

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0190.3
(329 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
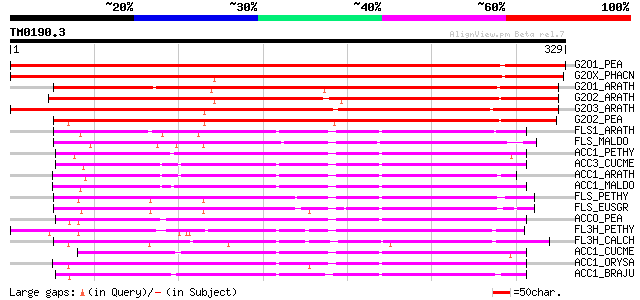
G2O1_PEA (Q9SQ80) Gibberellin 2-beta-dioxygenase 1 (EC 1.14.11.1... 528 e-150
G2OX_PHACN (Q9XG83) Gibberellin 2-beta-dioxygenase (EC 1.14.11.1... 417 e-116
G2O1_ARATH (Q8LEA2) Gibberellin 2-beta-dioxygenase 1 (EC 1.14.11... 373 e-103
G2O2_ARATH (Q9XFR9) Gibberellin 2-beta-dioxygenase 2 (EC 1.14.11... 370 e-102
G2O3_ARATH (O64692) Gibberellin 2-beta-dioxygenase 3 (EC 1.14.11... 342 1e-93
G2O2_PEA (Q9XHM5) Gibberellin 2-beta-dioxygenase 2 (EC 1.14.11.1... 305 1e-82
FLS1_ARATH (Q96330) Flavonol synthase 1 (EC 1.14.11.-) (FLS 1) 145 1e-34
FLS_MALDO (Q9XHG2) Flavonol synthase (EC 1.14.11.-) (FLS) 132 1e-30
ACC1_PETHY (Q08506) 1-aminocyclopropane-1-carboxylate oxidase 1 ... 132 1e-30
ACC3_CUCME (P54847) 1-aminocyclopropane-1-carboxylate oxidase 3 ... 131 2e-30
ACC1_ARATH (Q06588) 1-aminocyclopropane-1-carboxylate oxidase (E... 131 2e-30
ACC1_MALDO (Q00985) 1-aminocyclopropane-1-carboxylate oxidase 1 ... 131 3e-30
FLS_PETHY (Q07512) Flavonol synthase (EC 1.14.11.-) (FLS) 130 5e-30
FLS_EUSGR (Q9M547) Flavonol synthase (EC 1.14.11.-) (FLS) 130 5e-30
ACCO_PEA (P31239) 1-aminocyclopropane-1-carboxylate oxidase (EC ... 129 8e-30
FL3H_PETHY (Q07353) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 129 1e-29
FL3H_CALCH (Q05963) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 129 1e-29
ACC1_CUCME (Q04644) 1-aminocyclopropane-1-carboxylate oxidase 1 ... 128 2e-29
ACC1_ORYSA (Q40634) 1-aminocyclopropane-1-carboxylate oxidase 1 ... 128 2e-29
ACC1_BRAJU (Q09052) 1-aminocyclopropane-1-carboxylate oxidase (E... 127 3e-29
>G2O1_PEA (Q9SQ80) Gibberellin 2-beta-dioxygenase 1 (EC 1.14.11.13)
(Gibberellin 2-beta-hydroxylase 1) (Gibberellin
2-oxidase 1) (GA 2-oxidase 1) (SLENDER protein)
Length = 327
Score = 528 bits (1360), Expect = e-150
Identities = 256/329 (77%), Positives = 294/329 (88%), Gaps = 2/329 (0%)
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MVLLSK +EQY+YV+N M S +IP+VDLSKPDAK++IVKACE+FGFFKVINHG+ +
Sbjct: 1 MVLLSKPTSEQYTYVRNNMPITFSSSIPLVDLSKPDAKTLIVKACEDFGFFKVINHGIPL 60
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP 120
+AISQLESEA FFS+P TEKEK G ANPFGYGNKRIG NGD+GW+EYLLL TNQ+ N
Sbjct: 61 DAISQLESEAFKFFSLPQTEKEKAGPANPFGYGNKRIGLNGDIGWIEYLLLTTNQDHNFS 120
Query: 121 VHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNH 180
++ ++ +KFR +L DY CA+RNM CEILDLMAEGLKIQPKNV SKL+MDKQSD +FRVNH
Sbjct: 121 LYGEDIDKFRGLLKDYKCAMRNMACEILDLMAEGLKIQPKNVFSKLVMDKQSDCLFRVNH 180
Query: 181 YPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINV 240
YPACPE+ ++GENLIGFGEHTDPQIIS+LRSNNTSG QISLRDGSWISVPPD+SSFFINV
Sbjct: 181 YPACPELAINGENLIGFGEHTDPQIISILRSNNTSGFQISLRDGSWISVPPDHSSFFINV 240
Query: 241 GDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKE 300
GDSLQVMTNGRF+SV+HRVLANG RLSMIYF GPPLSEKIAPLPSL+KG +ESLYKE
Sbjct: 241 GDSLQVMTNGRFKSVRHRVLANGIDPRLSMIYFCGPPLSEKIAPLPSLMKG--KESLYKE 298
Query: 301 FTWFEYKNSSYGSRLADNRLGHFERIAAS 329
FTWFEYK+S+YGSRLADNRLG++ERIAA+
Sbjct: 299 FTWFEYKSSTYGSRLADNRLGNYERIAAT 327
>G2OX_PHACN (Q9XG83) Gibberellin 2-beta-dioxygenase (EC 1.14.11.13)
(Gibberellin 2-beta-hydroxylase) (Gibberellin 2-oxidase)
(GA 2-oxidase)
Length = 332
Score = 417 bits (1071), Expect = e-116
Identities = 207/332 (62%), Positives = 260/332 (77%), Gaps = 5/332 (1%)
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MV+LS+ A Q+ +K +TP IPVVDL+ PDAK++IV AC +FGFFK++NHGV +
Sbjct: 1 MVVLSQPALNQFFLLKPFKSTPLFTGIPVVDLTHPDAKNLIVNACRDFGFFKLVNHGVPL 60
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP 120
E ++ LE+EAL FF +EK++ G +PFGYG+KRIG NGDVGWVEYLLL TN +V P
Sbjct: 61 ELMANLENEALRFFKKSQSEKDRAGPPDPFGYGSKRIGPNGDVGWVEYLLLNTNPDVISP 120
Query: 121 ----VHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVF 176
+ +NP FR V+++Y+ AV+NM +L+LMAEGL I+ +N LS+LL D++SDS F
Sbjct: 121 KSLCIFRENPHHFRAVVENYITAVKNMCYAVLELMAEGLGIRQRNTLSRLLKDEKSDSCF 180
Query: 177 RVNHYPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSF 236
R+NHYP CPE+ NL+GFGEHTDPQIIS+LRSN+TSGLQI L DG+W+SVPPD +SF
Sbjct: 181 RLNHYPPCPEVQALNRNLVGFGEHTDPQIISVLRSNSTSGLQICLTDGTWVSVPPDQTSF 240
Query: 237 FINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEES 296
FINVGD+LQVMTNGRF+SVKHRVLA+ +SRLSMIYFGGP LSE IAPLPS++ G EE
Sbjct: 241 FINVGDALQVMTNGRFKSVKHRVLADTTKSRLSMIYFGGPALSENIAPLPSVMLKG-EEC 299
Query: 297 LYKEFTWFEYKNSSYGSRLADNRLGHFERIAA 328
LYKEFTW EYK ++Y SRLADNRL F++ AA
Sbjct: 300 LYKEFTWCEYKKAAYTSRLADNRLAPFQKSAA 331
>G2O1_ARATH (Q8LEA2) Gibberellin 2-beta-dioxygenase 1 (EC
1.14.11.13) (Gibberellin 2-beta-hydroxylase 1)
(Gibberellin 2-oxidase 1) (GA 2-oxidase 1)
Length = 329
Score = 373 bits (957), Expect = e-103
Identities = 184/306 (60%), Positives = 236/306 (76%), Gaps = 9/306 (2%)
Query: 27 IPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTGL 86
IPV+D+S P++K +VKACE+FGFFKVINHGVS E +S LE E ++FFS+P +EK +
Sbjct: 18 IPVIDMSDPESKHALVKACEDFGFFKVINHGVSAELVSVLEHETVDFFSLPKSEKTQVA- 76
Query: 87 ANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVN----VPVHSQNPEKFRCVLDDYLCAVRN 142
PFGYGN +IG NGDVGWVEYLL+ N + P ++P FR L++Y +VR
Sbjct: 77 GYPFGYGNSKIGRNGDVGWVEYLLMNANHDSGSGPLFPSLLKSPGTFRNALEEYTTSVRK 136
Query: 143 MGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACP---EMGLDGENLIGFGE 199
M ++L+ + +GL I+P+N LSKL+ D+ +DS+ R+NHYP CP + G+N+IGFGE
Sbjct: 137 MTFDVLEKITDGLGIKPRNTLSKLVSDQNTDSILRLNHYPPCPLSNKKTNGGKNVIGFGE 196
Query: 200 HTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRV 259
HTDPQIIS+LRSNNTSGLQI+L DGSWISVPPD++SFF NVGDSLQVMTNGRF+SV+HRV
Sbjct: 197 HTDPQIISVLRSNNTSGLQINLNDGSWISVPPDHTSFFFNVGDSLQVMTNGRFKSVRHRV 256
Query: 260 LANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGSRLADNR 319
LAN +SR+SMIYF GP L+++IAPL L+ ++E LY+EFTW EYKNS+Y SRL+DNR
Sbjct: 257 LANCKKSRVSMIYFAGPSLTQRIAPLTCLI-DNEDERLYEEFTWSEYKNSTYNSRLSDNR 315
Query: 320 LGHFER 325
L FER
Sbjct: 316 LQQFER 321
>G2O2_ARATH (Q9XFR9) Gibberellin 2-beta-dioxygenase 2 (EC
1.14.11.13) (Gibberellin 2-beta-hydroxylase 2)
(Gibberellin 2-oxidase 2) (GA 2-oxidase 2)
Length = 341
Score = 370 bits (950), Expect = e-102
Identities = 181/308 (58%), Positives = 244/308 (78%), Gaps = 10/308 (3%)
Query: 24 SFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEK 83
S +IPVV+L+ P+AK+ IVKACEEFGFFKV+NHGV E +++LE EA+ FF +P + K +
Sbjct: 28 SHSIPVVNLADPEAKTRIVKACEEFGFFKVVNHGVRPELMTRLEQEAIGFFGLPQSLKNR 87
Query: 84 TGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP----VHSQNPEKFRCVLDDYLCA 139
G P+GYGNKRIG NGDVGW+EYLLL N +++ P V Q P+ FR +++Y+
Sbjct: 88 AGPPEPYGYGNKRIGPNGDVGWIEYLLLNANPQLSSPKTSAVFRQTPQIFRESVEEYMKE 147
Query: 140 VRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLI--GF 197
++ + ++L+++AE L I+P++ LSK+L D++SDS R+NHYPA E + E ++ GF
Sbjct: 148 IKEVSYKVLEMVAEELGIEPRDTLSKMLRDEKSDSCLRLNHYPAAEE---EAEKMVKVGF 204
Query: 198 GEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKH 257
GEHTDPQIIS+LRSNNT+GLQI ++DGSW++VPPD+SSFFINVGD+LQVMTNGRF+SVKH
Sbjct: 205 GEHTDPQIISVLRSNNTAGLQICVKDGSWVAVPPDHSSFFINVGDALQVMTNGRFKSVKH 264
Query: 258 RVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGSRLAD 317
RVLA+ +SR+SMIYFGGPPLS+KIAPLP L+ +++ LYKEFTW +YK+S+Y S+L D
Sbjct: 265 RVLADTRRSRISMIYFGGPPLSQKIAPLPCLVP-EQDDWLYKEFTWSQYKSSAYKSKLGD 323
Query: 318 NRLGHFER 325
RLG FE+
Sbjct: 324 YRLGLFEK 331
>G2O3_ARATH (O64692) Gibberellin 2-beta-dioxygenase 3 (EC
1.14.11.13) (Gibberellin 2-beta-hydroxylase 3)
(Gibberellin 2-oxidase 3) (GA 2-oxidase 3)
Length = 335
Score = 342 bits (876), Expect = 1e-93
Identities = 175/329 (53%), Positives = 233/329 (70%), Gaps = 7/329 (2%)
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MV++ + A+ + N P IPV+DL+ DAK+ IVKACEEFGFFKVINHGV
Sbjct: 1 MVIVLQPASFDSNLYVNPKCKPRPVLIPVIDLTDSDAKTQIVKACEEFGFFKVINHGVRP 60
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTN----QE 116
+ ++QLE EA+NFF++ + K+K G +PFGYG KRIG NGD+GW+EY+LL N
Sbjct: 61 DLLTQLEQEAINFFALHHSLKDKAGPPDPFGYGTKRIGPNGDLGWLEYILLNANLCLESH 120
Query: 117 VNVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVF 176
+ P FR +++Y+ ++ M + L+++ E LKI+PK LS+L+ K+SDS
Sbjct: 121 KTTAIFRHTPAIFREAVEEYIKEMKRMSSKFLEMVEEELKIEPKEKLSRLVKVKESDSCL 180
Query: 177 RVNHYPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSF 236
R+NHYP E + E IGFGEHTDPQ+ISLLRSN+T GLQI ++DG+W+ V PD+SSF
Sbjct: 181 RMNHYPEKEETPVKEE--IGFGEHTDPQLISLLRSNDTEGLQICVKDGTWVDVTPDHSSF 238
Query: 237 FINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEES 296
F+ VGD+LQVMTNGRF+SVKHRV+ N +SR+SMIYF GPPLSEKIAPL S L +++
Sbjct: 239 FVLVGDTLQVMTNGRFKSVKHRVVTNTKRSRISMIYFAGPPLSEKIAPL-SCLVPKQDDC 297
Query: 297 LYKEFTWFEYKNSSYGSRLADNRLGHFER 325
LY EFTW +YK S+Y ++L D RLG FE+
Sbjct: 298 LYNEFTWSQYKLSAYKTKLGDYRLGLFEK 326
>G2O2_PEA (Q9XHM5) Gibberellin 2-beta-dioxygenase 2 (EC 1.14.11.13)
(Gibberellin 2-beta-hydroxylase 2) (Gibberellin
2-oxidase 2) (GA 2-oxidase 2)
Length = 345
Score = 305 bits (781), Expect = 1e-82
Identities = 154/314 (49%), Positives = 209/314 (66%), Gaps = 17/314 (5%)
Query: 27 IPVVDLS--KPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKT 84
IP +DLS + ++VKACEE+GFFKV+NH V E IS+L+ E + FFS +EK +
Sbjct: 20 IPTIDLSLERSQLSELVVKACEEYGFFKVVNHSVPKEVISRLDEEGIEFFSKNSSEKRQA 79
Query: 85 GLANPFGYGNKRIGHNGDVGWVEYLLLKTN----QEVNVPVHSQNPEKFRCVLDDYLCAV 140
G + PFGYG K IG NGD G +EYLLL +N E + + +P KF C+++DY+ AV
Sbjct: 80 GTSTPFGYGCKNIGPNGDKGELEYLLLHSNPISISERSKTIAKDHPIKFSCIVNDYIKAV 139
Query: 141 RNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDG--------- 191
+++ CEIL+L AEGL + K+ LSK++ D+ SDS+ R+NHYP ++G D
Sbjct: 140 KDLTCEILELAAEGLWVPDKSSLSKIIKDEHSDSLLRINHYPPVKKLGNDNWDPSKIQNS 199
Query: 192 -ENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNG 250
N IGFGEH+DPQI+++LRSNN GLQIS G WI VPPD S F++ VGD+LQV+TNG
Sbjct: 200 NNNNIGFGEHSDPQILTILRSNNVGGLQISTHHGLWIPVPPDPSEFYVMVGDALQVLTNG 259
Query: 251 RFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSS 310
RF SV+HRVL N + R+SM+YF PPL+ I+PL ++ LY+ FTW +YK ++
Sbjct: 260 RFVSVRHRVLTNTTKPRMSMMYFAAPPLNWLISPLSKMVT-AHSPCLYRPFTWAQYKQAA 318
Query: 311 YGSRLADNRLGHFE 324
Y RL D RL F+
Sbjct: 319 YALRLGDTRLDQFK 332
>FLS1_ARATH (Q96330) Flavonol synthase 1 (EC 1.14.11.-) (FLS 1)
Length = 336
Score = 145 bits (367), Expect = 1e-34
Identities = 94/292 (32%), Positives = 157/292 (53%), Gaps = 21/292 (7%)
Query: 27 IPVVDLSKPDAKSI---IVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEK 83
IPVVDLS PD +S+ +VKA EE+G F+V+NHG+ E I +L+ FF +P +EKE
Sbjct: 43 IPVVDLSDPDEESVRRAVVKASEEWGLFQVVNHGIPTELIRRLQDVGRKFFELPSSEKE- 101
Query: 84 TGLANP------FGYGNK-RIGHNGDVGWVEYLL--LKTNQEVNVPVHSQNPEKFRCVLD 134
+A P GYG K + G WV++L + VN +NP ++R V +
Sbjct: 102 -SVAKPEDSKDIEGYGTKLQKDPEGKKAWVDHLFHRIWPPSCVNYRFWPKNPPEYREVNE 160
Query: 135 DYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENL 194
+Y V+ + +L ++++GL ++ ++ L + L + ++ + ++N+YP CP L
Sbjct: 161 EYAVHVKKLSETLLGILSDGLGLK-RDALKEGLGGEMAEYMMKINYYPPCPRPDL----A 215
Query: 195 IGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRS 254
+G HTD I+LL N GLQ+ +D W S+ +++GD + ++NGR+++
Sbjct: 216 LGVPAHTDLSGITLLVPNEVPGLQV-FKDDHWFDAEYIPSAVIVHIGDQILRLSNGRYKN 274
Query: 255 VKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEY 306
V HR + ++R+S F PP + + PLP L G +K F + +Y
Sbjct: 275 VLHRTTVDKEKTRMSWPVFLEPPREKIVGPLPE-LTGDDNPPKFKPFAFKDY 325
>FLS_MALDO (Q9XHG2) Flavonol synthase (EC 1.14.11.-) (FLS)
Length = 337
Score = 132 bits (332), Expect = 1e-30
Identities = 86/300 (28%), Positives = 146/300 (48%), Gaps = 29/300 (9%)
Query: 27 IPVVDLSKPDAKSIIVKACE---EFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEK 83
+P++D S PD + +IV+ E +G ++++NH + E IS+L++ FF +P EKE
Sbjct: 41 VPIIDFSDPDEEKLIVQITEASSNWGMYQIVNHDIPSEVISKLQAVGKEFFELPQEEKEA 100
Query: 84 TGL----ANPFGYGNKRI-----GHNGDVGWVEYLLLKT--NQEVNVPVHSQNPEKFRCV 132
A+ GYG K G GWV+ L K VN +NP +R
Sbjct: 101 YAKPPDSASIEGYGTKLFKEISEGDTTKKGWVDNLFNKIWPPSVVNYQFWPKNPPSYREA 160
Query: 133 LDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGE 192
++Y + N+ ++ L++ GL ++ + L K + + ++N+YP CP L
Sbjct: 161 NEEYAKHLHNVVEKLFRLLSLGLGLEGQE-LKKAAGGDNLEYLLKINYYPPCPRPDL--- 216
Query: 193 NLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRF 252
+G HTD +++L N+ GLQ + +DG W V ++ I++GD +++M+NG++
Sbjct: 217 -ALGVVAHTDMSTVTILVPNDVQGLQ-ACKDGRWYDVKYIPNALVIHIGDQMEIMSNGKY 274
Query: 253 RSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYG 312
SV HR N ++R+S F PP + P P L+ + +YK YG
Sbjct: 275 TSVLHRTTVNKDKTRISWPVFLEPPADHVVGPHPQLVNAVNQP---------KYKTKKYG 325
>ACC1_PETHY (Q08506) 1-aminocyclopropane-1-carboxylate oxidase 1 (EC
1.14.17.4) (ACC oxidase 1) (Ethylene-forming enzyme)
(EFE)
Length = 319
Score = 132 bits (332), Expect = 1e-30
Identities = 87/288 (30%), Positives = 148/288 (51%), Gaps = 16/288 (5%)
Query: 28 PVVDLSKPDA------KSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK 81
P++ L K + +I ACE +GFF+++NHG+ E + +E + + ++
Sbjct: 5 PIISLDKVNGVERAATMEMIKDACENWGFFELVNHGIPREVMDTVEKMTKGHYKKCMEQR 64
Query: 82 EKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAVR 141
K +A+ G + D+ W LK N+ E++R V+ D+ +
Sbjct: 65 FKELVASKALEGVQ--AEVTDMDWESTFFLKHLPISNISEVPDLDEEYREVMRDFAKRLE 122
Query: 142 NMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLIGFGEHT 201
+ E+LDL+ E L ++ + + K + +V++YP CP+ L + G HT
Sbjct: 123 KLAEELLDLLCENLGLEKGYLKNAFYGSKGPNFGTKVSNYPPCPKPDL----IKGLRAHT 178
Query: 202 DPQ-IISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVL 260
D II L + + SGLQ+ L+DG WI VPP S +N+GD L+V+TNG+++SV HRV+
Sbjct: 179 DAGGIILLFQDDKVSGLQL-LKDGQWIDVPPMRHSIVVNLGDQLEVITNGKYKSVMHRVI 237
Query: 261 ANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEES--LYKEFTWFEY 306
A +R+S+ F P I P P+L++ EE+ +Y +F + +Y
Sbjct: 238 AQKDGARMSLASFYNPGSDAVIYPAPALVEKEAEENKQVYPKFVFDDY 285
>ACC3_CUCME (P54847) 1-aminocyclopropane-1-carboxylate oxidase 3 (EC
1.14.17.4) (ACC oxidase 3) (Ethylene-forming enzyme)
(EFE)
Length = 320
Score = 131 bits (330), Expect = 2e-30
Identities = 86/288 (29%), Positives = 156/288 (53%), Gaps = 17/288 (5%)
Query: 28 PVVDLSKPDAKSIIV------KACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK 81
PV++++ + +S + ACE +GFF+++NHG+S E + ++E + + ++
Sbjct: 6 PVINMNNLNGESRVSVLNQINDACENWGFFELVNHGISHELMDKVEKLTKEHYRKCMEQR 65
Query: 82 EKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAVR 141
K +A+ G + N D W L+ N+ E+++ V+ ++ +
Sbjct: 66 FKEMVASK-GLDSVETEIN-DTDWESTFFLRHLPVSNMSEIGDLDEEYKKVMKEFADELE 123
Query: 142 NMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSV-FRVNHYPACPEMGLDGENLIGFGEH 200
+ E+LDL+ E L ++ K L K+ + + +V++YP CP+ L + G H
Sbjct: 124 KLAEEVLDLLCENLGLE-KGYLKKVFYGSKGPNFGTKVSNYPPCPKPEL----IKGLRAH 178
Query: 201 TDPQ-IISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRV 259
TD +I L + + SGL + L+DG W+ VPP + S IN+GD L+V+TNG+++SV HRV
Sbjct: 179 TDAGGLILLFQDDKVSGLHV-LKDGKWVDVPPMHHSIVINLGDQLEVITNGKYKSVMHRV 237
Query: 260 LANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEES-LYKEFTWFEY 306
+A +R+S+ F P I P P+L++G +E++ LY +F + +Y
Sbjct: 238 IAQEDGNRMSIASFYNPGNDAVIYPAPALVEGEQEKTKLYPKFVFDDY 285
>ACC1_ARATH (Q06588) 1-aminocyclopropane-1-carboxylate oxidase (EC
1.14.17.4) (ACC oxidase) (Ethylene-forming enzyme) (EFE)
Length = 323
Score = 131 bits (330), Expect = 2e-30
Identities = 90/284 (31%), Positives = 147/284 (51%), Gaps = 19/284 (6%)
Query: 26 TIPVVDLSKPDAKSIIVK------ACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVT 79
+ P+++L K + + + ACE +GFF+ +NHG+S+E + ++E + +
Sbjct: 3 SFPIINLEKLNGEERAITMEKIKDACENWGFFECVNHGISLELLDKVEKMTKEHYKKCME 62
Query: 80 EKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCA 139
E+ K + N G + R N DV W LK N+ + +R ++ D+
Sbjct: 63 ERFKESIKNR-GLDSLRSEVN-DVDWESTFYLKHLPVSNISDVPDLDDDYRTLMKDFAGK 120
Query: 140 VRNMGCEILDLMAEGLKIQPKNVLSKLLM-DKQSDSVFRVNHYPACPEMGLDGENLIGFG 198
+ + E+LDL+ E L ++ K L K+ K+ +V++YP CP L + G
Sbjct: 121 IEKLSEELLDLLCENLGLE-KGYLKKVFYGSKRPTFGTKVSNYPPCPNPDL----VKGLR 175
Query: 199 EHTDPQ-IISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKH 257
HTD II L + + SGLQ+ L+DG W+ VPP S +N+GD L+V+TNG+++SV+H
Sbjct: 176 AHTDAGGIILLFQDDKVSGLQL-LKDGEWVDVPPVKHSIVVNLGDQLEVITNGKYKSVEH 234
Query: 258 RVLA-NGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKE 300
RVL+ + R+S+ F P I P P L+ GKE K+
Sbjct: 235 RVLSQTDGEGRMSIASFYNPGSDSVIFPAPELI--GKEAEKEKK 276
>ACC1_MALDO (Q00985) 1-aminocyclopropane-1-carboxylate oxidase 1 (EC
1.14.17.4) (ACC oxidase 1) (Ethylene-forming enzyme)
(EFE) (Protein AP4) (PAE12)
Length = 314
Score = 131 bits (329), Expect = 3e-30
Identities = 87/290 (30%), Positives = 154/290 (53%), Gaps = 17/290 (5%)
Query: 26 TIPVVDLSKPDAKSI------IVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVT 79
T PVVDLS + + I ACE +GFF+++NHG+S E + +E + + +
Sbjct: 3 TFPVVDLSLVNGEERAATLEKINDACENWGFFELVNHGMSTELLDTVEKMTKDHYKKTME 62
Query: 80 EKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCA 139
++ K +A G + + D+ W L+ N+ E++R + ++
Sbjct: 63 QRFKEMVAAK-GLDDVQ-SEIHDLDWESTFFLRHLPSSNISEIPDLEEEYRKTMKEFAVE 120
Query: 140 VRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSV-FRVNHYPACPEMGLDGENLIGFG 198
+ + ++LDL+ E L ++ K L K+ + + +V++YP CP+ L + G
Sbjct: 121 LEKLAEKLLDLLCENLGLE-KGYLKKVFYGSKGPNFGTKVSNYPPCPKPDL----IKGLR 175
Query: 199 EHTDPQ-IISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKH 257
H+D II L + + SGLQ+ L+DG W+ VPP + S IN+GD ++V+TNG+++SV H
Sbjct: 176 AHSDAGGIILLFQDDKVSGLQL-LKDGEWVDVPPMHHSIVINLGDQIEVITNGKYKSVMH 234
Query: 258 RVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEES-LYKEFTWFEY 306
RV+A +R+S+ F P I+P P++L+ E++ Y +F + +Y
Sbjct: 235 RVIAQSDGTRMSIASFYNPGNDSFISPAPAVLEKKTEDAPTYPKFVFDDY 284
>FLS_PETHY (Q07512) Flavonol synthase (EC 1.14.11.-) (FLS)
Length = 348
Score = 130 bits (327), Expect = 5e-30
Identities = 91/296 (30%), Positives = 150/296 (49%), Gaps = 21/296 (7%)
Query: 27 IPVVDLSKPDAKS---IIVKACEEFGFFKVINHGVSMEAISQLESEALNFFS-MPVTEKE 82
+PV+DL PD +I A +E+G F++INHG+ EAI+ L+ FF +P EKE
Sbjct: 55 VPVIDLRDPDENKMVKLIADASKEWGIFQLINHGIPDEAIADLQKVGKEFFEHVPQEEKE 114
Query: 83 ---KTGLANPF-GYGNKRIGH-NGDVGWVEYLLLKT--NQEVNVPVHSQNPEKFRCVLDD 135
KT +N GYG G GWV++L K VN +NP +R ++
Sbjct: 115 LIAKTPGSNDIEGYGTSLQKEVEGKKGWVDHLFHKIWPPSAVNYRYWPKNPPSYREANEE 174
Query: 136 YLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLI 195
Y +R + I ++ GL ++ ++ D + + ++N+YP CP L +
Sbjct: 175 YGKRMREVVDRIFKSLSLGLGLEGHEMIEAAGGD-EIVYLLKINYYPPCPRPDL----AL 229
Query: 196 GFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSV 255
G HTD I++L N GLQ+ +DG W V ++ +++GD +++++NG+++SV
Sbjct: 230 GVVAHTDMSYITILVPNEVQGLQV-FKDGHWYDVKYIPNALIVHIGDQVEILSNGKYKSV 288
Query: 256 KHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSY 311
HR N ++R+S F PP ++ P+P LL E+ +F +YK+ Y
Sbjct: 289 YHRTTVNKDKTRMSWPVFLEPPSEHEVGPIPKLL----SEANPPKFKTKKYKDYVY 340
>FLS_EUSGR (Q9M547) Flavonol synthase (EC 1.14.11.-) (FLS)
Length = 334
Score = 130 bits (327), Expect = 5e-30
Identities = 90/297 (30%), Positives = 154/297 (51%), Gaps = 25/297 (8%)
Query: 27 IPVVDLSKPDAKSII---VKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKE- 82
+PV+DLS D K I+ +A +E+G F+V+NHG+ E I +L+ +FF +P EKE
Sbjct: 43 VPVIDLSDSDEKKIVGLVSEASKEWGIFQVVNHGIPNEVIRKLQEVGKHFFELPQEEKEL 102
Query: 83 ---KTGLANPFGYGNKRIGH-NGDVGWVEYLLLKT--NQEVNVPVHSQNPEKFRCVLDDY 136
G + GYG + +G GWV++L K +N +NP +R ++Y
Sbjct: 103 IAKPEGSQSIEGYGTRLQKEVDGKKGWVDHLFHKIWPPSAINYQFWPKNPPAYREANEEY 162
Query: 137 LCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVF--RVNHYPACPEMGLDGENL 194
++ + + ++ GL ++P + D D V+ ++N+YP CP L
Sbjct: 163 AKRLQLVVDNLFKYLSLGLDLEPNSFKDGAGGD---DLVYLMKINYYPPCPRPDL----A 215
Query: 195 IGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRS 254
+G HTD I++L N GLQ+ +DG W ++ +++GD +++M+NG+++S
Sbjct: 216 LGVA-HTDMSAITVLVPNEVPGLQV-YKDGHWYDCKYIPNALIVHIGDQVEIMSNGKYKS 273
Query: 255 VKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSY 311
V HR N ++R+S F PP ++ P+P L+ EE+ K F +YK+ +Y
Sbjct: 274 VYHRTTVNKEKTRMSWPVFLEPPPDHEVGPIPKLV---NEENPAK-FKTKKYKDYAY 326
>ACCO_PEA (P31239) 1-aminocyclopropane-1-carboxylate oxidase (EC
1.14.17.4) (ACC oxidase) (Ethylene-forming enzyme) (EFE)
Length = 317
Score = 129 bits (325), Expect = 8e-30
Identities = 87/287 (30%), Positives = 147/287 (50%), Gaps = 16/287 (5%)
Query: 28 PVVDLSK---PDAKS---IIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK 81
P+VD+ K D KS +I ACE +GFF+ +NHG+S+E + +E + + ++
Sbjct: 5 PIVDMGKLNTEDRKSTMELIKDACENWGFFECVNHGISIEMMDTVEKLTKEHYKKCMEQR 64
Query: 82 EKTGLANPFGYGNKRIGHN-GDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAV 140
K +A G + + D+ W L+ ++ + +R V+ ++ +
Sbjct: 65 FKEMVATK---GLECVQSEIDDLDWESTFFLRHLPVSSISEIPDLDDDYRKVMKEFALKL 121
Query: 141 RNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLIGFGEH 200
+ E+LDL+ E L ++ + K + +V++YP CP+ L + G H
Sbjct: 122 EELAEELLDLLCENLGLEKGYLKKAFYGSKGPNFGTKVSNYPPCPKPEL----IKGLRAH 177
Query: 201 TDPQ-IISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRV 259
TD II L + + SGLQ+ L+D WI VPP S IN+GD L+V+TNG+++SV HRV
Sbjct: 178 TDAGGIILLFQDDKVSGLQL-LKDDQWIDVPPMRHSIVINLGDQLEVITNGKYKSVMHRV 236
Query: 260 LANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEY 306
+A +R+S+ F P I+P +LLK + +Y +F + +Y
Sbjct: 237 IAQTDGARMSIASFYNPGDDAVISPASTLLKENETSEVYPKFVFDDY 283
>FL3H_PETHY (Q07353) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
(Fragment)
Length = 369
Score = 129 bits (324), Expect = 1e-29
Identities = 99/331 (29%), Positives = 164/331 (48%), Gaps = 41/331 (12%)
Query: 1 MVLLSKQATEQYSYVKNCMATP------CSFTIPVVDLSKPDAKS--------IIVKACE 46
+ L+++ T Q S++++ P S IP++ L D ++ IVKACE
Sbjct: 9 LTALAEEKTLQTSFIRDEDERPKVAYNQFSNEIPIISLEGIDDETGKRAEICDKIVKACE 68
Query: 47 EFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGH------N 100
++G F+V++HGV E ISQ+ + A FF++P EK + ++ G K+ G
Sbjct: 69 DWGVFQVVDHGVDAEVISQMTTFAKEFFALPPEEKLRFDMS-----GGKKGGFIVSSHLQ 123
Query: 101 GDV--GW---VEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGL 155
G+V W V Y T + PE + V Y + + C++LD+++E +
Sbjct: 124 GEVVQDWREIVTYFSYPTRAR-DYSRWPDKPEGWIAVTQKYSEKLMELACKLLDVLSEAM 182
Query: 156 KIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTS 215
++ K L+K +D V VN YP CPE L +G HTDP I+LL +
Sbjct: 183 GLE-KEALTKACVDMDQKVV--VNFYPKCPEPDL----TLGLKRHTDPGTITLLLQDQVG 235
Query: 216 GLQISLRDG-SWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFG 274
GLQ + +G +WI+V P +F +N+GD ++NGRF++ H+ + N SRLS+ F
Sbjct: 236 GLQATKDNGKTWITVQPVEGAFVVNLGDHGHFLSNGRFKNADHQAVVNSNSSRLSIATFQ 295
Query: 275 GPPLSEKIAPLPSLLKGGKEESLYKEFTWFE 305
P + PL ++ G++ + + T+ E
Sbjct: 296 NPAPEAIVYPLK--IREGEKSIMDEPITFAE 324
>FL3H_CALCH (Q05963) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 356
Score = 129 bits (324), Expect = 1e-29
Identities = 91/308 (29%), Positives = 156/308 (50%), Gaps = 25/308 (8%)
Query: 27 IPVVDLS------KPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTE 80
IPV+ L+ + + IVKACE++G F+V++HGV + +S + A +FF +P E
Sbjct: 38 IPVISLAGIDGCRRAEICDEIVKACEDWGIFQVVDHGVDTKLLSDMTGLARDFFHLPTQE 97
Query: 81 K---EKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPEK---FRCVLD 134
K + TG + + W E ++ + + +S+ P+K +R V +
Sbjct: 98 KLRFDMTGGKKGGFIVSSHLQGEAVQDWRE-IVTYFSYPIKARDYSRWPDKPNEWRAVTE 156
Query: 135 DYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENL 194
+Y + + C++L++++E + ++ K L+K +D V VN+YP CP+ L
Sbjct: 157 EYSKVLMGLACKLLEVLSEAMGLE-KEALTKACVDMDQKVV--VNYYPKCPQPDLT---- 209
Query: 195 IGFGEHTDPQIISLLRSNNTSGLQISLRDG--SWISVPPDYSSFFINVGDSLQVMTNGRF 252
+G HTDP I+LL + GLQ + RDG SWI+V P +F +N+GD ++NGRF
Sbjct: 210 LGLKRHTDPGTITLLLQDQVGGLQAT-RDGGESWITVKPVEGAFVVNLGDHGHYLSNGRF 268
Query: 253 RSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYG 312
++ H+ + N SRLS+ F P + PL + G++ + + T+ E
Sbjct: 269 KNADHQAVVNSSTSRLSIATFQNPAPEAIVYPLK--INEGEKSIMEEPMTFMEMYKKKMS 326
Query: 313 SRLADNRL 320
+ L RL
Sbjct: 327 TDLELARL 334
>ACC1_CUCME (Q04644) 1-aminocyclopropane-1-carboxylate oxidase 1 (EC
1.14.17.4) (ACC oxidase 1) (Ethylene-forming enzyme)
(EFE) (PMEL1)
Length = 318
Score = 128 bits (322), Expect = 2e-29
Identities = 84/273 (30%), Positives = 144/273 (51%), Gaps = 16/273 (5%)
Query: 41 IVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEK-TGLANPFGYGNKRIGH 99
I AC+ +GFF+++NHG+ E + +E + + + E+ K T L+ +
Sbjct: 24 IEDACQNWGFFELVNHGIPHEFLDMVEKMTRDHYKKCMEERFKETVLSKGLEAAQAEVN- 82
Query: 100 NGDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQP 159
D+ W L+ E N+ S E+++ ++ ++ + N+ E+LDL+ E L ++
Sbjct: 83 --DMDWESTFFLRHLPESNISQMSDLDEEYKKIMKEFAKKLENLAEELLDLLCENLGLE- 139
Query: 160 KNVLSKLLMDKQSDSV-FRVNHYPACPEMGLDGENLIGFGEHTDPQ-IISLLRSNNTSGL 217
K L K + + +V++YP CP+ L + G HTD II L + + SGL
Sbjct: 140 KGYLKKAFYGSKGPTFGTKVSNYPPCPKPDL----IKGLRAHTDAGGIILLFQDDKVSGL 195
Query: 218 QISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLA-NGFQSRLSMIYFGGP 276
Q+ L+DG+WI VPP + +N+GD L+V+TNGR++SV HRVL R+S+ F P
Sbjct: 196 QL-LKDGNWIDVPPMRHAIVVNLGDQLEVITNGRYKSVMHRVLTQTSGTGRMSIASFYNP 254
Query: 277 PLSEKIAPLPSLLKGGKEE---SLYKEFTWFEY 306
I P P+L++ ++E +Y +F + +Y
Sbjct: 255 GSDAVIYPAPALVEKDQDEEKKEVYPKFVFEDY 287
>ACC1_ORYSA (Q40634) 1-aminocyclopropane-1-carboxylate oxidase 1 (EC
1.14.17.4) (ACC oxidase 1) (Ethylene-forming enzyme)
(EFE)
Length = 322
Score = 128 bits (321), Expect = 2e-29
Identities = 82/290 (28%), Positives = 143/290 (49%), Gaps = 15/290 (5%)
Query: 26 TIPVVDLS------KPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVT 79
T PV+++ +P A + ACE +GFF+++NHG+S E + ++E + +
Sbjct: 6 TFPVINMELLAGEERPAAMEQLDDACENWGFFEILNHGISTELMDEVEKMTKDHYKRVRE 65
Query: 80 EKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCA 139
++ + G + + W ++ E N+ + +R ++ +
Sbjct: 66 QRFLEFASKTLKEGCDDVNKAEKLDWESTFFVRHLPESNIADIPDLDDDYRRLMKRFAAE 125
Query: 140 VRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVF--RVNHYPACPEMGLDGENLIGF 197
+ + +LDL+ E L ++ K L+K F +V+ YP CP L + G
Sbjct: 126 LETLAERLLDLLCENLGLE-KGYLTKAFRGPAGAPTFGTKVSSYPPCPRPDL----VKGL 180
Query: 198 GEHTDPQ-IISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVK 256
HTD II L + ++ GLQ+ L+DG W+ VPP S +N+GD L+V+TNGR++SV
Sbjct: 181 RAHTDRGGIILLFQDDSVGGLQL-LKDGEWVDVPPMRHSIVVNLGDQLEVITNGRYKSVM 239
Query: 257 HRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEY 306
HRV+A +R+S+ F P I+P P+L+K + Y +F + +Y
Sbjct: 240 HRVVAQTDGNRMSIASFYNPGSDGVISPAPALVKEEEAVVAYPKFVFEDY 289
>ACC1_BRAJU (Q09052) 1-aminocyclopropane-1-carboxylate oxidase (EC
1.14.17.4) (ACC oxidase) (Ethylene-forming enzyme) (EFE)
Length = 320
Score = 127 bits (320), Expect = 3e-29
Identities = 89/287 (31%), Positives = 148/287 (51%), Gaps = 19/287 (6%)
Query: 28 PVVDLSK------PDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK 81
PVVDLSK ++I ACE +GFF+++NHG+ + + E + + + +K
Sbjct: 8 PVVDLSKLIGEERDQTMALINDACENWGFFEIVNHGLPHDLMDNAEKMTKEHYKISMEQK 67
Query: 82 EKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAVR 141
L + +R DV W L+ + N+ +++R + D+ +
Sbjct: 68 FNDMLKSKGLENLER--EVEDVDWESTFYLRHLPQSNLYDIPDMSDEYRTAMKDFGKRLE 125
Query: 142 NMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSV-FRVNHYPACPEMGLDGENLIGFGEH 200
N+ ++LDL+ E L ++ K L K+ + + +V++YPACP+ E + G H
Sbjct: 126 NLAEDLLDLLCENLGLE-KGYLKKVFHGTKGPTFGTKVSNYPACPKP----EMIKGLRAH 180
Query: 201 TDPQ-IISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRV 259
TD II L + + +GLQ+ L+DG WI VPP S IN+GD L+V+TNGR++S+ HRV
Sbjct: 181 TDAGGIILLFQDDKVTGLQL-LKDGDWIDVPPLNHSIVINLGDQLEVITNGRYKSMMHRV 239
Query: 260 LANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEY 306
+ +R+S+ F P +I+P SL +E+ Y F + +Y
Sbjct: 240 VTQKEGNRMSIASFYNPGSDAEISPASSL---ACKETEYPSFVFDDY 283
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,086,819
Number of Sequences: 164201
Number of extensions: 1711949
Number of successful extensions: 3994
Number of sequences better than 10.0: 79
Number of HSP's better than 10.0 without gapping: 59
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 3760
Number of HSP's gapped (non-prelim): 92
length of query: 329
length of database: 59,974,054
effective HSP length: 111
effective length of query: 218
effective length of database: 41,747,743
effective search space: 9101007974
effective search space used: 9101007974
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0190.3


