

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
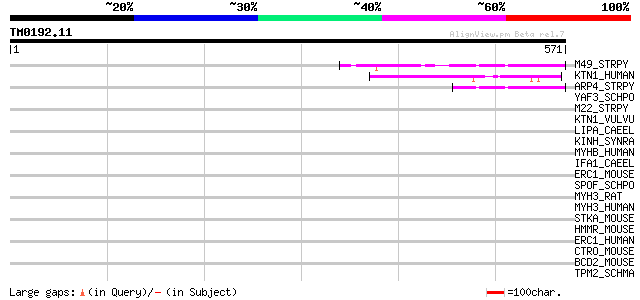
Query= TM0192.11
(571 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
M49_STRPY (P16947) M protein, serotype 49 precursor 47 1e-04
KTN1_HUMAN (Q86UP2) Kinectin (Kinesin receptor) (CG-1 antigen) 46 2e-04
ARP4_STRPY (P13050) IgA receptor precursor 44 9e-04
YAF3_SCHPO (Q09857) Hypothetical protein C29E6.03c in chromosome I 44 0.001
M22_STRPY (P50469) M protein, serotype 2.2 precursor 42 0.006
KTN1_VULVU (O97961) Kinectin 42 0.006
LIPA_CAEEL (Q21049) Liprin alpha (LAR-interacting protein alpha)... 41 0.007
KINH_SYNRA (O43093) Kinesin heavy chain (Synkin) 41 0.010
MYHB_HUMAN (P35749) Myosin heavy chain, smooth muscle isoform (S... 40 0.016
IFA1_CAEEL (P90901) Intermediate filament protein ifa-1 (Interme... 40 0.016
ERC1_MOUSE (Q99MI1) ERC protein 1 (Rab6-interacting protein 2) (... 40 0.016
SPOF_SCHPO (Q10411) Sporulation-specific protein 15 40 0.021
MYH3_RAT (P12847) Myosin heavy chain, fast skeletal muscle, embr... 39 0.028
MYH3_HUMAN (P11055) Myosin heavy chain, fast skeletal muscle, em... 39 0.028
STKA_MOUSE (O55098) Serine/threonine-protein kinase 10 (EC 2.7.1... 39 0.037
HMMR_MOUSE (Q00547) Hyaluronan mediated motility receptor (Intra... 39 0.037
ERC1_HUMAN (Q8IUD2) ERC protein 1 (ELKS protein) 39 0.037
CTRO_MOUSE (P49025) Citron protein (Rho-interacting, serine/thre... 39 0.037
BCD2_MOUSE (Q921C5) Cytoskeleton-like bicaudal D protein homolog 2 39 0.048
TPM2_SCHMA (P42638) Tropomyosin 2 (TMII) 38 0.062
>M49_STRPY (P16947) M protein, serotype 49 precursor
Length = 389
Score = 47.0 bits (110), Expect = 1e-04
Identities = 58/239 (24%), Positives = 106/239 (44%), Gaps = 31/239 (12%)
Query: 340 KRHQRDNEVCENSNPVRQGTGEKESRTALREKEGHFD-----LAEKEARPNLGEKEKEVP 394
K+HQ + E RQ E+ R RE E + + E + E ++
Sbjct: 115 KKHQEYKQEQEE----RQKNQEQLERKYQREVEKRYQEQLQKQQQLETEKQISEASRKSL 170
Query: 395 SDPSTMLRDATDHVEFLLSRIN--HDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQ 452
S R+A VE L+ + H L++E K I DA ++ ++ +
Sbjct: 171 SRDLEASREAKKKVEADLAALTAEHQKLKEE---KQISDASRQGLS-------------R 214
Query: 453 KFGASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARF 512
AS +K++AD AL EH K + +K +++A+R ++ D + A + VEA
Sbjct: 215 DLEASREAKKKVEADLAALTAEHQKLKE--EKQISDASRQGLSRDLEASREAKKKVEA-- 270
Query: 513 ELVCRSERITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDLLKKK 571
+L + ++ LE+ KE ++ +L+ K + A + E +L+ +L ++ +L K K
Sbjct: 271 DLAEANSKLQALEKLNKELEEGKKLSEKEKAELQARLEAEAKALKEQLAKQAEELAKLK 329
>KTN1_HUMAN (Q86UP2) Kinectin (Kinesin receptor) (CG-1 antigen)
Length = 1357
Score = 46.2 bits (108), Expect = 2e-04
Identities = 56/221 (25%), Positives = 101/221 (45%), Gaps = 32/221 (14%)
Query: 371 KEGHFDLAEKEARPNLGEKEKE--VPSDPSTMLRDATDHVEFLLS-RINHDYLEKEVFGK 427
K+ L E++ + + K +E + +D + L+ ++ L+S + N D +E+
Sbjct: 678 KDDKIRLLEEQLQHEISNKMEEFKILNDQNKALKSEVQKLQTLVSEQPNKDVVEQMEKCI 737
Query: 428 GIDDAGQECVASLFHTGCI-FAHAFQKFGASTAEGEKLQADYTALRTEH------AKCDD 480
D + V L TG I A ++ A E L + L+ + A +
Sbjct: 738 QEKDEKLKTVEELLETGLIQVATKEEELNAIRTENSSLTKEVQDLKAKQNDQVSFASLVE 797
Query: 481 RLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSER-ITDLEEEIKETKQEA---- 535
L KV+ E D K+ S+ EL+EA V E+ + DL++EIK K+E
Sbjct: 798 ELKKVIHEK-------DGKIKSV-EELLEAELLKVANKEKTVQDLKQEIKALKEEIGNVQ 849
Query: 536 -----QLALAAK----DDALALKDQELTSLRAELEEKKRDL 567
QL++ +K + L K++++ +++A LEEK++DL
Sbjct: 850 LEKAQQLSITSKVQELQNLLKGKEEQMNTMKAVLEEKEKDL 890
>ARP4_STRPY (P13050) IgA receptor precursor
Length = 386
Score = 44.3 bits (103), Expect = 9e-04
Identities = 33/116 (28%), Positives = 62/116 (53%), Gaps = 4/116 (3%)
Query: 456 ASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELV 515
AS +K++AD AL EH K + DK +++A+R ++ D + A + VEA +L
Sbjct: 215 ASREAKKKVEADLAALTAEHQKLKE--DKQISDASRQGLSRDLEASREAKKKVEA--DLA 270
Query: 516 CRSERITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDLLKKK 571
+ ++ LE+ KE ++ +L+ K + A + E +L+ +L ++ +L K K
Sbjct: 271 EANSKLQALEKLNKELEEGKKLSEKEKAELQARLEAEAKALKEQLAKQAEELAKLK 326
>YAF3_SCHPO (Q09857) Hypothetical protein C29E6.03c in chromosome I
Length = 1044
Score = 43.9 bits (102), Expect = 0.001
Identities = 54/230 (23%), Positives = 96/230 (41%), Gaps = 19/230 (8%)
Query: 335 ENPKKKRHQRDNEVCENSNPVRQGTGEKESRTALREKEGHFDLAEKEARPNLGEKEKEVP 394
E+ KK N + E N ++ + + +RE E L++K NLG KE +
Sbjct: 749 ESKNKKLENDLNLLTEKLN--KKNADTESFKNTIREAE----LSKKALNDNLGNKENII- 801
Query: 395 SDPSTMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQKF 454
SD L + + ++ L S++N D + E + I A E L I + +
Sbjct: 802 SDLKNKLSEESTRLQELQSQLNQDKNQIETLNERISAAADE----LSSMESINKNQANEL 857
Query: 455 GASTAEGEKLQADYT---ALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEAR 511
+ + LQ L EH + L+K L A + T+ K+L ++ +E
Sbjct: 858 KLAKQKCSNLQEKINFGNKLAKEHTEKISSLEKDLEAATKTASTLSKELKTVKSE----N 913
Query: 512 FELVCRSERITDLEEEIKETK-QEAQLALAAKDDALALKDQELTSLRAEL 560
L S + E+ + K +E ALA ++ L +D+E+ L+ ++
Sbjct: 914 DSLKSVSNDDQNKEKSVNNEKFKEVSQALAEANEKLNARDEEIERLKVDI 963
>M22_STRPY (P50469) M protein, serotype 2.2 precursor
Length = 372
Score = 41.6 bits (96), Expect = 0.006
Identities = 65/244 (26%), Positives = 103/244 (41%), Gaps = 27/244 (11%)
Query: 326 AKFPPNFNTENPKKKRHQRDN-EVCENSNPVRQGTGEKESRTALREKEGHFDLAEKEARP 384
AK+ N E K K+ + N E+ EN ++ G E +EKE K +
Sbjct: 75 AKYLEKINAEEEKNKKLEAINKELNENYYKLQDGIDALE-----KEKEDLKTTLAKTTKE 129
Query: 385 N-LGEKEKEVPSDPSTMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHT 443
N + E ++ S R A +E H LE E K + + Q AS
Sbjct: 130 NEISEASRKGLSRDLEASRTAKKELE-----AKHQKLEAE--NKKLTEGNQVSEASRKGL 182
Query: 444 GCIFAHAFQKFGASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSI 503
+ A+ +KL+ D+ AL +H K + D ++E +R ++ D +
Sbjct: 183 SNDLEASRAAKKELEAKYQKLETDHQALEAKHQKLE--ADYQVSETSRKGLSRDLEASRE 240
Query: 504 ANELVEARFELVCRSERITDLEEEIKETKQEAQLALAAKDDA--------LALKDQELTS 555
AN+ V + EL +++ LEE K +++E + L AK DA LA + +EL
Sbjct: 241 ANKKVTS--ELTQAKAQLSALEESKKLSEKE-KAELQAKLDAQGKALKEQLAKQTEELAK 297
Query: 556 LRAE 559
LRAE
Sbjct: 298 LRAE 301
>KTN1_VULVU (O97961) Kinectin
Length = 1330
Score = 41.6 bits (96), Expect = 0.006
Identities = 54/233 (23%), Positives = 106/233 (45%), Gaps = 32/233 (13%)
Query: 362 KESRTALREKEGHFDLAEKEARPNLGEKEKE--VPSDPSTMLRDATDHVEFLLS-RINHD 418
++ + ++ K+ L E++ + + K +E + +D + L+ ++ L+S + N D
Sbjct: 669 EKMQKSIHVKDDQIRLLEEQLQCEISNKMEEFKILNDQNKALQLEVQKLQILVSEQPNKD 728
Query: 419 YLEKEVFGKGIDDAGQECVASLFHTGCI-FAHAFQKFGASTAEGEKLQADYTALRTEH-- 475
+E+ D + V L TG I A ++ A E L + L+ +
Sbjct: 729 VVEQMEKCIQEKDEKLKTVEELLETGLIQVATKEEELNAIRTENSSLTKEVQDLKAKQND 788
Query: 476 ----AKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSER-ITDLEEEIKE 530
A + L KV+ E D K+ S+ EL+EA V E+ I DL++EI+
Sbjct: 789 QVSFASLVEELKKVIHEK-------DGKIKSV-EELLEAEVLKVANKEKTIQDLKQEIEA 840
Query: 531 TKQEA---------QLALAAK----DDALALKDQELTSLRAELEEKKRDLLKK 570
K+E QL++ ++ + L K++++ +++ LEEK++DL +
Sbjct: 841 LKEEIGNIQLEKAQQLSITSQIQELQNLLKGKEEQMNTMKTVLEEKEKDLASR 893
>LIPA_CAEEL (Q21049) Liprin alpha (LAR-interacting protein alpha)
(Synapse defective protein 2)
Length = 1139
Score = 41.2 bits (95), Expect = 0.007
Identities = 29/109 (26%), Positives = 54/109 (48%), Gaps = 14/109 (12%)
Query: 469 TALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSERITDLEEE- 527
T + T +A+ +D+L TE AR+K + +V + N++ E + + RIT E
Sbjct: 264 TEITTRNAELEDQL----TEDAREKHAAQESIVRLKNQICELDAQRTDQETRITTFESRF 319
Query: 528 ---------IKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDL 567
I++ + + LA KD A+ L ++++ SL+ LE ++ L
Sbjct: 320 LTAQRESTCIRDLNDKLEHQLANKDAAVRLNEEKVHSLQERLELAEKQL 368
>KINH_SYNRA (O43093) Kinesin heavy chain (Synkin)
Length = 935
Score = 40.8 bits (94), Expect = 0.010
Identities = 53/245 (21%), Positives = 101/245 (40%), Gaps = 45/245 (18%)
Query: 350 ENSNPVRQGTGEKESRTALR-EKEGHFDLAEKEARPNLGEKEKEVPSDPSTMLRDATDHV 408
+++ + + +K++ A + E EG EKE L +K + + + M D++
Sbjct: 603 QHTKTISDLSADKDAMEAKKIELEGRLGALEKEYEELL---DKTIAEEEANMQNADVDNL 659
Query: 409 EFLLSRINHDYLEK-EVFGKGIDDAGQECVASLFHTGCIFAHAFQKFGASTAEGEKLQAD 467
L +++ Y EK EV K IDD +E + EKL A
Sbjct: 660 SALKTKLEAQYAEKKEVQQKEIDDLKRE------------------LDRKQSGHEKLSAA 701
Query: 468 YTALRTEHAKCDDRLDKVLTEAARDKIT-------IDKKLVSIANELVEARF-------E 513
T LR + + L + +A +D I++ S+A +L + +
Sbjct: 702 MTDLRAANDQLQAALSEQPFQAPQDNSDMTEKEKDIERTRKSMAQQLADFEVMKKALMRD 761
Query: 514 LVCRSERITDLEEEIKETKQEAQLALAAKDDALALKD--------QELTSLRAELEEKKR 565
L R E++ +LE + ET+++ L A ++ K ++LT+++ +L E+
Sbjct: 762 LQNRCEKVVELEMSLDETREQYNNVLRASNNKAQQKKMAFLERNLEQLTNVQKQLVEQNA 821
Query: 566 DLLKK 570
L K+
Sbjct: 822 SLKKE 826
>MYHB_HUMAN (P35749) Myosin heavy chain, smooth muscle isoform (SMMHC)
Length = 1972
Score = 40.0 bits (92), Expect = 0.016
Identities = 44/216 (20%), Positives = 88/216 (40%), Gaps = 6/216 (2%)
Query: 355 VRQGTGEKESRTALREKEGHFDLAEKEARPNLGEKEKEVPSDPSTMLRDATDHVEFLLSR 414
VR ++E L E E L E+E R + E++ + L + + E +
Sbjct: 911 VRLAAKKQELEEILHEMEAR--LEEEEDRGQQLQAERKKMAQQMLDLEEQLEEEEAARQK 968
Query: 415 INHDYLEKEVFGKGIDD---AGQECVASLFHTGCIFAHAFQKFGASTAEGEKLQADYTAL 471
+ + + E K ++D + L + + AE E+ + T L
Sbjct: 969 LQLEKVTAEAKIKKLEDEILVMDDQNNKLSKERKLLEERISDLTTNLAEEEEKAKNLTKL 1028
Query: 472 RTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSERITDLEEEIKET 531
+ +H L+ L + + + ++K + + + ++ +I +L+ ++ +
Sbjct: 1029 KNKHESMISELEVRLKKEEKSRQELEKLKRKLEGDASDFHEQIADLQAQIAELKMQLAKK 1088
Query: 532 KQEAQLALAAKDDALALKDQELTSLRAELEEKKRDL 567
++E Q ALA DD +A K+ L +R ELE DL
Sbjct: 1089 EEELQAALARLDDEIAQKNNALKKIR-ELEGHISDL 1123
Score = 35.8 bits (81), Expect = 0.31
Identities = 53/228 (23%), Positives = 88/228 (38%), Gaps = 47/228 (20%)
Query: 345 DNEVCENSNPVRQGTGEKESRTALREKEGHF-----DLAEKEARPNLGEKEKEVPSDPST 399
D+E+ + +N +++ +RE EGH DL + A N EK+K
Sbjct: 1100 DDEIAQKNNALKK----------IRELEGHISDLQEDLDSERAARNKAEKQK-------- 1141
Query: 400 MLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQKFGASTA 459
RD + +E L + + D L+ QE A + A +
Sbjct: 1142 --RDLGEELEALKTELE-DTLDSTA-------TQQELRAKREQEVTVLKKALDE------ 1185
Query: 460 EGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSE 519
E +A +R +HA+ + L + L + R K +DK ++ E + EL +
Sbjct: 1186 ETRSHEAQVQEMRQKHAQAVEELTEQLEQFKRAKANLDKNKQTLEKENADLAGELRVLGQ 1245
Query: 520 RITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDL 567
++E K+ K EAQ+ L K + RAEL +K L
Sbjct: 1246 AKQEVEH--KKKKLEAQV------QELQSKCSDGERARAELNDKVHKL 1285
Score = 31.2 bits (69), Expect = 7.6
Identities = 29/111 (26%), Positives = 52/111 (46%), Gaps = 19/111 (17%)
Query: 461 GEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSER 520
G++LQA+ + + +++L++ EAAR K+ ++K V +
Sbjct: 938 GQQLQAERKKMAQQMLDLEEQLEE--EEAARQKLQLEK----------------VTAEAK 979
Query: 521 ITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDLLKKK 571
I LE+EI + L+ + L + +LT+ AE EEK ++L K K
Sbjct: 980 IKKLEDEIL-VMDDQNNKLSKERKLLEERISDLTTNLAEEEEKAKNLTKLK 1029
>IFA1_CAEEL (P90901) Intermediate filament protein ifa-1
(Intermediate filament protein A1) (IF-A1) (Cel IF A1)
Length = 575
Score = 40.0 bits (92), Expect = 0.016
Identities = 32/107 (29%), Positives = 55/107 (50%), Gaps = 5/107 (4%)
Query: 464 LQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSERITD 523
++ ++ E A+ +L+ +D+ ID LV+++N +EA L+ R RI
Sbjct: 140 MEGQLKKMQDELAEMRRKLEDATKGREQDRAKIDALLVTLSN--LEAEISLLKR--RIAQ 195
Query: 524 LEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDLLKK 570
LE+E+K KQE Q L+ A DQE T R + + + + LL++
Sbjct: 196 LEDEVKRIKQENQRLLSELQRARTDLDQE-TLNRIDYQNQVQTLLEE 241
>ERC1_MOUSE (Q99MI1) ERC protein 1 (Rab6-interacting protein 2)
(Rab6IP2) (CAZ-associated structural protein 2) (CAST2)
Length = 1120
Score = 40.0 bits (92), Expect = 0.016
Identities = 64/229 (27%), Positives = 100/229 (42%), Gaps = 30/229 (13%)
Query: 347 EVCENSNPVRQGTGEKESRTALREKEGHFDLAEKEARPNLGEKEKEVPSDPSTMLRDATD 406
EV N + E ES T+ + K+ + +A NL KE+ + ML +A
Sbjct: 767 EVENEKNDKDKKIAELESLTSRQVKDQNKKVA------NLKHKEQVEKKKSAQMLEEARR 820
Query: 407 HVEFLL--SRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQKFGASTAEGEKL 464
+ L S+ D L K+ DD +E +L + I TAE E +
Sbjct: 821 REDSLSDSSQQLQDSLRKK------DDRIEELEEALRESVQI-----------TAEREMV 863
Query: 465 QADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELV-CRSERITD 523
A + RT K + L + + ++ ++ KL S L E L R+ER
Sbjct: 864 LAQEESARTNAEKQVEELLMAMEKVKQELESMKAKLSSTQQSLAEKETHLTNLRAERRKH 923
Query: 524 LEEEIKETKQEAQL-ALAAKDDALALKDQELTSLRAELEEKKRDLLKKK 571
LEE + E KQEA L A++ KD +AL EL+S + + +E+ L ++K
Sbjct: 924 LEE-VLEMKQEALLAAISEKDANIAL--LELSSSKKKTQEEVAALKREK 969
Score = 35.0 bits (79), Expect = 0.53
Identities = 21/78 (26%), Positives = 39/78 (49%), Gaps = 3/78 (3%)
Query: 496 IDKKLVSIANELVEARFELVCRSERITDLEEEIKETKQEAQL---ALAAKDDALALKDQE 552
+ ++ + EL EL+ ++ L + ++KQ ++ +L AK+ A+ E
Sbjct: 464 LQSEIGQVKQELSRKDTELLALQTKLETLTNQFSDSKQHIEVLKESLTAKEQRAAILQTE 523
Query: 553 LTSLRAELEEKKRDLLKK 570
+ +LR LEEK+ L KK
Sbjct: 524 VDALRLRLEEKETMLNKK 541
Score = 32.3 bits (72), Expect = 3.4
Identities = 53/219 (24%), Positives = 99/219 (45%), Gaps = 28/219 (12%)
Query: 369 REKEGHFDLAEKEARPNLGEKEKEVPSDPS---TMLRDATDHVEFLLSRINHDYLEKEVF 425
REK+ D +K+ + +L EK + D S L D +H L S +
Sbjct: 641 REKQEEIDTYKKDLK-DLREKVSLLQGDLSEKEASLLDIKEHASSLASSGLKKDSRLKTL 699
Query: 426 GKGIDDAGQECVA-----SLFHTGCIFAHAFQKFGASTAEGEKLQADYTALRTEHAKCD- 479
++ +EC+ H + A A + ++L+ + + + E +K
Sbjct: 700 EIALEQKKEECLKMESQLKKAHEATLEARASPEMSDRI---QQLEREISRYKDESSKAQT 756
Query: 480 --DRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSERITDLE-EEIKETKQEAQ 536
DRL ++L E +K DKK+ + E + +R ++ +++++ +L+ +E E K+ AQ
Sbjct: 757 EVDRLLEILKEVENEKNDKDKKIAEL--ESLTSR-QVKDQNKKVANLKHKEQVEKKKSAQ 813
Query: 537 LALAA--KDDALALKDQEL-TSLR------AELEEKKRD 566
+ A ++D+L+ Q+L SLR ELEE R+
Sbjct: 814 MLEEARRREDSLSDSSQQLQDSLRKKDDRIEELEEALRE 852
>SPOF_SCHPO (Q10411) Sporulation-specific protein 15
Length = 1957
Score = 39.7 bits (91), Expect = 0.021
Identities = 61/290 (21%), Positives = 123/290 (42%), Gaps = 20/290 (6%)
Query: 288 GIVDPPVDSLTVEDKTIIGFFAQLPILDCSKLAAARRLAKFPPNFNTENPKKK---RHQR 344
G +D ++L+ +DK + +QL S A +LA+ + +N K K + ++
Sbjct: 406 GSIDEYQNNLSSKDKMVKQVSSQLEEARSSLAHATGKLAEINSERDFQNKKIKDFEKIEQ 465
Query: 345 DNEVCENSNPVRQGTGEKESRTALREKEGHFDLAEKEARPNLGEKEKEVPSDPSTMLRDA 404
D C NS+ + E + ++AL +K+ D R + E++K S S++
Sbjct: 466 DLRACLNSS-----SNELKEKSALIDKK---DQELNNLREQIKEQKKVSESTQSSLQSLQ 517
Query: 405 TDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQKFGASTAEGEKL 464
D L + H+ E ++ + Q +++ H + + A+ A +L
Sbjct: 518 RD---ILNEKKKHEVYESQL--NELKGELQTEISNSEHLSSQLSTLAAEKEAAVATNNEL 572
Query: 465 QADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSERITDL 524
+L+T ++L K + + ++ S +L E+ EL + IT
Sbjct: 573 SESKNSLQTLCNAFQEKLAKSVMQLKENEQNFSSLDTSF-KKLNESHQELENNHQTITKQ 631
Query: 525 EEEIKETKQEAQLALA---AKDDALALKDQELTSLRAELEEKKRDLLKKK 571
++ Q+ QL A K+ L+ ++ +L + +LEE + L+KK+
Sbjct: 632 LKDTSSKLQQLQLERANFEQKESTLSDENNDLRTKLLKLEESNKSLIKKQ 681
Score = 34.7 bits (78), Expect = 0.69
Identities = 24/97 (24%), Positives = 45/97 (45%), Gaps = 10/97 (10%)
Query: 480 DRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSERITDLE----------EEIK 529
+ L+ +L + +R ++ +K+ SI + L + FEL E++ L+ E IK
Sbjct: 1442 NELEHMLDDTSRKNSSLMEKIESINSSLDDKSFELASAVEKLGALQKLHSESLSLMENIK 1501
Query: 530 ETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRD 566
QEA+ + + + D E+T+ + E K D
Sbjct: 1502 SQLQEAKEKIQVDESTIQELDHEITASKNNYEGKLND 1538
Score = 31.6 bits (70), Expect = 5.8
Identities = 34/174 (19%), Positives = 74/174 (41%), Gaps = 19/174 (10%)
Query: 393 VPSDPSTMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQ 452
+ S + L D T+ ++ Y+EK V K +D+ Q V
Sbjct: 1061 ISSQTNKSLEDKTNQLK---------YIEKNV-QKLLDEKDQRNVE--------LEELTS 1102
Query: 453 KFGASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARF 512
K+G E +++ + ALR + K D L + ++K ++L + NEL+ +
Sbjct: 1103 KYGKLGEENAQIKDELLALRKKSKKQHD-LCANFVDDLKEKSDALEQLTNEKNELIVSLE 1161
Query: 513 ELVCRSERITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRD 566
+ +E + + ++ + + +L+ D+ +++ +L + EL+ K+D
Sbjct: 1162 QSNSNNEALVEERSDLANRLSDMKKSLSDSDNVISVIRSDLVRVNDELDTLKKD 1215
>MYH3_RAT (P12847) Myosin heavy chain, fast skeletal muscle,
embryonic
Length = 1940
Score = 39.3 bits (90), Expect = 0.028
Identities = 35/111 (31%), Positives = 56/111 (49%), Gaps = 10/111 (9%)
Query: 458 TAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCR 517
+AE EK A ++ E K D L K +EA R ++ ++KLV++ E + + ++
Sbjct: 843 SAETEKEMA---TMKEEFQKTKDELAK--SEAKRKEL--EEKLVTLVQEKNDLQLQVQAE 895
Query: 518 SERITDLEEEIKET-KQEAQLALAAKDDALALKDQELTSLRAELEEKKRDL 567
SE + D EE + K + QL K+ +D+E + AEL KKR L
Sbjct: 896 SENLLDAEERCDQLIKAKFQLEAKIKEVTERAEDEE--EINAELTAKKRKL 944
>MYH3_HUMAN (P11055) Myosin heavy chain, fast skeletal muscle,
embryonic (Muscle embryonic myosin heavy chain) (SMHCE)
Length = 1940
Score = 39.3 bits (90), Expect = 0.028
Identities = 35/111 (31%), Positives = 56/111 (49%), Gaps = 10/111 (9%)
Query: 458 TAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCR 517
+AE EK A ++ E K D L K +EA R ++ ++KLV++ E + + ++
Sbjct: 843 SAETEKEMA---TMKEEFQKTKDELAK--SEAKRKEL--EEKLVTLVQEKNDLQLQVQAE 895
Query: 518 SERITDLEEEIKET-KQEAQLALAAKDDALALKDQELTSLRAELEEKKRDL 567
SE + D EE + K + QL K+ +D+E + AEL KKR L
Sbjct: 896 SENLLDAEERCDQLIKAKFQLEAKIKEVTERAEDEE--EINAELTAKKRKL 944
>STKA_MOUSE (O55098) Serine/threonine-protein kinase 10 (EC
2.7.1.37) (Lymphocyte-oriented kinase)
Length = 966
Score = 38.9 bits (89), Expect = 0.037
Identities = 32/115 (27%), Positives = 61/115 (52%), Gaps = 12/115 (10%)
Query: 462 EKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSERI 521
EK++ D++ R E AK ++ E RD ++L + E V++ E + R +R
Sbjct: 624 EKMEQDHSVRRKEEAK------RIRLEQDRDYAKFQEQLKQMKKE-VKSEVEKLPRQQRK 676
Query: 522 TDLEEEIKETKQEAQLA----LAAKDDALALKDQELTSL-RAELEEKKRDLLKKK 571
++++++E Q+ Q +A + + L L ++LT+ R E+ +K+RD L KK
Sbjct: 677 ESMKQKMEEHSQKKQRLDRDFVAKQKEDLELAMRKLTTENRREICDKERDCLSKK 731
>HMMR_MOUSE (Q00547) Hyaluronan mediated motility receptor
(Intracellular hyaluronic acid binding protein)
(Receptor for hyaluronan-mediated motility)
Length = 794
Score = 38.9 bits (89), Expect = 0.037
Identities = 24/82 (29%), Positives = 47/82 (57%), Gaps = 7/82 (8%)
Query: 484 KVLTEAARDKITIDKKLVSIANELVEARFELVCRSERITDLEEEIK---ETKQEAQLALA 540
K LTE+ + ++ KLVSI E ++ + C +E++ + +EI + ++ ++ +A
Sbjct: 201 KDLTESKGKIVQLEGKLVSIEKEKIDEK----CETEKLLEYIQEISCASDQVEKCKVDIA 256
Query: 541 AKDDALALKDQELTSLRAELEE 562
++ L KD+E+ SL+ LEE
Sbjct: 257 QLEEDLKEKDREILSLKQSLEE 278
Score = 30.8 bits (68), Expect = 9.9
Identities = 24/113 (21%), Positives = 51/113 (44%), Gaps = 6/113 (5%)
Query: 465 QADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSERITDL 524
Q + AL E K ++ + + + ++ KL + +L E++ ++V ++ +
Sbjct: 161 QKNMRALSLELMKLRNKRETKMRSMMVKQEGMELKLQATQKDLTESKGKIVQLEGKLVSI 220
Query: 525 EEEIKETKQEAQLALAAKDDALALKDQ------ELTSLRAELEEKKRDLLKKK 571
E+E + K E + L + DQ ++ L +L+EK R++L K
Sbjct: 221 EKEKIDEKCETEKLLEYIQEISCASDQVEKCKVDIAQLEEDLKEKDREILSLK 273
>ERC1_HUMAN (Q8IUD2) ERC protein 1 (ELKS protein)
Length = 1116
Score = 38.9 bits (89), Expect = 0.037
Identities = 38/116 (32%), Positives = 59/116 (50%), Gaps = 5/116 (4%)
Query: 458 TAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELV-C 516
TAE E + A + RT K + L + + ++ ++ KL S L E L
Sbjct: 853 TAEREMVLAQEESARTNAEKQVEELLMAMEKVKQELESMKAKLSSTQQSLAEKETHLTNL 912
Query: 517 RSERITDLEEEIKETKQEAQL-ALAAKDDALALKDQELTSLRAELEEKKRDLLKKK 571
R+ER LEE + E KQEA L A++ KD +AL EL+S + + +E+ L ++K
Sbjct: 913 RAERRKHLEE-VLEMKQEALLAAISEKDANIAL--LELSSSKKKTQEEVAALKREK 965
Score = 34.7 bits (78), Expect = 0.69
Identities = 21/78 (26%), Positives = 39/78 (49%), Gaps = 3/78 (3%)
Query: 496 IDKKLVSIANELVEARFELVCRSERITDLEEEIKETKQEAQL---ALAAKDDALALKDQE 552
+ ++ + EL EL+ ++ L + ++KQ ++ +L AK+ A+ E
Sbjct: 464 LQAEIGQVKQELSRKDTELLALQTKLETLTNQFSDSKQHIEVLKESLTAKEQRAAILQTE 523
Query: 553 LTSLRAELEEKKRDLLKK 570
+ +LR LEEK+ L KK
Sbjct: 524 VDALRLRLEEKETMLNKK 541
>CTRO_MOUSE (P49025) Citron protein (Rho-interacting,
serine/threonine kinase 21)
Length = 1597
Score = 38.9 bits (89), Expect = 0.037
Identities = 45/215 (20%), Positives = 93/215 (42%), Gaps = 26/215 (12%)
Query: 368 LREKEGHF-------------DLAEKEARPNLGEKEKEVPSDPSTMLRDATDHVEFLLSR 414
L++KE H+ DLA+KE+ N+ ++ +E + +L + + + S+
Sbjct: 297 LKQKEQHYEEKIKVLDNQIKKDLADKESLENMMQRHEEEAHEKGKILSEQKAMINAMDSK 356
Query: 415 INHDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQKFGASTAEGEKLQADYTALRTE 474
I LE+ + + +A + S T + + L+ L +
Sbjct: 357 IRS--LEQRIV--ELSEANKLAANSSLFTQRNMKAQEEMISELRQQKFYLETQAGKLEAQ 412
Query: 475 HAKCDDRLDKVLTEAARDK---ITIDKKLVSIANELVEARFELVCRSERITDLEEEIKET 531
+ K +++L+K+ + DK + ++ +L ++ E E + EL ++T+L+ ++E
Sbjct: 413 NRKLEEQLEKISHQDHSDKSRLLELETRLREVSLEHEEQKLEL---KRQLTELQLSLQER 469
Query: 532 KQE---AQLALAAKDDALALKDQELTSLRAELEEK 563
+ + Q A AA + L EL AE EE+
Sbjct: 470 ESQLTALQAARAALESQLRQAKTELEETTAEAEEE 504
>BCD2_MOUSE (Q921C5) Cytoskeleton-like bicaudal D protein homolog 2
Length = 820
Score = 38.5 bits (88), Expect = 0.048
Identities = 33/127 (25%), Positives = 62/127 (47%), Gaps = 10/127 (7%)
Query: 454 FGASTAEGEKLQADYTALRTEHAKC----DDRLDKVLTEAARDKITIDKKLVSIANELVE 509
+ A +E E+L+ + T H K + R + ++ E+A + +K++ + EL +
Sbjct: 68 YEAIRSEMEQLKEAFGQAHTNHKKVAADGESREESLIQESASKEQYYVRKVLELQTELKQ 127
Query: 510 ARFELV---CRSERITDLEEEIKETKQEAQLALA-AKDDALALKDQELTSLR--AELEEK 563
R L +ER+T + +E+KE Q ++ +DD K +E L+ +ELEE+
Sbjct: 128 LRNVLTNTQSENERLTSVAQELKEINQNVEIQRGRLRDDIKEYKFREARLLQDYSELEEE 187
Query: 564 KRDLLKK 570
L K+
Sbjct: 188 NISLQKQ 194
>TPM2_SCHMA (P42638) Tropomyosin 2 (TMII)
Length = 284
Score = 38.1 bits (87), Expect = 0.062
Identities = 55/230 (23%), Positives = 90/230 (38%), Gaps = 40/230 (17%)
Query: 344 RDNEVCENSNPVRQGTGEKES-RTALREKEGHFDLAEKEARPNLGEKEKEVPSDPSTMLR 402
+D EV E ++Q +KE+ +T L E + +K A E E EV S +R
Sbjct: 39 KDEEVAEVLKKIQQVDTDKETAQTQLAETNTKLEETDKRAT----EAEAEVAS-LQKRIR 93
Query: 403 DATDHVEFLLSRINHDYLEKEVFGKGID--DAGQECVASLFHTGCIFAHAFQKFGASTAE 460
D +E +R+ ++ E K D D G++ + + + A+
Sbjct: 94 QLEDELESTETRLQEATVKLEEASKAADESDRGRKVLEN----------------RTFAD 137
Query: 461 GEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSER 520
E++ L+ +D D+ EAAR KL EL A L +
Sbjct: 138 EERINQLEEQLKESTFMAEDA-DRKYDEAAR-------KLAITEVELERAESRLEAAESK 189
Query: 521 ITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDLLKK 570
IT+LEEE++ + +L + +QE EE RDL ++
Sbjct: 190 ITELEEELRIVGNNVK--------SLEISEQEAAQREEAYEENIRDLTER 231
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.134 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 69,177,050
Number of Sequences: 164201
Number of extensions: 3083001
Number of successful extensions: 9568
Number of sequences better than 10.0: 222
Number of HSP's better than 10.0 without gapping: 24
Number of HSP's successfully gapped in prelim test: 204
Number of HSP's that attempted gapping in prelim test: 9099
Number of HSP's gapped (non-prelim): 685
length of query: 571
length of database: 59,974,054
effective HSP length: 116
effective length of query: 455
effective length of database: 40,926,738
effective search space: 18621665790
effective search space used: 18621665790
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0192.11


