

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0185.17
(469 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
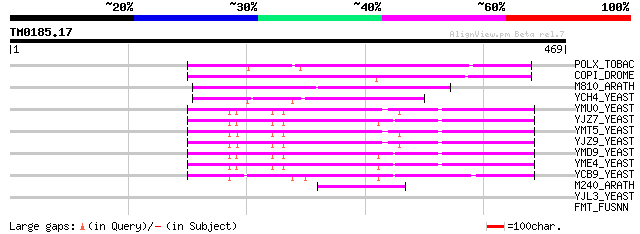
Score E
Sequences producing significant alignments: (bits) Value
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 221 3e-57
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 158 3e-38
M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810... 107 9e-23
YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein 78 4e-14
YMU0_YEAST (Q04670) Transposon Ty1 protein B 52 3e-06
YJZ7_YEAST (P47098) Transposon Ty1 protein B 52 4e-06
YMT5_YEAST (Q04214) Transposon Ty1 protein B 51 7e-06
YJZ9_YEAST (P47100) Transposon Ty1 protein B 51 7e-06
YMD9_YEAST (Q03434) Transposon Ty1 protein B 50 1e-05
YME4_YEAST (Q04711) Transposon Ty1 protein B 50 2e-05
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 48 6e-05
M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240... 47 1e-04
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 39 0.022
FMT_FUSNN (Q8RDM3) Methionyl-tRNA formyltransferase (EC 2.1.2.9) 35 0.41
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 221 bits (563), Expect = 3e-57
Identities = 120/301 (39%), Positives = 181/301 (59%), Gaps = 14/301 (4%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRD--GEIIGLSQ 208
++ LYVDDMLI G D + K K LS SFDMKDLG A ILG+KI+R+ + LSQ
Sbjct: 1003 IILLLYVDDMLIVGKDKGLIAKLKGDLSKSFDMKDLGPAQQILGMKIVRERTSRKLWLSQ 1062
Query: 209 SHYIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCP--------VAQLEYSKAIRCLMYA 260
YIE+VL++F+ + K VSTP + KL K CP +A++ YS A+ LMYA
Sbjct: 1063 EKYIERVLERFNMKNAKPVSTPLAGHLKLSK-KMCPTTVEEKGNMAKVPYSSAVGSLMYA 1121
Query: 261 MTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSVLEGYTYA 320
M CTRPDIA+ VG +SR+ NP K+HW V +L+YL GT L + G +L+GYT A
Sbjct: 1122 MVCTRPDIAHAVGVVSRFLENPGKEHWEAVKWILRYLRGTTGDCLCFGGSDPILKGYTDA 1181
Query: 321 SWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLL 380
+ + S++G +F GGA+SW SK Q C+A ST AE+IA KE + L+ L
Sbjct: 1182 DMAGDIDNRKSSTGYLFTFSGGAISWQSKLQKCVALSTTEAEYIAATETGKEMIWLKRFL 1241
Query: 381 YEILIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVR 440
E+ + K + ++CDSQ + + + +Y+ +++HI +R+ +++++ + + ++ +
Sbjct: 1242 QELGLHQK---EYVVYCDSQSAIDLSKNSMYHARTKHIDVRYHWIREMVDDESLKVLKIS 1298
Query: 441 T 441
T
Sbjct: 1299 T 1299
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 158 bits (399), Expect = 3e-38
Identities = 95/295 (32%), Positives = 154/295 (52%), Gaps = 6/295 (2%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSH 210
+ LYVDD++I D+ + K +L F M DL E +GI+I + I LSQS
Sbjct: 1082 IYVLLYVDDVVIATGDMTRMNNFKRYLMEKFRMTDLNEIKHFIGIRIEMQEDKIYLSQSA 1141
Query: 211 YIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQLEYSKAIRCLMYAMTCTRPDIAY 270
Y++K+L KF+ +C +VSTP + I CLMY M CTRPD+
Sbjct: 1142 YVKKILSKFNMENCNAVSTPLPSKINYELLNSDEDCNTPCRSLIGCLMYIMLCTRPDLTT 1201
Query: 271 VVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYN---GYPSVLEGYTYASWVTCVK 327
V LSRY+ + + W + RVL+YL GTI+ LI+ + + + GY + W
Sbjct: 1202 AVNILSRYSSKNNSELWQNLKRVLRYLKGTIDMKLIFKKNLAFENKIIGYVDSDWAGSEI 1261
Query: 328 DHASTSG*IFNL-GGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIW 386
D ST+G +F + + W +K+Q +A S+ AE++AL +E + L+ LL I I
Sbjct: 1262 DRKSTTGYLFKMFDFNLICWNTKRQNSVAASSTEAEYMALFEAVREALWLKFLLTSINI- 1320
Query: 387 PKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRT 441
KL + I+ D+Q +S A + + +++HI +++ ++ + N VI + ++ T
Sbjct: 1321 -KLENPIKIYEDNQGCISIANNPSCHKRAKHIDIKYHFAREQVQNNVICLEYIPT 1374
>M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810
(ORF240b)
Length = 240
Score = 107 bits (266), Expect = 9e-23
Identities = 68/219 (31%), Positives = 108/219 (49%), Gaps = 2/219 (0%)
Query: 155 LYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSHYIEK 214
LYVDD+L+ G+ + LS++F MKDLG LGI+I + LSQ+ Y E+
Sbjct: 5 LYVDDILLTGSSNTLLNMLIFQLSSTFSMKDLGPVHYFLGIQIKTHPSGLFLSQTKYAEQ 64
Query: 215 VLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQLEYSKAIRCLMYAMTCTRPDIAYVVGR 274
+L DCK +STP ++ + L Y +T TRPDI+Y V
Sbjct: 65 ILNNAGMLDCKPMSTPLPLKLNSSVSTAKYPDPSDFRSIVGALQY-LTLTRPDISYAVNI 123
Query: 275 LSRYTCNPSKDHWHVVNRVLKYLNGTINYSL-IYNGYPSVLEGYTYASWVTCVKDHASTS 333
+ + P+ + ++ RVL+Y+ GTI + L I+ ++ + + W C ST+
Sbjct: 124 VCQRMHEPTLADFDLLKRVLRYVKGTIFHGLYIHKNSKLNVQAFCDSDWAGCTSTRRSTT 183
Query: 334 G*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKE 372
G LG +SW +K+Q ++ S+ E+ ALA + E
Sbjct: 184 GFCTFLGCNIISWSAKRQPTVSRSSTETEYRALALTAAE 222
>YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein
Length = 308
Score = 78.2 bits (191), Expect = 4e-14
Identities = 65/203 (32%), Positives = 91/203 (44%), Gaps = 11/203 (5%)
Query: 155 LYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIR--DGEIIGLSQSHYI 212
+YVDD+L+ ++ K L+ + MKDLG+ D LG+ I + +G+I LS YI
Sbjct: 86 VYVDDLLVAAPSPKIYDRVKQELTKLYSMKDLGKVDKFLGLNIHQSTNGDIT-LSLQDYI 144
Query: 213 EKVLKKFDHFDCKSVSTPFDQNTKL----QPHKRCPVAQLEYSKAIRCLMYAMTCTRPDI 268
K + + K TP + L PH + Y + L++ RPDI
Sbjct: 145 AKAASESEINTFKLTQTPLCNSKPLFETTSPHLKDITP---YQSIVGQLLFCANTGRPDI 201
Query: 269 AYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIY-NGYPSVLEGYTYASWVTCVK 327
+Y V LSR+ P H RVL+YL T + L Y +G L Y AS
Sbjct: 202 SYPVSLLSRFLREPRAIHLESARRVLRYLYTTRSMCLKYRSGSQVALTVYCDASHGAIHD 261
Query: 328 DHASTSG*IFNLGGGAVSWGSKK 350
ST G + L G V+W SKK
Sbjct: 262 LPHSTGGYVTLLAGAPVTWSSKK 284
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 52.4 bits (124), Expect = 3e-06
Identities = 72/319 (22%), Positives = 133/319 (41%), Gaps = 33/319 (10%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMK--DLGEAD-----VILGIKI-IRDGE 202
V CL+VDDM++F DLN +K + L +D K +LGE+D ILG++I + G+
Sbjct: 993 VTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRGK 1052
Query: 203 IIGLSQSHYIEKVLKKFD---HFDCKSVSTP------FDQNTKLQPHKRCPVAQLEYSKA 253
+ L + + + + K + + + +S P DQ + E K
Sbjct: 1053 YMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHEMQKL 1112
Query: 254 IRCLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV 313
I Y R D+ Y + L+++ PSK + +++++ T + LI++ V
Sbjct: 1113 IGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWNTRDKQLIWHKSKPV 1172
Query: 314 LEGYTYASWVTCVKD--------HASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIA 365
+ + + D + S G I+ L G + S K + ST AE
Sbjct: 1173 KP----TNKLVVISDASYGNQPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAE--- 1225
Query: 366 LAACSKETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYS-QVYNGKSRHIVLRHSH 424
+ A S+ L NL + + K + DS+ T+S S ++R +
Sbjct: 1226 IHAISESVPLLNNLSHLVQELNKKPITKGLLTDSKSTISIIISNNEEKFRNRFFGTKAMR 1285
Query: 425 VKDLITNGVISIVFVRTVK 443
++D ++ + + ++ T K
Sbjct: 1286 LRDEVSGNHLHVCYIETKK 1304
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 51.6 bits (122), Expect = 4e-06
Identities = 75/316 (23%), Positives = 134/316 (41%), Gaps = 27/316 (8%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMK--DLGEAD-----VILGIKI-IRDGE 202
V CL+VDDM++F DLN +K + L +D K +LGE+D ILG++I + G+
Sbjct: 1420 VTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRGK 1479
Query: 203 IIGLSQSHYIEKVLKKFD---HFDCKSVSTP------FDQNTKLQPHKRCPVAQLEYSKA 253
+ L + + + + K + + + +S P DQ+ E K
Sbjct: 1480 YMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQDELEIDEDEYKEKVHEMQKL 1539
Query: 254 IRCLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGY--- 310
I Y R D+ Y + L+++ PS+ + +++++ T + LI++
Sbjct: 1540 IGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHKNKPT 1599
Query: 311 --PSVLEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAA 368
+ L + AS+ + S G IF L G + S K + ST AE + A
Sbjct: 1600 EPDNKLVAISDASYGN-QPYYKSQIGNIFLLNGKVIGGKSTKASLTCTSTTEAE---IHA 1655
Query: 369 CSKETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYS-QVYNGKSRHIVLRHSHVKD 427
S+ L NL Y I K + DS+ T+S S ++R + ++D
Sbjct: 1656 ISESVPLLNNLSYLIQELNKKPIIKGLLTDSRSTISIIKSTNEEKFRNRFFGTKAMRLRD 1715
Query: 428 LITNGVISIVFVRTVK 443
++ + + ++ T K
Sbjct: 1716 EVSGNNLYVYYIETKK 1731
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 50.8 bits (120), Expect = 7e-06
Identities = 72/319 (22%), Positives = 132/319 (40%), Gaps = 33/319 (10%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMK--DLGEADV-----ILGIKI-IRDGE 202
V CL+VDDM++F +LN ++ L +D K +LGE+D ILG++I + G+
Sbjct: 993 VTICLFVDDMVLFSKNLNSNKRIIDKLKMQYDTKIINLGESDEEIQYDILGLEIKYQRGK 1052
Query: 203 IIGLSQSHYIEKVLKKFD---HFDCKSVSTP------FDQNTKLQPHKRCPVAQLEYSKA 253
+ L + + + + K + + + +S P DQ + E K
Sbjct: 1053 YMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHEMQKL 1112
Query: 254 IRCLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV 313
I Y R D+ Y + L+++ PSK + +++++ T + LI++ V
Sbjct: 1113 IGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWNTRDKQLIWHKSKPV 1172
Query: 314 LEGYTYASWVTCVKD--------HASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIA 365
+ + + D + S G I+ L G + S K + ST AE
Sbjct: 1173 KP----TNKLVVISDASYGNQPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAE--- 1225
Query: 366 LAACSKETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYS-QVYNGKSRHIVLRHSH 424
+ A S+ L NL Y I K + DS+ T+S S ++R +
Sbjct: 1226 IHAISESVPLLNNLSYLIQELDKKPITKGLLTDSKSTISIIISNNEEKFRNRFFGTKAMR 1285
Query: 425 VKDLITNGVISIVFVRTVK 443
++D ++ + + ++ T K
Sbjct: 1286 LRDEVSGNHLHVCYIETKK 1304
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 50.8 bits (120), Expect = 7e-06
Identities = 72/319 (22%), Positives = 132/319 (40%), Gaps = 33/319 (10%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMK--DLGEADV-----ILGIKI-IRDGE 202
V CL+VDDM++F +LN ++ L +D K +LGE+D ILG++I + G+
Sbjct: 1420 VTICLFVDDMVLFSKNLNSNKRIIEKLKMQYDTKIINLGESDEEIQYDILGLEIKYQRGK 1479
Query: 203 IIGLSQSHYIEKVLKKFD---HFDCKSVSTP------FDQNTKLQPHKRCPVAQLEYSKA 253
+ L + + + + K + + + +S P DQ + E K
Sbjct: 1480 YMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHEMQKL 1539
Query: 254 IRCLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV 313
I Y R D+ Y + L+++ PSK + +++++ T + LI++ V
Sbjct: 1540 IGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWNTRDKQLIWHKSKPV 1599
Query: 314 LEGYTYASWVTCVKD--------HASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIA 365
+ + + D + S G I+ L G + S K + ST AE
Sbjct: 1600 KP----TNKLVVISDASYGNQPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAE--- 1652
Query: 366 LAACSKETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYS-QVYNGKSRHIVLRHSH 424
+ A S+ L NL Y I K + DS+ T+S S ++R +
Sbjct: 1653 IHAISESVPLLNNLSYLIQELDKKPITKGLLTDSKSTISIIISNNEEKFRNRFFGTKAMR 1712
Query: 425 VKDLITNGVISIVFVRTVK 443
++D ++ + + ++ T K
Sbjct: 1713 LRDEVSGNHLHVCYIETKK 1731
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 50.4 bits (119), Expect = 1e-05
Identities = 74/316 (23%), Positives = 134/316 (41%), Gaps = 27/316 (8%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMK--DLGEAD-----VILGIKI-IRDGE 202
V CL+VDDM++F DLN +K + L +D K +LGE+D ILG++I + G+
Sbjct: 993 VTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRGK 1052
Query: 203 IIGLSQSHYIEKVLKKFD---HFDCKSVSTP------FDQNTKLQPHKRCPVAQLEYSKA 253
+ L + + + + K + + + +S P DQ+ E K
Sbjct: 1053 YMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQDELEIDEDEYKEKVHEMQKL 1112
Query: 254 IRCLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGY--- 310
I Y R D+ Y + L+++ PS+ + +++++ T + LI++
Sbjct: 1113 IGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHKNKPT 1172
Query: 311 --PSVLEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAA 368
+ L + AS+ + S G I+ L G + S K + ST AE + A
Sbjct: 1173 EPDNKLVAISDASYGN-QPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAE---IHA 1228
Query: 369 CSKETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYS-QVYNGKSRHIVLRHSHVKD 427
S+ L NL Y I K + DS+ T+S S ++R + ++D
Sbjct: 1229 ISESVPLLNNLSYLIQELNKKPIIKGLLTDSRSTISIIKSTNEEKFRNRFFGTKAMRLRD 1288
Query: 428 LITNGVISIVFVRTVK 443
++ + + ++ T K
Sbjct: 1289 EVSGNNLYVYYIETKK 1304
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 49.7 bits (117), Expect = 2e-05
Identities = 72/316 (22%), Positives = 134/316 (41%), Gaps = 27/316 (8%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMK--DLGEAD-----VILGIKI-IRDGE 202
V CL+VDDM++F DLN +K + L +D K +LGE+D ILG++I + G+
Sbjct: 993 VTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRGK 1052
Query: 203 IIGLSQSHYIEKVLKKFD---HFDCKSVSTP------FDQNTKLQPHKRCPVAQLEYSKA 253
+ L + + + + K + + + +S P DQ+ E K
Sbjct: 1053 YMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQDELEIDEDEYKEKVHEMQKL 1112
Query: 254 IRCLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGY--- 310
I Y R D+ Y + L+++ PS+ + +++++ T + LI++
Sbjct: 1113 IGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHKNKPT 1172
Query: 311 --PSVLEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAA 368
+ L + AS+ + S G I+ L G + S K + ST AE + A
Sbjct: 1173 EPDNKLVAISDASYGN-QPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAE---IHA 1228
Query: 369 CSKETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYS-QVYNGKSRHIVLRHSHVKD 427
S+ L NL + + K + DS+ T+S S ++R + ++D
Sbjct: 1229 ISESVPLLNNLSHLVQELNKKPITKGLLTDSKSTISIIISNNEEKFRNRFFGTKAMRLRD 1288
Query: 428 LITNGVISIVFVRTVK 443
++ + + ++ T K
Sbjct: 1289 EVSGNHLHVCYIETKK 1304
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 47.8 bits (112), Expect = 6e-05
Identities = 72/318 (22%), Positives = 132/318 (40%), Gaps = 31/318 (9%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMK--DLGEADVILGIKIIRDGEIIGLSQ 208
V CL+VDDM++F DLN +K + L +D K +LGE+D + I+ G I +
Sbjct: 1435 VTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDIL--GLEIKYQR 1492
Query: 209 SHYIEKVLKKFDHFDCKSVSTPFDQNTKL-----QPHKRCPVAQL------------EYS 251
S Y++ ++K ++ P + K QP +L E
Sbjct: 1493 SKYMKLGMEKSLTEKLPKLNVPLNPKGKKLRAPGQPGHYIDQDELEIDEDEYKEKVHEMQ 1552
Query: 252 KAIRCLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGY- 310
K I Y R D+ Y + L+++ PS+ + +++++ T + LI++
Sbjct: 1553 KLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHKNK 1612
Query: 311 ----PSVLEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIAL 366
+ L + AS+ + S G IF L G + S K + ST AE A+
Sbjct: 1613 PTKPDNKLVAISDASYGN-QPYYKSQIGNIFLLNGKVIGGKSTKASLTCTSTTEAEIHAV 1671
Query: 367 AACSKETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYS-QVYNGKSRHIVLRHSHV 425
+ L +L+ E+ P + + DS+ T+S S ++R + +
Sbjct: 1672 SEAIPLLNNLSHLVQELNKKPIIK---GLLTDSRSTISIIKSTNEEKFRNRFFGTKAMRL 1728
Query: 426 KDLITNGVISIVFVRTVK 443
+D ++ + + ++ T K
Sbjct: 1729 RDEVSGNNLYVYYIETKK 1746
>M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240
(ORF111a)
Length = 111
Score = 47.0 bits (110), Expect = 1e-04
Identities = 22/75 (29%), Positives = 38/75 (50%), Gaps = 1/75 (1%)
Query: 261 MTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV-LEGYTY 319
+T TRPD+ + V RLS+++ V +VL Y+ GT+ L Y+ + L+ +
Sbjct: 3 LTITRPDLTFAVNRLSQFSSASRTAQMQAVYKVLHYVKGTVGQGLFYSATSDLQLKAFAD 62
Query: 320 ASWVTCVKDHASTSG 334
+ W +C S +G
Sbjct: 63 SDWASCPDTRRSVTG 77
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 39.3 bits (90), Expect = 0.022
Identities = 38/168 (22%), Positives = 84/168 (49%), Gaps = 20/168 (11%)
Query: 149 KAVMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEA--DV----ILGIKIIRDGE 202
K +M +YVDD +I ++ +++ + L ++F++K G DV ILG+ ++ +
Sbjct: 1454 KNLMIAVYVDDCVIAASNEQRLDEFINKLKSNFELKITGTLIDDVLDTDILGMDLVYNKR 1513
Query: 203 I--IGLSQSHYIEKVLKKFDHFDCKSV---STPFDQNTKLQPHKR-CPVAQLEYSKAIRC 256
+ I L+ +I ++ KK++ + K + S P K+ P K +++ E+ + +
Sbjct: 1514 LGTIDLTLKSFINRMDKKYNE-ELKKIRKSSIPHMSTYKIDPKKDVLQMSEEEFRQGVLK 1572
Query: 257 LM-------YAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYL 297
L Y R DI + V +++R P + ++++ ++++YL
Sbjct: 1573 LQQLLGELNYVRHKCRYDIEFAVKKVARLVNYPHERVFYMIYKIIQYL 1620
>FMT_FUSNN (Q8RDM3) Methionyl-tRNA formyltransferase (EC 2.1.2.9)
Length = 310
Score = 35.0 bits (79), Expect = 0.41
Identities = 22/81 (27%), Positives = 41/81 (50%), Gaps = 4/81 (4%)
Query: 149 KAVMTCLYVDDMLIFGTDL----NEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEII 204
K+ ++ +YV++ L G + E+ +FLS +KD+G ++ I++I+ GE+
Sbjct: 129 KSGISIMYVEEELDAGDVILQEETEISDEDTFLSLHDRLKDMGADLLLKAIELIKKGEVK 188
Query: 205 GLSQSHYIEKVLKKFDHFDCK 225
Q + +K F DCK
Sbjct: 189 AQKQDKKLVTFVKPFRKEDCK 209
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.351 0.154 0.546
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,650,604
Number of Sequences: 164201
Number of extensions: 1863568
Number of successful extensions: 8148
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 8124
Number of HSP's gapped (non-prelim): 15
length of query: 469
length of database: 59,974,054
effective HSP length: 114
effective length of query: 355
effective length of database: 41,255,140
effective search space: 14645574700
effective search space used: 14645574700
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0185.17


