 260 PWRIWGLDAMYV--IMWVTYVAFLFVFFGALLAVWCHRRRYFVSEYT--PIDSNIAFSVN 315
Query: 282 YARNQEKDENLPMQMIEDVPQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAI 341
+ E P+ D + + K+G+ R+P ++ +LA
Sbjct: 316 SSDKGEASCCDPLGAAFD--------------DCLRRMFTKWGAFCVRNPTCIIFFSLAF 361
Query: 342 VLLLCLGLIRFKVETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNST 401
+ + GL+ +V T P +LW P S+A EK++FD H PF+R EQLI+ ++
Sbjct: 362 ITVCSSGLVFVQVTTNPVELWSAPHSQARLEKEYFDKHFGPFFRTEQLIIQAPNTSVHIY 421
Query: 402 SPRIVSAD-------NIRFLFEV---QKKVDAIRANYSGLMVSLQDICMKPL---DKDCA 448
P AD N L +V Q +++I A+Y+ V+LQDIC+ PL +K+C
Sbjct: 422 EPYPAGADVPFGPPLNKEILHQVLNLQIAIESITASYNNETVTLQDICVAPLSPYNKNCT 481
Query: 449 TQSVLQYFK-----MDPRNFDD----SGAVEHLNYCFQQYSSADQ-------CMSAFKAP 492
SVL YF+ +D + DD + H YC + +S + C+ F P
Sbjct: 482 IMSVLNYFQNSHAVLDSQVGDDFYIYADYHTHFLYCVRAPASLNDTSLLHGPCLGTFGGP 541
Query: 493 LDPSTVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMA 552
+ P VLGG+ ++Y+ A+A ++T+PVNN ++ +A AWEK FI VK+ P
Sbjct: 542 VFPWLVLGGYDDQNYNNATALVITFPVNNYYNDT-ERLQRAWAWEKEFISFVKNYKNP-- 598
Query: 553 QSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKV 612
NLT++F++E SIE+EL RES +D T+++SY+VMF YISL LG S + SK+
Sbjct: 599 ---NLTISFTAERSIEDELNRESNSDVFTVIISYVVMFLYISLALGHIQSCSRLLVDSKI 655
Query: 613 LLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQP- 671
LG++G+++V+ SV S+ IFS +G+ TLI++EVIPFLVLAVGVDN+ ILV +R
Sbjct: 656 SLGIAGILIVLSSVACSLGIFSYMGMPLTLIVIEVIPFLVLAVGVDNIFILVQTYQRDER 715
Query: 672 -LELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFVL 730
E L+ ++ L EV P++ L+S SE AF G+ SMPA FS+FA +AVL+DF+L
Sbjct: 716 LQEETLDQQLGRILGEVAPTMFLSSFSETSAFFFGALSSMPAVHTFSLFAGMAVLIDFLL 775
Query: 731 QVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAPIL 790
Q+T FV+L+ LD +R E +D C++ AD +G L R+ K AP+L
Sbjct: 776 QITCFVSLLGLDIKRQEKNHLDILCCVR----GADDGQG-SHASESYLFRFFKNYFAPLL 830
Query: 791 SIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPL 850
++ +V+A+FV S+A+ +++ GL+Q + +P DSY+ F ++++YL GPP+
Sbjct: 831 LKDWLRPIVVAVFVGVLSFSVAVVNKVDIGLDQSLSMPNDSYVIANFKSLAQYLHSGPPV 890
Query: 851 YFVV-KNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLV 909
YFV+ + YNYSS N +C C++DSL+ +I A+ + + + +SW+DD+
Sbjct: 891 YFVLEEGYNYSSRKGQ-NMVCGGMGCDNDSLVQQIFNAAELDTYTRVGFAPSSWIDDYFD 949
Query: 910 WISPEAFGCCRKF-TNGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTST 968
W+SP++ CCR + +C P C C+ T R
Sbjct: 950 WVSPQS-SCCRLYNVTHQFCNASVMDPTCV---------RCRPLT------PEGKQRPQG 993
Query: 969 MQFRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDYV 1028
+F LP FLS P+ C KGGH AY S+V++ G D I A+ F TYHT L DY
Sbjct: 994 KEFMKFLPMFLSDNPNPKCGKGGHAAYGSAVNIVG-DDTYIGATYFMTYHTILKTSADYT 1052
Query: 1029 NSMRAAREFSSKVSDSLKVASGD 1051
++M+ AR +S ++++++ D
Sbjct: 1053 DAMKKARLIASNITETMRSKGSD 1075
Score = 98.2 bits (243), Expect = 1e-19
Identities = 49/99 (49%), Positives = 73/99 (73%), Gaps = 1/99 (1%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVAS-GDKDQRVKEALGTMGASVFSGITLTKLVGVIVLY 1083
V+ V S + EF S ++ + +++ G + R +EAL MG+SVFSGITLTK G++VL
Sbjct: 1155 VNLVMSCGISVEFCSHITRAFTMSTKGSRVSRAEEALAHMGSSVFSGITLTKFGGIVVLA 1214
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGP 1122
F+++++F I+YF+MYL++VLLG HGL+FLPV+LS GP
Sbjct: 1215 FAKSQIFEIFYFRMYLAMVLLGATHGLIFLPVLLSYIGP 1253
>NPC1_HUMAN (O15118) Niemann-Pick C1 protein precursor
Length = 1278
Score = 555 bits (1430), Expect = e-157
Identities = 370/1100 (33%), Positives = 566/1100 (50%), Gaps = 110/1100 (10%)
Query: 2 YDICGTRSDGKVLNCPF-GSPAVKPDDLLSSKIQSMCPTIT-GNV--CCTKAQFDTLQTQ 57
Y CG K NC + G P P D +Q +CP GNV CC Q TL+
Sbjct: 28 YGECGIAYGDKRYNCEYSGPPKPLPKDGYDL-VQELCPGFFFGNVSLCCDVRQLQTLKDN 86
Query: 58 VQQAIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTS----VDKAGGNS--TVGGIDY 111
+Q + FL CP+C N LNLFCELTCSP QS F+NVT+ VD + V + Y
Sbjct: 87 LQLPLQFLSRCPSCFYNLLNLFCELTCSPRQSQFLNVTATEDYVDPVTNQTKTNVKELQY 146
Query: 112 FVSDAFGEGLYESCKDVKFGSMNSRAIQFI---GAGAQNFKEWFAFIGRKAAPNSPGSPY 168
+V +F +Y +C+DV+ S N +A+ + A A N W ++ K ++ +P+
Sbjct: 147 YVGQSFANAMYNACRDVEAPSSNDKALGLLCGKDADACNATNWIEYMFNK---DNGQAPF 203
Query: 169 AIMFRPNATKSSGMKPMNVSAYSCSDT----SLGCSCGDCPSSSVCSNSASTTINKANSC 224
I + GM+PMN + C ++ + CSC DC S VC A
Sbjct: 204 TITPVFSDFPVHGMEPMNNATKGCDESVDEVTAPCSCQDC--SIVCGPKPQPPPPPAPWT 261
Query: 225 SIKVGSLTVKCVDFILAVLYIILICVFLG-----WALYHRIRERKMTYRTEPVSNVISGG 279
+ + ++ V I+ + Y+ + VF G W R+R P+ + I+
Sbjct: 262 ILGLDAMYV-----IMWITYMAFLLVFFGAFFAVWCY----RKRYFVSEYTPIDSNIAFS 312
Query: 280 VLYARNQEKDENLPMQMIEDVPQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTL 339
V + E P+ + +G + + ++GS R+P V+ +L
Sbjct: 313 VNASDKGEASCCDPVS--------------AAFEGCLRRLFTRWGSFCVRNPGCVIFFSL 358
Query: 340 AIVLLLCLGLIRFKVETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMN 399
+ GL+ +V T P LW P S+A EK++FD H PF+R EQLI+ +
Sbjct: 359 VFITACSSGLVFVRVTTNPVDLWSAPSSQARLEKEYFDQHFGPFFRTEQLIIRAPLTDKH 418
Query: 400 STSPRIVSAD-------NIRFLFEV---QKKVDAIRANYSGLMVSLQDICMKPL---DKD 446
P AD +I+ L +V Q ++ I A+Y V+LQDIC+ PL + +
Sbjct: 419 IYQPYPSGADVPFGPPLDIQILHQVLDLQIAIENITASYDNETVTLQDICLAPLSPYNTN 478
Query: 447 CATQSVLQYFK-----MDPRNFDD----SGAVEHLNYCFQQYSSA-------DQCMSAFK 490
C SVL YF+ +D + DD + H YC + +S D C+ F
Sbjct: 479 CTILSVLNYFQNSHSVLDHKKGDDFFVYADYHTHFLYCVRAPASLNDTSLLHDPCLGTFG 538
Query: 491 APLDPSTVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLP 550
P+ P VLGG+ ++Y+ A+A ++T+PVNN ++ + +A AWEK FI VK+ P
Sbjct: 539 GPVFPWLVLGGYDDQNYNNATALVITFPVNNYYNDT-EKLQRAQAWEKEFINFVKNYKNP 597
Query: 551 MAQSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISS 610
NLT++F++E SIE+EL RES +D T+++SY +MF YISL LG + S
Sbjct: 598 -----NLTISFTAERSIEDELNRESDSDVFTVVISYAIMFLYISLALGHIKSCRRLLVDS 652
Query: 611 KVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQ 670
KV LG++G+++V+ SV S+ +FS +G+ TLI++EVIPFLVLAVGVDN+ ILV A +R
Sbjct: 653 KVSLGIAGILIVLSSVACSLGVFSYIGLPLTLIVIEVIPFLVLAVGVDNIFILVQAYQRD 712
Query: 671 P--LELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDF 728
L+ ++ L EV PS+ L+S SE +AF +G+ MPA FS+FA LAV +DF
Sbjct: 713 ERLQGETLDQQLGRVLGEVAPSMFLSSFSETVAFFLGALSVMPAVHTFSLFAGLAVFIDF 772
Query: 729 VLQVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAP 788
+LQ+T FV+L+ LD +R E R+D F C++ D Q L R+ K ++P
Sbjct: 773 LLQITCFVSLLGLDIKRQEKNRLDIFCCVR-----GAEDGTSVQASESCLFRFFKNSYSP 827
Query: 789 ILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGP 848
+L ++ +VIAIFV SIA+ +++ GL+Q + +P DSY+ YF ++S+YL GP
Sbjct: 828 LLLKDWMRPIVIAIFVGVLSFSIAVLNKVDIGLDQSLSMPDDSYMVDYFKSISQYLHAGP 887
Query: 849 PLYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFL 908
P+YFV++ + + S N +C CN+DSL+ +I A+ + + I +SW+DD+
Sbjct: 888 PVYFVLEEGHDYTSSKGQNMVCGGMGCNNDSLVQQIFNAAQLDNYTRIGFAPSSWIDDYF 947
Query: 909 VWISPEAFGCCR-KFTNGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTS 967
W+ P++ CCR +C + S V AC C R
Sbjct: 948 DWVKPQS-SCCRVDNITDQFC------------NASVVDPACVRCRPLTPEGKQRPQGGD 994
Query: 968 TMQFRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDY 1027
M+F LP FLS P+ C KGGH AY+S+V++ + A+ F TYHT L D+
Sbjct: 995 FMRF---LPMFLSDNPNPKCGKGGHAAYSSAVNILLGHGTRVGATYFMTYHTVLQTSADF 1051
Query: 1028 VNSMRAAREFSSKVSDSLKV 1047
+++++ AR +S V++++ +
Sbjct: 1052 IDALKKARLIASNVTETMGI 1071
Score = 99.8 bits (247), Expect = 4e-20
Identities = 50/99 (50%), Positives = 73/99 (73%), Gaps = 1/99 (1%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVA-SGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLY 1083
V+ V S + EF S ++ + V+ G + +R +EAL MG+SVFSGITLTK G++VL
Sbjct: 1155 VNLVMSCGISVEFCSHITRAFTVSMKGSRVERAEEALAHMGSSVFSGITLTKFGGIVVLA 1214
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGP 1122
F+++++F I+YF+MYL++VLLG HGL+FLPV+LS GP
Sbjct: 1215 FAKSQIFQIFYFRMYLAMVLLGATHGLIFLPVLLSYIGP 1253
>NPC1_CAEEL (Q19127) Niemann-Pick C1 protein homolog 1 precursor
Length = 1383
Score = 314 bits (805), Expect = 8e-85
Identities = 265/1086 (24%), Positives = 471/1086 (42%), Gaps = 113/1086 (10%)
Query: 32 KIQSMCP-TITGN--VCCTKAQFDTLQTQVQQAIPFLVGCPACLRNFLNLFCELTCSPNQ 88
K+ CP +TG+ +CCT +Q + L Q+ QA L CP+C NF L+CE TCSPNQ
Sbjct: 60 KMVEFCPHLLTGDNKLCCTPSQAEGLTKQIAQARHILGRCPSCFDNFAKLWCEFTCSPNQ 119
Query: 89 SLFINVTS---VDKAGG--------NSTVGGIDYFVSDAFGEGLYESCKDVKFGSMNSRA 137
F++++ ++K G + V ++Y +S F EG++ SCKDV FG +
Sbjct: 120 QDFVSISEMKPIEKKEGFTPEYQPAEAYVNTVEYRLSTDFAEGMFSSCKDVTFGGQPALR 179
Query: 138 IQFIGAGAQNFKEWFAFIGRKAAP-NSPGSPYAIMFRPNATKSSGMKP-MNVSAYSCSDT 195
+ W FIG + N P +++ P T S MNV+
Sbjct: 180 VMCTSTPC-TLTNWLEFIGTQNLDLNIPIHTKFLLYDPIKTPPSDRSTYMNVN------- 231
Query: 196 SLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVK-CVDFILAVLYIILICVFLGW 254
GC CS S AN + G + + C +A L I ++ F+G
Sbjct: 232 FTGCDKSARVGWPACSTSECNKEEYANLIDLDDGKTSGQTCNVHGIACLNIFVMLAFIG- 290
Query: 255 ALYHRIRERKMTYRTEPVSNVISGGVLYARNQEKDENLPMQMIEDVPQNRNGVRL--SVV 312
++ ++ G ++ E NL + RN ++ + +
Sbjct: 291 ----------------SLAVLLCVGFVFTSYDEDYTNLRQTQSGEESPKRNRIKRTGAWI 334
Query: 313 QGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVETRPEKLWVGPGSKAAQE 372
+M N R G + R+P + + A+++ G+I K T +W P S+A QE
Sbjct: 335 HNFMENNARDIGMMAGRNPKSHFFIGCAVLIFCLPGMIYHKESTNVVDMWSSPRSRARQE 394
Query: 373 KQFFDSHLAPFYRIEQLILATVPDHMNSTS--PRIVSADNIRFLFEVQKKVDAIRANYS- 429
+ F+++ R +Q++L + D +S + D LF++ + I S
Sbjct: 395 EMVFNANFGRPQRYQQIMLLSHRDFQSSGKLYGPVFHKDIFEELFDILNAIKNISTQDSD 454
Query: 430 GLMVSLQDICMKPLDK--DCATQSVLQYFKMDPRNFD-----------DSGA-------- 468
G ++L D+C +P+ DC S YF+ + + D D A
Sbjct: 455 GRTITLDDVCYRPMGPGYDCLIMSPTNYFQGNKEHLDMKSNKEETVSEDDDAFDYFSSEA 514
Query: 469 -----VEHLNYCF-----QQYSSADQCMSAFKAPLDPSTVLGGFSGKDYSGASAFIVTYP 518
+ H+ C Q+ S CM + P P+ V G + ++ A++ ++T
Sbjct: 515 TTDEWMNHMAACIDQPMSQKTKSGLSCMGTYGGPSAPNMVFGK-NSTNHQAANSIMMTIL 573
Query: 519 VNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKRESTAD 578
V + E KA WEK F++ K+ +S + +F +E SI +E++ ++ +
Sbjct: 574 VTQRTEP---EIQKAELWEKEFLKFCKEY---REKSPKVIFSFMAERSITDEIENDAKDE 627
Query: 579 AITILVSYLVMFAYISLTLGD----TPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFS 634
+T++++ + Y++ +LG S + S++ LG+ VI+ +LS S IFS
Sbjct: 628 IVTVVIALAFLIGYVTFSLGRYFVCENQLWSILVHSRICLGMLSVIINLLSSFCSWGIFS 687
Query: 635 ALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELPLEGR------ISNALVEVG 688
G+ + V F+V +GV ++V +Q + +P + +
Sbjct: 688 MFGIHPVKNALVVQFFVVTLLGVCRTFMVVKYYAQQRVSMPYMSPDQCPEIVGMVMAGTM 747
Query: 689 PSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAED 748
P++ +SL +F +G F +PA R F ++A LAVL+D VL T F+AL V D+QR +
Sbjct: 748 PAMFSSSLGCAFSFFIGGFTDLPAIRTFCLYAGLAVLIDVVLHCTIFLALFVWDTQRELN 807
Query: 749 KRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAPILSIWGVKIVVIAIFVAFAL 808
+ + F ++ G ++ + ++ AP L +I+ IF+A +
Sbjct: 808 GKPEFFFPYQIKDLLGAYLIGRQRATDTFMTQFFHFQVAPFLMHRMTRIITGIIFIASFI 867
Query: 809 ASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYFVVKNYNYSSESTHTNQ 868
++ LS++I G +Q + SY+ +F + ++ +GPP++F V N+
Sbjct: 868 TTVILSSKISVGFDQSMAFTEKSYISTHFRYLDKFFDVGPPVFFTVDGELDWHRPDVQNK 927
Query: 869 LCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYC 928
C+ C+ S N ++ A E +Y++ +W+D++L WIS ++ CC+ + +
Sbjct: 928 FCTFPGCSDTSFGNIMNYAVGHTEQTYLSGEMYNWIDNYLEWISRKS-PCCKVYVH---- 982
Query: 929 PPDDQPPCCAPEDDSCVSG-ACKDC------TTCFRHSDLRNDRTSTMQFRDKLPWFLSA 981
D C + S + AC+ C + S + R S F L FL
Sbjct: 983 --DPNTFCSTNRNKSALDDKACRTCMDFDYVANSYPKSSIMYHRPSIEVFYRHLRHFLED 1040
Query: 982 LPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPL--NKQVDYVNSMRAAREFSS 1039
P+++C GG ++ ++ G IQAS F T+H L + D++ +M AR S
Sbjct: 1041 TPNSECVFGGRASFKDAISFT--SRGRIQASQFMTFHKKLSISNSSDFIKAMDTARMVSR 1098
Query: 1040 KVSDSL 1045
++ S+
Sbjct: 1099 RLERSI 1104
Score = 44.3 bits (103), Expect = 0.002
Identities = 34/140 (24%), Positives = 68/140 (48%), Gaps = 17/140 (12%)
Query: 996 TSSVDLKGYDSGII-QASSF-------RTYHTPLN--KQVDYVNSMRAAREFSSKV---- 1041
T +D+KG +I Q S++ ++ P+N + V S EFS V
Sbjct: 1148 TLGIDVKGAACAVICQVSNYFHIVAFMYIFNIPVNALSATNLVMSSGILIEFSVNVLKGY 1207
Query: 1042 SDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSL 1101
+ SL+ + D R + +G++G + SG +T + L + ++ +Y+F+++L
Sbjct: 1208 ACSLRQRAKD---RAESTVGSIGPIILSGPVVTMAGSTMFLSGAHLQIITVYFFKLFLIT 1264
Query: 1102 VLLGFLHGLVFLPVVLSIFG 1121
++ +H L+ LP++L+ G
Sbjct: 1265 IVSSAVHALIILPILLAFGG 1284
>NPC2_CAEEL (P34389) Niemann-Pick C1 protein homolog 2 precursor
Length = 1274
Score = 283 bits (725), Expect = 1e-75
Identities = 256/1058 (24%), Positives = 448/1058 (42%), Gaps = 139/1058 (13%)
Query: 44 VCCTKAQFDTLQTQVQQAIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDKAGGN 103
VCCT+ Q + ++ A L CP+C NF L+C+ TCSP+QS F+ V ++ G
Sbjct: 85 VCCTELQLKGMTDRISNAATILGSCPSCFDNFAKLWCQFTCSPDQSKFMKV--METTGPK 142
Query: 104 STVGGIDYFVSDAFGEGLYESCKDVKFGSMNSRAIQFIGAGAQ-NFKEWFAFIGRKAAPN 162
+ V +++ V+ F EGLYESC+ F N A++ + G + +F+ ++ F+G K
Sbjct: 143 NVVVKMEFKVNRDFVEGLYESCRHTWFA--NGLALRLMSLGGKVSFENFYGFMGTKNLAQ 200
Query: 163 SPGSPYAIMFRPNATKSSGMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNSASTTINKAN 222
S P F+ + K++ P S C DCP+++ I+K
Sbjct: 201 S--IPINTEFQFSRMKNAMNIPTTPCHKSAGPKVPACGAIDCPTNA----HQLVDISKVE 254
Query: 223 SCSIKVGSLTVKCVDFILAVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLY 282
KV +++L + + + V L + L + R
Sbjct: 255 HLGTKVFHPHFPDFEWLLKICGCLALTVLLVFILKYSCHRRS------------------ 296
Query: 283 ARNQEKDENLPMQMIEDVPQNRNGVRLSVVQGYMSNFYR----KYGSLVARHPINVLALT 338
P +G + + +G + + +Y + V +HP+ ++L
Sbjct: 297 -----------------APNGEDGCYVDLGKGNLEVQFEGLCARYANAVIKHPLIFVSLG 339
Query: 339 LAIVLLLCLGLIRFKVETRP-EKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDH 397
L + C G +F T +++ G EK+F S P +RIEQ+ + P
Sbjct: 340 LIVAAACCSGNFKFHSLTHSVDQVSAADGETRRNEKKFIHS-FGPNHRIEQIFINLPP-- 396
Query: 398 MNSTSPRIVSADNIRFLFEVQKKVDAIRANYSGLMVSLQDICMKPLDKD--CATQSVLQY 455
T+ + + +F++ + + A Y V L DIC KP+ K+ CA S Y
Sbjct: 397 ---TTKSMFNMPLFEEMFQLVGNIQNLTACYGNSSVKLDDICYKPIGKNHGCAIMSPTNY 453
Query: 456 FKMDPRNFDDSGAV------------EHLNYCFQQ------YSSADQCMSAFKAPLDPST 497
F+ NF+++G EHL YC + YS C F P+DP
Sbjct: 454 FQNKWTNFENAGPPTIDDEIFDDQHWEHLKYCIRNPLTVSTYSEMS-CFGEFSGPIDPIL 512
Query: 498 VLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMAQSRNL 557
V GG S + GA + + + G E +A+AWE AF+ ++ + ++
Sbjct: 513 VFGG-SNESIKGAEMYYTARTIMITVLIRGPED-QAIAWETAFLNMMS-----RYEMKHA 565
Query: 558 TLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISS----KVL 613
F +E+S+ EE+ D I +++ + ++ LG P S +S+ K+L
Sbjct: 566 NFTFMTETSVAEEIHTAVETDKIVSVIACAAVLIWVITMLGINHWPESSILSALVHHKLL 625
Query: 614 LGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQP-- 671
+ +S V++ ++SV S+ +FS GV +T + V+ F++ +G++ + +++ +
Sbjct: 626 ISISAVMISVISVWCSIGMFSLFGVHATDNAIVVLFFVITCLGINRIFVIIRTFQANGHC 685
Query: 672 LELP------LEGRISNALVEVGPSITLASL--SEVLAFAVGSF----ISMPACRVFSMF 719
LP + RISN + P + SL S L A G +SMPA VF+
Sbjct: 686 YGLPNISYREMNHRISNVMRRSIPIVLTNSLICSTCLFLAGGVLPYVSVSMPAVEVFARH 745
Query: 720 AALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPCI------KVHSFHADPDKGIRQR 773
A LA+L+D + + L D++R + + +P K++ D +R
Sbjct: 746 AGLAILMDTAFYLLVMLPLFQYDARREMSGKCEIWPWYELSNESKINLCMEAVDGNLRSP 805
Query: 774 KPGLLARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYL 833
+ K AP+L +I + F + + + +E G Q + SYL
Sbjct: 806 -----VDWFKLAIAPLLLKKICRIWIATFFFVSLIIACYCTLCLEFGFNQVMAFSETSYL 860
Query: 834 QGYFNNVSEYLRIGPPLYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVP-- 891
+F N++E L IGPPL+FVV+ + N+ C+++ C+ +S+ N+I +
Sbjct: 861 TKHFQNMNENLNIGPPLWFVVEGDVKWHDPKMQNKFCTLAGCDDNSMGNKIRSLAYAENY 920
Query: 892 ETSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPCCAPEDDSCVSGACKD 951
+ +Y+ WLD +L ++ P G C K +C P + C SC S +
Sbjct: 921 KGNYLHGDVNIWLDSYLQFMHPR--GSCCKMDGKQFCDPSNATHC-----SSCSSSSVAS 973
Query: 952 CTTCFRHSDLRNDRTSTMQFRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGIIQA 1011
T T+ +F L FL PS CA GG +++L +G IQ+
Sbjct: 974 LT------------TTEYEFYRNLHHFLETPPSIQCAHGGMALAKPAINLT--RNGKIQS 1019
Query: 1012 SSFRTYHTPLN--KQVDYVNSMRAAREFSSKVSDSLKV 1047
+ F T+ LN + ++ R A+ + + L++
Sbjct: 1020 AYFSTFFKKLNLSDSIQLYDAWRFAKYLADDIERELEI 1057
Score = 44.3 bits (103), Expect = 0.002
Identities = 22/83 (26%), Positives = 46/83 (54%)
Query: 1052 KDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLV 1111
+++R AL + G++ SGI ++ L F+ + V + Y+ + + L+ +HG+V
Sbjct: 1167 REERAFAALVSNGSTTLSGIFPAIMITAGCLSFADSRVLITYFCNQLVGIGLVCAVHGVV 1226
Query: 1112 FLPVVLSIFGPPSRCTIIEQEEN 1134
++P +L+IFG + +EE+
Sbjct: 1227 YMPTLLAIFGSDFYQNVSSEEES 1249
>PTC1_MOUSE (Q61115) Patched protein homolog 1 (PTC1) (PTC)
Length = 1434
Score = 127 bits (318), Expect = 2e-28
Identities = 83/266 (31%), Positives = 134/266 (50%), Gaps = 21/266 (7%)
Query: 493 LDPSTVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMA 552
+ P + F G DY V++ E A AW++ ++++V + P
Sbjct: 353 MTPKQMYEHFRGYDY-----------VSHINWNEDRAAAILEAWQRTYVEVVHQSVAP-- 399
Query: 553 QSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKV 612
S L F++ +++++ LK S I + YL+M AY LT+ S +
Sbjct: 400 NSTQKVLPFTT-TTLDDILKSFSDVSVIRVASGYLLMLAYACLTMLRWDCSKS-----QG 453
Query: 613 LLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKR--Q 670
+GL+GV+LV LSV + + S +G+ +V+PFL L VGVD++ +L HA Q
Sbjct: 454 AVGLAGVLLVALSVAAGLGLCSLIGISFNAATTQVLPFLALGVGVDDVFLLAHAFSETGQ 513
Query: 671 PLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFVL 730
+P E R L G S+ L S+S V AF + + I +PA R FS+ AA+ V+ +F +
Sbjct: 514 NKRIPFEDRTGECLKRTGASVALTSISNVTAFFMAALIPIPALRAFSLQAAVVVVFNFAM 573
Query: 731 QVTAFVALIVLDSQRAEDKRVDCFPC 756
+ F A++ +D R ED+R+D F C
Sbjct: 574 VLLIFPAILSMDLYRREDRRLDIFCC 599
Score = 61.6 bits (148), Expect = 1e-08
Identities = 38/87 (43%), Positives = 51/87 (57%), Gaps = 1/87 (1%)
Query: 1036 EFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYF 1095
EF+ V+ + A GDK+ R AL M A V G T L+GV++L S + V Y+F
Sbjct: 1081 EFTVHVALAFLTAIGDKNHRAMLALEHMFAPVLDGAVST-LLGVLMLAGSEFDFIVRYFF 1139
Query: 1096 QMYLSLVLLGFLHGLVFLPVVLSIFGP 1122
+ L +LG L+GLV LPV+LS FGP
Sbjct: 1140 AVLAILTVLGVLNGLVLLPVLLSFFGP 1166
Score = 32.7 bits (73), Expect = 5.7
Identities = 22/86 (25%), Positives = 43/86 (49%), Gaps = 9/86 (10%)
Query: 778 LARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDS----YL 833
L+ + ++ +AP L K+VVI +F+ S+ +TR+ GL+ ++PR++ ++
Sbjct: 718 LSSFAEKHYAPFLLKPKAKVVVILLFLGLLGVSLYGTTRVRDGLDLTDIVPRETREYDFI 777
Query: 834 QGYFNNVSEYLRIGPPLYFVVKNYNY 859
F S Y +Y V + +Y
Sbjct: 778 AAQFKYFSFY-----NMYIVTQKADY 798
>PTC1_HUMAN (Q13635) Patched protein homolog 1 (PTC1) (PTC)
Length = 1447
Score = 124 bits (312), Expect = 1e-27
Identities = 75/233 (32%), Positives = 123/233 (52%), Gaps = 10/233 (4%)
Query: 526 EGNETAKAVAWEKAFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKRESTADAITILVS 585
E A AW++ ++++V + AQ+ + + +++++ LK S I +
Sbjct: 389 EDKAAAILEAWQRTYVEVVHQSV---AQNSTQKVLSFTTTTLDDILKSFSDVSVIRVASG 445
Query: 586 YLVMFAYISLTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIM 645
YL+M AY LT+ S + +GL+GV+LV LSV + + S +G+
Sbjct: 446 YLLMLAYACLTMLRWDCSKS-----QGAVGLAGVLLVALSVAAGLGLCSLIGISFNAATT 500
Query: 646 EVIPFLVLAVGVDNMCILVHAVKR--QPLELPLEGRISNALVEVGPSITLASLSEVLAFA 703
+V+PFL L VGVD++ +L HA Q +P E R L G S+ L S+S V AF
Sbjct: 501 QVLPFLALGVGVDDVFLLAHAFSETGQNKRIPFEDRTGECLKRTGASVALTSISNVTAFF 560
Query: 704 VGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPC 756
+ + I +PA R FS+ AA+ V+ +F + + F A++ +D R ED+R+D F C
Sbjct: 561 MAALIPIPALRAFSLQAAVVVVFNFAMVLLIFPAILSMDLYRREDRRLDIFCC 613
Score = 62.0 bits (149), Expect = 9e-09
Identities = 38/87 (43%), Positives = 52/87 (59%), Gaps = 1/87 (1%)
Query: 1036 EFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYF 1095
EF+ V+ + A GDK++R AL M A V G T L+GV++L S + V Y+F
Sbjct: 1095 EFTVHVALAFLTAIGDKNRRAVLALEHMFAPVLDGAVST-LLGVLMLAGSEFDFIVRYFF 1153
Query: 1096 QMYLSLVLLGFLHGLVFLPVVLSIFGP 1122
+ L +LG L+GLV LPV+LS FGP
Sbjct: 1154 AVLAILTILGVLNGLVLLPVLLSFFGP 1180
Score = 32.3 bits (72), Expect = 7.5
Identities = 22/86 (25%), Positives = 43/86 (49%), Gaps = 9/86 (10%)
Query: 778 LARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDS----YL 833
L+ + ++ +AP L K+VVI +F+ S+ +TR+ GL+ ++PR++ ++
Sbjct: 732 LSSFAEKHYAPFLLKPKAKVVVIFLFLGLLGVSLYGTTRVRDGLDLTDIVPRETREYDFI 791
Query: 834 QGYFNNVSEYLRIGPPLYFVVKNYNY 859
F S Y +Y V + +Y
Sbjct: 792 AAQFKYFSFY-----NMYIVTQKADY 812
>PTC1_CHICK (Q90693) Patched protein homolog 1 (PTC1) (PTC)
Length = 1442
Score = 124 bits (312), Expect = 1e-27
Identities = 75/233 (32%), Positives = 123/233 (52%), Gaps = 10/233 (4%)
Query: 526 EGNETAKAVAWEKAFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKRESTADAITILVS 585
E A AW++ ++++V + AQ+ + + +++++ LK S I +
Sbjct: 389 EDKAAAILEAWQRMYVEVVHQSV---AQNSTQKVLSFTTTTLDDILKSFSDVSVIRVASG 445
Query: 586 YLVMFAYISLTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIM 645
YL+M AY LT+ S+ +GL+GV+LV LSV + + S +G+
Sbjct: 446 YLLMLAYACLTMLRWD-----CAKSQGAVGLAGVLLVALSVAAGLGLCSLIGISFNAATT 500
Query: 646 EVIPFLVLAVGVDNMCILVHAVKR--QPLELPLEGRISNALVEVGPSITLASLSEVLAFA 703
+V+PFL L VGVD++ +L HA Q +P E R L G S+ L S+S V AF
Sbjct: 501 QVLPFLALGVGVDDVFLLAHAFSETGQNKRIPFEDRTGECLKRTGASVALTSISNVTAFF 560
Query: 704 VGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPC 756
+ + I +PA R FS+ AA+ V+ +F + + F A++ +D R ED+R+D F C
Sbjct: 561 MAALIPIPALRAFSLQAAVVVVFNFAMVLLIFPAILSMDLYRREDRRLDIFCC 613
Score = 61.6 bits (148), Expect = 1e-08
Identities = 37/87 (42%), Positives = 52/87 (59%), Gaps = 1/87 (1%)
Query: 1036 EFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYF 1095
EF+ ++ + A GDK++R AL M A V G T L+GV++L S + V Y+F
Sbjct: 1094 EFTVHIALAFLTAIGDKNRRAVLALEHMFAPVLDGAVST-LLGVLMLAGSEFDFIVRYFF 1152
Query: 1096 QMYLSLVLLGFLHGLVFLPVVLSIFGP 1122
+ L +LG L+GLV LPV+LS FGP
Sbjct: 1153 AVLAILTILGVLNGLVLLPVLLSFFGP 1179
Score = 33.5 bits (75), Expect = 3.4
Identities = 23/86 (26%), Positives = 43/86 (49%), Gaps = 9/86 (10%)
Query: 778 LARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDS----YL 833
L+ + ++ +AP L K+VVI +F+ S+ +TR+ GL+ ++PRD+ ++
Sbjct: 731 LSTFAEKHYAPFLLKPKAKVVVIFLFLGLLGLSLYGTTRVRDGLDLTDIVPRDTREYDFI 790
Query: 834 QGYFNNVSEYLRIGPPLYFVVKNYNY 859
F S Y +Y V + +Y
Sbjct: 791 AAQFKYFSFY-----NMYIVTQKADY 811
>PTC1_BRARE (Q98864) Patched protein homolog 1 (Patched 1) (PTC1)
Length = 1220
Score = 121 bits (303), Expect = 1e-26
Identities = 74/240 (30%), Positives = 129/240 (52%), Gaps = 8/240 (3%)
Query: 517 YPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKREST 576
Y +++ E TA +W++ F+++V + P S N+ AFS+ +++ + +K S
Sbjct: 363 YEIHDINWNEDKATAILESWQRKFVEVVHGSI-PQNSSSNV-YAFST-TTLNDIMKSFSD 419
Query: 577 ADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSAL 636
I + YL+M AY +T+ S+ +GL+GV+LV LSV + + S L
Sbjct: 420 VSVIRVAGGYLLMLAYACVTMLRWD-----CAKSQGAVGLAGVLLVALSVAAGLGLCSLL 474
Query: 637 GVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELPLEGRISNALVEVGPSITLASL 696
G+ +V+P L L +GVD+M +L H+ +P + R + L G S+ L S+
Sbjct: 475 GLSFNAATTQVLPSLALGIGVDDMFLLGHSFTETRSNIPFKERTGDCLRRTGTSVALTSV 534
Query: 697 SEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPC 756
+ ++AF + + + +PA R FS+ AA+ V+ +F + + F A++ LD R EDKR+D C
Sbjct: 535 NNMIAFFMAALVPIPALRAFSLQAAVVVVFNFAMALLIFPAILSLDLHRREDKRLDILCC 594
Score = 60.1 bits (144), Expect = 3e-08
Identities = 34/89 (38%), Positives = 53/89 (59%), Gaps = 1/89 (1%)
Query: 1036 EFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYF 1095
EF+ ++ A GD++ R A+ M A V G ++ L+GV++L S + + Y+F
Sbjct: 1084 EFTVHIALGFLTAIGDRNTRSAVAMEHMFAPVIDG-AISTLLGVLMLAGSEFDFIMRYFF 1142
Query: 1096 QMYLSLVLLGFLHGLVFLPVVLSIFGPPS 1124
+ L LLG L+GLV LPV+LS+ GPP+
Sbjct: 1143 AVLAILTLLGILNGLVLLPVLLSLMGPPA 1171
Score = 35.8 bits (81), Expect = 0.68
Identities = 22/92 (23%), Positives = 41/92 (43%)
Query: 778 LARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYF 837
L+ + +E +AP+L K VV+ +FVA S+ +T + GL ++PRD+ +
Sbjct: 721 LSSFAREKYAPLLLKPETKTVVVVVFVALLSLSLYGTTMVHDGLYLTDIVPRDTQEYEFI 780
Query: 838 NNVSEYLRIGPPLYFVVKNYNYSSESTHTNQL 869
+Y + ++Y+ QL
Sbjct: 781 TAQFKYFSFYNMYLVTMDGFDYARSQRQLLQL 812
Score = 35.0 bits (79), Expect = 1.2
Identities = 29/123 (23%), Positives = 47/123 (37%), Gaps = 3/123 (2%)
Query: 319 FYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVETRPEKLWVGPGSKAAQEKQFFDS 378
F G + RH VL + L + L +GL +ET EKLWV GS+ ++E ++
Sbjct: 70 FLFSLGCHIQRHCGKVLFIGLLVFGALSVGLRVAAIETDIEKLWVEAGSRVSKELRYTKE 129
Query: 379 HLAPFYRIEQLILATVPDHMNSTSPRIVSADNIRFLFEVQKKVDAIRANYSGLMVSLQDI 438
+L P I++ + + E ++ + G L I
Sbjct: 130 KQGEESVFTSQMLIQTP---KQEGTNILTQEALLLHLEAALSASKVQVSLYGKSWDLNKI 186
Query: 439 CMK 441
C K
Sbjct: 187 CFK 189
>PTC2_MOUSE (O35595) Patched protein homolog 2 (PTC2)
Length = 1182
Score = 118 bits (295), Expect = 1e-25
Identities = 71/222 (31%), Positives = 124/222 (54%), Gaps = 8/222 (3%)
Query: 535 AWEKAFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYIS 594
AW++ F+QL + E LP S+ + AFSS +++++ L+ S ++ YL+M AY
Sbjct: 354 AWQRRFVQLAQ-EALPANASQQIH-AFSS-TTLDDILRAFSEVSTTRVVGGYLLMLAYAC 410
Query: 595 LTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLA 654
+T+ S+ +GL+GV+LV L+V + + + LG+ +V+PFL L
Sbjct: 411 VTMLRWD-----CAQSQGAVGLAGVLLVALAVASGLGLCALLGITFNAATTQVLPFLALG 465
Query: 655 VGVDNMCILVHAVKRQPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACR 714
+GVD++ +L HA + P + PL R+ L G S+ L S++ ++AF + + + +PA R
Sbjct: 466 IGVDDIFLLAHAFTKAPPDTPLPERMGECLRSTGTSVALTSVNNMVAFFMAALVPIPALR 525
Query: 715 VFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPC 756
FS+ AA+ V +F + F A++ LD +R +R+D C
Sbjct: 526 AFSLQAAIVVGCNFAAVMLVFPAILSLDLRRRHRQRLDVLCC 567
Score = 60.5 bits (145), Expect = 3e-08
Identities = 38/113 (33%), Positives = 60/113 (52%), Gaps = 1/113 (0%)
Query: 1028 VNSMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRT 1087
V S+ EF+ V+ + G ++ R AL A V G T L+G+++L S
Sbjct: 1023 VASIGIGVEFTVHVALGFLTSHGSRNLRAASALEQTFAPVTDGAVST-LLGLLMLAGSNF 1081
Query: 1088 EVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGPPSRCTIIEQEENRSSTSS 1140
+ + Y+F + L LLG LHGL+ LPV+LSI GPP + + +E ++ S+
Sbjct: 1082 DFIIRYFFVVLTVLTLLGLLHGLLLLPVLLSILGPPPQVVQVYKESPQTLNSA 1134
Score = 35.0 bits (79), Expect = 1.2
Identities = 32/132 (24%), Positives = 49/132 (36%), Gaps = 7/132 (5%)
Query: 312 VQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVETRPEKLWVGPGSKAAQ 371
++ Y G + +H VL L L L LGL +ET E+LWV GS+ +Q
Sbjct: 36 LRAYFQGLLFSLGCRIQKHCGKVLFLGLVAFGALALGLRVAVIETDLEQLWVEVGSRVSQ 95
Query: 372 EKQFFDSHLA--PFYRIEQLILATVPDHMNSTSPRIVSADNIRFLFEVQKKVDAIRANYS 429
E + L Y + LI + N +P + + ++ +
Sbjct: 96 ELHYTKEKLGEEAAYTSQMLIQTAHQEGGNVLTPEALDLH-----LQAALTASKVQVSLY 150
Query: 430 GLMVSLQDICMK 441
G L IC K
Sbjct: 151 GKSWDLNKICYK 162
>PTC2_HUMAN (Q9Y6C5) Patched protein homolog 2 (PTC2)
Length = 1203
Score = 113 bits (283), Expect = 3e-24
Identities = 72/222 (32%), Positives = 121/222 (54%), Gaps = 8/222 (3%)
Query: 535 AWEKAFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYIS 594
AW++ F+QL + E LP S+ + AFSS +++++ L S A ++ YL+M AY
Sbjct: 354 AWQRRFVQLAQ-EALPENASQQIH-AFSS-TTLDDILHAFSEVSAARVVGGYLLMLAYAC 410
Query: 595 LTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLA 654
+T+ S+ +GL+GV+LV L+V + + + LG+ +V+PFL L
Sbjct: 411 VTMLRWD-----CAQSQGSVGLAGVLLVALAVASGLGLCALLGITFNAATTQVLPFLALG 465
Query: 655 VGVDNMCILVHAVKRQPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACR 714
+GVD++ +L HA PL+ R+ L G S+ L S++ + AF + + + +PA R
Sbjct: 466 IGVDDVFLLAHAFTEALPGTPLQERMGECLQRTGTSVVLTSINNMAAFLMAALVPIPALR 525
Query: 715 VFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPC 756
FS+ AA+ V FV + F A++ LD +R +R+D C
Sbjct: 526 AFSLQAAIVVGCTFVAVMLVFPAILSLDLRRRHCQRLDVLCC 567
Score = 58.9 bits (141), Expect = 7e-08
Identities = 37/96 (38%), Positives = 51/96 (52%), Gaps = 1/96 (1%)
Query: 1028 VNSMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRT 1087
V S+ EF+ V+ G ++ R AL A V G ++ L+G+++L S
Sbjct: 1023 VASVGIGVEFTVHVALGFLTTQGSRNLRAAHALEHTFAPVTDG-AISTLLGLLMLAGSHF 1081
Query: 1088 EVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGPP 1123
+ V Y+F L LLG LHGLV LPV+LSI GPP
Sbjct: 1082 DFIVRYFFAALTVLTLLGLLHGLVLLPVLLSILGPP 1117
Score = 36.6 bits (83), Expect = 0.40
Identities = 33/132 (25%), Positives = 49/132 (37%), Gaps = 7/132 (5%)
Query: 312 VQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVETRPEKLWVGPGSKAAQ 371
++ Y G + RH VL L L L LGL +ET E+LWV GS+ +Q
Sbjct: 36 LRAYFQGLLFSLGCGIQRHCGKVLFLGLLAFGALALGLRMAIIETNLEQLWVEVGSRVSQ 95
Query: 372 EKQFFDSHLA--PFYRIEQLILATVPDHMNSTSPRIVSADNIRFLFEVQKKVDAIRANYS 429
E + L Y + LI + N +P + + ++ +
Sbjct: 96 ELHYTKEKLGEEAAYTSQMLIQTARQEGENILTPEALGLH-----LQAALTASKVQVSLY 150
Query: 430 GLMVSLQDICMK 441
G L IC K
Sbjct: 151 GKSWDLNKICYK 162
>SCAP_HUMAN (Q12770) Sterol regulatory element binding protein
cleavage-activating protein (SREBP cleavage-activating
protein) (SCAP)
Length = 1278
Score = 99.0 bits (245), Expect = 7e-20
Identities = 113/482 (23%), Positives = 209/482 (42%), Gaps = 65/482 (13%)
Query: 316 MSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVE-------TRPEKLWVGP--- 365
+S + +G L A +PI ++ T +L C L++ + T P K + P
Sbjct: 11 ISRAFYNHGLLCASYPIPIILFTGFCILACCYPLLKLPLPGTGPVEFTTPVKDYSPPPVD 70
Query: 366 -----GSKAAQEKQFFDSHLAPFYRIEQLIL--ATVPDHMNSTSPRIVSADNIRFLFEVQ 418
G Q + + AP ++Q+ + + P H N + + + R V+
Sbjct: 71 SDRKQGEPTEQPEWYVG---APVAYVQQIFVKSSVFPWHKNLLAVDVFRSPLSRAFQLVE 127
Query: 419 KKVDAIRANYSGLMVSLQDICMKPLD--------------KDCATQSVLQYFKMDPRNFD 464
+ + + + SG+ SL+++C++ D C S +++ D F
Sbjct: 128 EIRNHVLRDSSGIR-SLEELCLQVTDLLPGLRKLRNLLPEHGCLLLSPGNFWQNDWERFH 186
Query: 465 DSGAVEHLNYCFQQYSSADQCMSAFKAPLDPSTVLGGFSGKDYSGASAFIVTYPVNNAID 524
+ + Q Q + K +L G GK YSG S + V+ I
Sbjct: 187 ADPDI--IGTIHQHEPKTLQTSATLK------DLLFGVPGK-YSGVSLYTRKRMVSYTI- 236
Query: 525 EEGNETAKAVAWEKAFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKRE-STADAITIL 583
T + F+ ++ L+ + S N +L +ES + K E A+ I ++
Sbjct: 237 -----TLVFQHYHAKFLGSLRARLMLLHPSPNCSLR--AESLVHVHFKEEIGVAELIPLV 289
Query: 584 VSYLVMFAYISLTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVKSTLI 643
+Y+++FAYI + + SK L L+ V+ V+ S+L SV + + G+ TL
Sbjct: 290 TTYIILFAYIYFSTRKID-----MVKSKWGLALAAVVTVLSSLLMSVGLCTLFGLTPTLN 344
Query: 644 IMEVIPFLVLAVGVDNMCILVHAVKRQPLELPLEGRISNALVEVGPSITLASLSEVLAFA 703
E+ P+LV+ +G++N+ +L +V P++L ++ RI+ L SI +E+
Sbjct: 345 GGEIFPYLVVVIGLENVLVLTKSVVSTPVDLEVKLRIAQGLSSESWSIMKNMATELGIIL 404
Query: 704 VGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAE----DKRVD---CFPC 756
+G F +PA + F +FA + ++ DF LQ+ F ++ +D +R E +KR+ C P
Sbjct: 405 IGYFTLVPAIQEFCLFAVVGLVSDFFLQMLFFTTVLSIDIRRMELADLNKRLPPEACLPS 464
Query: 757 IK 758
K
Sbjct: 465 AK 466
>SCAP_CRIGR (P97260) Sterol regulatory element binding protein
cleavage-activating protein (SREBP cleavage-activating
protein) (SCAP)
Length = 1276
Score = 96.3 bits (238), Expect = 4e-19
Identities = 112/482 (23%), Positives = 208/482 (42%), Gaps = 65/482 (13%)
Query: 316 MSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVE-------TRPEKLWVGP--- 365
+S + +G L A +PI ++ T +L C L++ + + P K + P
Sbjct: 11 ISQAFYNHGLLCASYPIPIILFTGLCILACCYPLLKLPLPGTGPVEFSTPVKDYSPPPVD 70
Query: 366 -----GSKAAQEKQFFDSHLAPFYRIEQLIL--ATVPDHMNSTSPRIVSADNIRFLFEVQ 418
G + Q + + AP I+Q+ + + P H N + + R V+
Sbjct: 71 SDHKQGEPSEQPEWYVG---APVAYIQQIFVKSSVSPWHKNLLAVDVFRLPLSRAFQLVE 127
Query: 419 KKVDAIRANYSGLMVSLQDICMKPLD--------------KDCATQSVLQYFKMDPRNFD 464
+ + + + SG SL+++C++ D C S +++ D F
Sbjct: 128 EIRNHVLRDSSGTK-SLEEVCLQVTDLLPGLRKLRNLLPEHGCLLLSPGNFWQNDWERFH 186
Query: 465 DSGAVEHLNYCFQQYSSADQCMSAFKAPLDPSTVLGGFSGKDYSGASAFIVTYPVNNAID 524
+ + Q Q + K +L G GK YSG S + V+ I
Sbjct: 187 ADPDI--IGTIHQHEPKTLQTSATLK------DLLFGVPGK-YSGVSLYTRKRTVSYTI- 236
Query: 525 EEGNETAKAVAWEKAFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKRE-STADAITIL 583
T + F+ ++ L+ + S N +L +E+ + K E A+ I ++
Sbjct: 237 -----TLVFQRYHAKFLSSLRARLMLLHPSPNCSLR--AENLVHVHFKEEIGIAELIPLV 289
Query: 584 VSYLVMFAYISLTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVKSTLI 643
+Y+++FAYI + + SK L L+ V+ V+ S+L SV + + G+ TL
Sbjct: 290 TTYIILFAYIYFSTRKID-----MVKSKWGLALAAVVTVLSSLLMSVGLCTLFGLTPTLN 344
Query: 644 IMEVIPFLVLAVGVDNMCILVHAVKRQPLELPLEGRISNALVEVGPSITLASLSEVLAFA 703
E+ P+LV+ +G++N+ +L +V P++L ++ RI+ L SI +E+
Sbjct: 345 GGEIFPYLVVVIGLENVLVLTKSVVSTPVDLEVKLRIAQGLSSESWSIMKNVATELGIIL 404
Query: 704 VGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAE----DKRV---DCFPC 756
+G F +PA + F +FA + ++ DF LQ+ F ++ +D +R E +KR+ C P
Sbjct: 405 IGYFTLVPAIQEFCLFAVVGLVSDFFLQMFFFTTVLSIDIRRMELADLNKRLPPESCLPS 464
Query: 757 IK 758
K
Sbjct: 465 AK 466
>PATC_DROME (P18502) Patched protein (Hedgehog receptor)
Length = 1286
Score = 93.6 bits (231), Expect = 3e-18
Identities = 87/344 (25%), Positives = 156/344 (45%), Gaps = 59/344 (17%)
Query: 535 AWEKAFIQLVKDELLPMAQ-SRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYI 593
AW++ F + V+ L ++ + N + S +++++ L + S A++I++ V Y
Sbjct: 384 AWQRNFSREVEQLLRKQSRIATNYDIYVFSSAALDDILAKFSHPSALSIVIGVAVTVLYA 443
Query: 594 SLTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVL 653
TL P + + +G++GV+L+ S + + + LG+ +V+PFL L
Sbjct: 444 FCTLLRWRDP----VRGQSSVGVAGVLLMCFSTAAGLGLSALLGIVFNAASTQVVPFLAL 499
Query: 654 AVGVDNMCILVHAV----KRQPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFIS 709
+GVD++ +L A +R+ +L L+ +VGPSI ++ S +F +FI
Sbjct: 500 GLGVDHIFMLTAAYAESNRREQTKLILK--------KVGPSILFSACSTAGSFFAAAFIP 551
Query: 710 MPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVD----CFPCIKVHSFHAD 765
+PA +VF + AA+ + + + F A+I LD +R R D CFP K A
Sbjct: 552 VPALKVFCLQAAIVMCSNLAAALLVFPAMISLDLRRRTAGRADIFCCCFPVWKEQPKVAP 611
Query: 766 P--------DKGIRQRK----------------------PG--------LLARYMKEVHA 787
P +G R K PG LA + + +
Sbjct: 612 PVLPLNNNNGRGARHPKSCNNNRVPLPAQNPLLEQRADIPGSSHSLASFSLATFAFQHYT 671
Query: 788 PILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDS 831
P L VK + + F+A ++S+ STR++ GL+ ++P+DS
Sbjct: 672 PFLMRSWVKFLTVMGFLAALISSLYASTRLQDGLDIIDLVPKDS 715
Score = 37.7 bits (86), Expect = 0.18
Identities = 35/166 (21%), Positives = 68/166 (40%), Gaps = 12/166 (7%)
Query: 281 LYARNQEKDENLPMQMIED--VPQNRNGVRL-SVVQGYMSNFYRKYGSLVARHPINVLAL 337
LY R D + + I+ +R + L SV Q ++ GS V +H VL +
Sbjct: 25 LYIRTSWVDAQVALDQIDKGKARGSRTAIYLRSVFQSHLETL----GSSVQKHAGKVLFV 80
Query: 338 TLAIVLLLCLGLIRFKVETRPEKLWVGPGSKAAQEKQFFDSHLA-PFYRIEQLILATVPD 396
+ ++ C+GL ++ ++ +LW+ G + E + + QL++ T D
Sbjct: 81 AILVLSTFCVGLKSAQIHSKVHQLWIQEGGRLEAELAYTQKTIGEDESATHQLLIQTTHD 140
Query: 397 HMNSTSPRIVSADNIRFLFEVQKKVDAIRANYSGLMVSLQDICMKP 442
+ ++ + EV K A++ + L+D+C P
Sbjct: 141 ----PNASVLHPQALLAHLEVLVKATAVKVHLYDTEWGLRDMCNMP 182
Score = 35.8 bits (81), Expect = 0.68
Identities = 24/103 (23%), Positives = 52/103 (50%), Gaps = 1/103 (0%)
Query: 1037 FSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQ 1096
F+ +S + G++ +RV+ ++ + G+ LT V V +L S E + ++
Sbjct: 1024 FNVLISLGFMTSVGNRQRRVQLSMQMSLGPLVHGM-LTSGVAVFMLSTSPFEFVIRHFCW 1082
Query: 1097 MYLSLVLLGFLHGLVFLPVVLSIFGPPSRCTIIEQEENRSSTS 1139
+ L ++ +G + L+ P++LS+ GP + +E + S+ S
Sbjct: 1083 LLLVVLCVGACNSLLVFPILLSMVGPEAELVPLEHPDRISTPS 1125
>PTC1_CAEEL (Q09614) Patched protein homolog 1
Length = 1405
Score = 86.7 bits (213), Expect = 3e-16
Identities = 86/371 (23%), Positives = 162/371 (43%), Gaps = 44/371 (11%)
Query: 528 NETAKAV---AWEKAFIQLVKDELLPMAQSRN--LTLAFSSESSIEEELKRESTADAITI 582
NETA AW++ F + + + + + N TL + +SI + L+ + I
Sbjct: 596 NETAAEQVLQAWQRNFTKSLYNHKANVDEDGNERRTLHPLASTSIADMLEEFCQFNYTII 655
Query: 583 LVSYLVMFAYISLTLG--DTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVKS 640
L Y +M AY +T D P++ S + L L+GV++V + + + + + G++
Sbjct: 656 LAGYALMLAYAIVTQARFDNCLPAT---ESSMGLALAGVLVVTFASVAGLGLATWFGIEF 712
Query: 641 TLIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELPLEGRISNALVEVGPSITLASLSEVL 700
+++PFL L +GVDNM +L+H + ++ + E G SI S++ +L
Sbjct: 713 NAATTQIVPFLTLGIGVDNMFMLLHNYRDVVKLAGGHAEMAILMRETGMSILCTSINNIL 772
Query: 701 AFAVGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPCI--- 757
+F G+ + +PA R F +++ + +F+ +T + A+I +D +R + +R D C+
Sbjct: 773 SFLTGTLLPIPALRSFCAQSSILLTFNFIAILTIYPAIISIDLRRKKAQRRDLVCCLYGD 832
Query: 758 ------------KVHS---FHADPDKGIRQRKPGL---------------LARYMKEVHA 787
K+ S A + I Q+ G+ L +++ +
Sbjct: 833 TREESYSMISKPKIQSKRIIGAPSEASIMQQFDGITQAQMASSDDPAPWSLHSFIRYYYI 892
Query: 788 PILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIG 847
P +S K+ +I A AS + GLE VLP + + +Y
Sbjct: 893 PFISKPASKVAIIVGCCALLGASFIGMRQSTLGLELGDVLPEHTAPAQFLRARDKYFSF- 951
Query: 848 PPLYFVVKNYN 858
P++ V+K N
Sbjct: 952 YPMFAVIKGPN 962
Score = 58.9 bits (141), Expect = 7e-08
Identities = 33/98 (33%), Positives = 55/98 (55%), Gaps = 1/98 (1%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYF 1084
V + ++ EF+ V S A G + QR A+ + V G + + L+G+++L F
Sbjct: 1236 VTLITAVGIGVEFTVHVVVSFLTALGTRSQRTSSAVDRVFVPVIHG-SFSTLLGILMLGF 1294
Query: 1085 SRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGP 1122
S E V Y+F + +L+ +G ++GL+ LPV+LS FGP
Sbjct: 1295 SEFEFVVKYFFIVMTALICIGIINGLILLPVLLSWFGP 1332
>YMB5_CAEEL (Q03602) Hypothetical protein F54G8.5 in chromosome III
Length = 413
Score = 67.0 bits (162), Expect = 3e-10
Identities = 48/159 (30%), Positives = 83/159 (52%), Gaps = 6/159 (3%)
Query: 677 EGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVTAFV 736
E R +AL E S+ L SL++ L+FA+GS A RVF + A+A+L F+ QVT F
Sbjct: 9 EQRYIHALTESAASLFLTSLTDGLSFAIGSISDFHAVRVFCTYCAMAILFMFLFQVTFFN 68
Query: 737 ALIVLDSQRAEDKRVDCFPCIKVHSFHADP--DKGIRQRKPGLLARYMKEVHAPILSIWG 794
A++ L +R + F C ++ +K ++A+++ ++ P+ S
Sbjct: 69 AVMSLCCRREVSGKHPVFCCYEITPPEETQVLEKQSNFDFSSIVAKHLFKILNPLPS--- 125
Query: 795 VKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYL 833
+I V IF+A+ SI + + GL+ +++ P DSY+
Sbjct: 126 -RIGVFFIFLAYLFISIHFAIGLPLGLDLKLLAPDDSYV 163
>HMDH_STRPU (P16393) 3-hydroxy-3-methylglutaryl-coenzyme A reductase
(EC 1.1.1.34) (HMG-CoA reductase)
Length = 932
Score = 54.7 bits (130), Expect = 1e-06
Identities = 33/132 (25%), Positives = 66/132 (50%), Gaps = 1/132 (0%)
Query: 610 SKVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKR 669
SK +LG++G+ + S L S A+ G++ T + E +PF +L + + L
Sbjct: 89 SKYILGIAGLFTIFSSFLFSSAVIHLFGLELTGL-NEALPFFLLLIDLTKASALTKFALS 147
Query: 670 QPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFV 729
+ + I+ + +GP+ITL ++ L ++G+ S+ VF F L+++ ++
Sbjct: 148 STTQNEVVDNIARGMAILGPTITLDTVVTTLVISIGTMSSIRKMEVFCCFGILSLIANYF 207
Query: 730 LQVTAFVALIVL 741
+ +T F A + L
Sbjct: 208 VFMTFFPACLSL 219
>HMDH_PHYBL (Q12649) 3-hydroxy-3-methylglutaryl-coenzyme A reductase
(EC 1.1.1.34) (HMG-CoA reductase)
Length = 1176
Score = 52.4 bits (124), Expect = 7e-06
Identities = 46/183 (25%), Positives = 84/183 (45%), Gaps = 15/183 (8%)
Query: 567 IEEELKRESTADAITILVSYLVMFA-YISLTLGDTPHPSSFYISSKVLL-GLSGVILVML 624
++E + D I ILV Y++M A +ISL + S + +++ V+ G +L +L
Sbjct: 290 VKELIDLADNIDIIVILVGYIMMIATFISLYVNMRAMGSRYTLATAVVFNGFFSFMLALL 349
Query: 625 SVLGSVAIFSALGVKS-TLIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELPLEGR---- 679
+V ALGV +++ E IPFL + +G + L V + E PL +
Sbjct: 350 TV-------RALGVDVYPVVLAEAIPFLAVTIGFERPFKLTKRVFQFSKETPLTKQEIRT 402
Query: 680 -ISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVTAFVAL 738
I A+ V I E++ +G+ + F + +A+ + DF++ T + A+
Sbjct: 403 TIMRAVDTVALPIARDCFMEIIVLVLGAKSGISGLEEFCLLSAILLAYDFIIMFTWYTAV 462
Query: 739 IVL 741
+ L
Sbjct: 463 LAL 465
>HMDH_BLAGE (P54960) 3-hydroxy-3-methylglutaryl-coenzyme A reductase
(EC 1.1.1.34) (HMG-CoA reductase)
Length = 856
Score = 52.0 bits (123), Expect = 9e-06
Identities = 31/134 (23%), Positives = 67/134 (49%), Gaps = 1/134 (0%)
Query: 608 ISSKVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAV 667
+ SK +LG++G+ V S + S ++ + LG + + + F +L + + +L
Sbjct: 84 LGSKYILGIAGLFTVFSSFVFSSSVINFLG-SDVSDLKDALFFFLLLIDLSKATVLAQFA 142
Query: 668 KRQPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLD 727
+ ++ I+ + +GP+ITL ++ E L VG + V FA ++V+++
Sbjct: 143 LSSRSQDEVKHNIARGIAMLGPTITLDTVVETLVIGVGMLSGVRRLEVLCCFACMSVIVN 202
Query: 728 FVLQVTAFVALIVL 741
+V+ +T + A + L
Sbjct: 203 YVVFMTFYPACLSL 216
>HMDH_XENLA (P20715) 3-hydroxy-3-methylglutaryl-coenzyme A reductase
(EC 1.1.1.34) (HMG-CoA reductase)
Length = 883
Score = 51.6 bits (122), Expect = 1e-05
Identities = 36/175 (20%), Positives = 75/175 (42%), Gaps = 6/175 (3%)
Query: 576 TADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSA 635
++D I + ++ + YI + + SK +LG++G+ + S + S +
Sbjct: 59 SSDIIILTITRCIAILYIYFQFQNLRQ-----LGSKYILGIAGLFTIFSSFVFSTVVIHF 113
Query: 636 LGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELPLEGRISNALVEVGPSITLAS 695
L K + E +PF +L + + L + + I+ + +GP+ TL +
Sbjct: 114 LD-KELTGLNEALPFFLLLIDLSKASALAKFALSSNSQDEVRDNIARGMAILGPTFTLEA 172
Query: 696 LSEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKR 750
L E L VG+ + + F ++VL ++ +T F A + L + + + R
Sbjct: 173 LVECLVIGVGTMSGVRQLEIMCCFGCMSVLANYFAFMTFFPACVSLVLELSRESR 227
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 128,621,115
Number of Sequences: 164201
Number of extensions: 5338845
Number of successful extensions: 13902
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 34
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 13708
Number of HSP's gapped (non-prelim): 136
length of query: 1140
length of database: 59,974,054
effective HSP length: 121
effective length of query: 1019
effective length of database: 40,105,733
effective search space: 40867741927
effective search space used: 40867741927
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0193.4


