

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.7
(1813 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
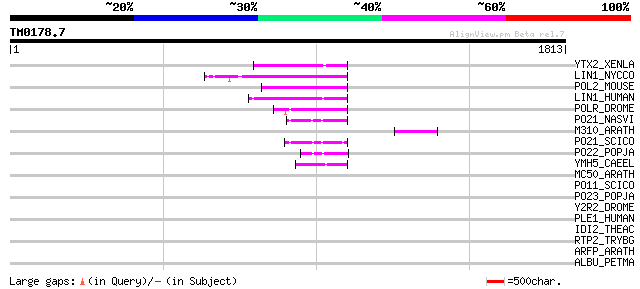
Score E
Sequences producing significant alignments: (bits) Value
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 142 1e-32
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 121 2e-26
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 119 8e-26
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 118 1e-25
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 97 4e-19
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 73 8e-12
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 68 2e-10
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 61 3e-08
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 54 3e-06
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 53 9e-06
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 43 0.007
PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type... 41 0.026
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 40 0.059
Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I retro... 38 0.29
PLE1_HUMAN (Q15149) Plectin 1 (PLTN) (PCN) (Hemidesmosomal prote... 34 3.2
IDI2_THEAC (Q9HLX2) Isopentenyl-diphosphate delta-isomerase (EC ... 34 4.2
RTP2_TRYBG (P15594) Retrotransposable element SLACS 132 kDa prot... 33 7.2
ARFP_ARATH (Q93YR9) Auxin response factor 16 33 7.2
ALBU_PETMA (Q91274) Serum albumin SDS-1 precursor 33 9.4
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein (ORF
2)
Length = 1308
Score = 142 bits (357), Expect = 1e-32
Identities = 89/313 (28%), Positives = 162/313 (51%), Gaps = 9/313 (2%)
Query: 796 FQKARENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDRQLLAEEIQNYFQSL 855
F ++R +C DR + +F+A + R +I L DG+ +D + + + ++++Q+L
Sbjct: 357 FVRSRMQLLCDMDRGSRFFYALEKKKGNRKQITCLFAEDGTPLEDPEAIRDRARSFYQNL 416
Query: 856 FAADPDNTDVNMQ-HVMMPRLTEAERHKAEAEVIKEEVFQALLRMRSYSALGPDGFHPFF 914
F+ DP + D + +P ++E + + E + +E+ QAL M + G DG F
Sbjct: 417 FSPDPISPDACEELWDGLPVVSERRKERLETPITLDELSQALRLMPHNKSPGLDGLTIEF 476
Query: 915 FKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVYK 974
F+ +W+ +G D +++ +AF+ G + S ++ L+PK +K +RP+SL + YK
Sbjct: 477 FQFFWDTLGPDFHRVLTEAFKKGELPLSCRRAVLSLLPKKGDLRLIKNWRPVSLLSTDYK 536
Query: 975 LITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIH---HMGASMAKKGTLAF 1031
++ K + R++ +L +I P Q+ +PG I DN FL ++++H G S+A
Sbjct: 537 IVAKAISLRLKSVLAEVIHPDQSYTVPGRTIFDNVFLIRDLLHFARRTGLSLA-----FL 591
Query: 1032 KIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKPGRGL 1091
+D EKA+D V +L TL + F + V + + + + N S GRG+
Sbjct: 592 SLDQEKAFDRVDHQYLIGTLQAYSFGPQFVGYLKTMYASAECLVKINWSLTAPLAFGRGV 651
Query: 1092 RKGDPLSPYLFVL 1104
R+G PLS L+ L
Sbjct: 652 RQGCPLSGQLYSL 664
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 121 bits (303), Expect = 2e-26
Identities = 114/493 (23%), Positives = 215/493 (43%), Gaps = 40/493 (8%)
Query: 635 YTWFRKQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHRIHSDHCPLLINCSGPEIVH- 693
YT+F + ++D +G + F + +E + I SDH + + + +H
Sbjct: 190 YTFFSSAHGTY---SKIDHILGHKSNLSKFKK--IEIIPCIFSDHHGIKVELNNNRNLHT 244
Query: 694 HDRPFRF-----LASWVEHPQYKEVV-------------QSAWTGETETVAGKLNKVRTK 735
H + ++ +WV KE+ Q+ W + GK ++
Sbjct: 245 HTKTWKLNNLMLKDTWVIDEIKKEITKFLEQNNNQDTNYQNLWDTAKAVLRGKFIALQAF 304
Query: 736 SCKFNKEVYGSIFQRKRWVEGRLRGVQRELNYRVTFDMTRLETELQAEYRFLLRQEELLW 795
K +E ++ G L+ +++E + + T+++AE + + +
Sbjct: 305 LKKTEREEVNNLM-------GHLKQLEKEEHSNPKPSRRKEITKIRAELNEIENKRIIQQ 357
Query: 796 FQKARENRVCFDDRNTAYFHAHTIIRRKRNK--IHRLKLNDGSWCDDRQLLAEEIQNYFQ 853
K++ F+ N + R+KR K I ++ + D + + + Y++
Sbjct: 358 INKSKS--WFFEKINKIDKPLANLTRKKRVKSLISSIRNGNDEITTDPSEIQKILNEYYK 415
Query: 854 SLFAADPDNT---DVNMQHVMMPRLTEAERHKAEAEVIKEEVFQALLRMRSYSALGPDGF 910
L++ +N D ++ +PRL++ E + E+ + + + GPDGF
Sbjct: 416 KLYSHKYENLKEIDQYLEACHLPRLSQKEVEMLNRPISSSEIASTIQNLPKKKSPGPDGF 475
Query: 911 HPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKE-FRPISLC 969
F++ + E + L L ++ + G++ ++ E I LIPK T KE +RPISL
Sbjct: 476 TSEFYQTFKEELVPILLNLFQNIEKEGILPNTFYEANITLIPKPGKDPTRKENYRPISLM 535
Query: 970 NVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGASMAKKGTL 1029
N+ K++ K++ NRI+ + +II Q F+PG N + +I H+ + K +
Sbjct: 536 NIDAKILNKILTNRIQQHIKKIIHHDQVGFIPGSQGWFNIRKSINVIQHIN-KLKNKDHM 594
Query: 1030 AFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKPGR 1089
ID EKA+D++ F+ TL + G ++LI S +++ NG +L SF
Sbjct: 595 ILSIDAEKAFDNIQHPFMIRTLKKIGIEGTFLKLIEAIYSKPTANIILNGVKLKSFPLRS 654
Query: 1090 GLRKGDPLSPYLF 1102
G R+G PLSP LF
Sbjct: 655 GTRQGCPLSPLLF 667
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 119 bits (298), Expect = 8e-26
Identities = 77/285 (27%), Positives = 140/285 (49%), Gaps = 5/285 (1%)
Query: 822 RKRNKIHRLKLNDGSWCDDRQLLAEEIQNYFQSLFAADPDNTDVN---MQHVMMPRLTEA 878
R + I++++ G D + + I+++++ L++ +N D + +P+L +
Sbjct: 411 RDKILINKIRNEKGDITTDPEEIQNTIRSFYKRLYSTKLENLDEMDKFLDRYQVPKLNQD 470
Query: 879 ERHKAEAEVIKEEVFQALLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGM 938
+ + + +E+ + + + + GPDGF F++ + E + L KL G
Sbjct: 471 QVDHLNSPISPKEIEAVINSLPTKKSPGPDGFSAEFYQTFKEDLIPILHKLFHKIEVEGT 530
Query: 939 IDDSLLETLIVLIPKVDS-PSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQN 997
+ +S E I LIPK P+ ++ FRPISL N+ K++ K++ NRI+ + II P Q
Sbjct: 531 LPNSFYEATITLIPKPQKDPTKIENFRPISLMNIDAKILNKILANRIQEHIKAIIHPDQV 590
Query: 998 SFLPGMGIMDNAFLAQEIIHHMGASMAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFP 1057
F+PGM N + +IH++ + K + +D EKA+D + F+ L + G
Sbjct: 591 GFIPGMQGWFNIRKSINVIHYIN-KLKDKNHMIISLDAEKAFDKIQHPFMIKVLERSGIQ 649
Query: 1058 NRLVQLIMRCVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSPYLF 1102
+ +I S ++ NG +L + G R+G PLSPYLF
Sbjct: 650 GPYLNMIKAIYSKPVANIKVNGEKLEAIPLKSGTRQGCPLSPYLF 694
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 118 bits (296), Expect = 1e-25
Identities = 91/330 (27%), Positives = 161/330 (48%), Gaps = 11/330 (3%)
Query: 779 ELQAEYRFLLRQEELLWFQKARENRVCFDDR-NTAYFHAHTIIRRKR--NKIHRLKLNDG 835
+++AE + + Q+ L QK E+R F ++ N +I++KR N+I +K + G
Sbjct: 341 KIRAELKEIETQKTL---QKINESRSWFFEKINKIDRPLARLIKKKREKNQIDTIKNDRG 397
Query: 836 SWCDDRQLLAEEIQNYFQSLFAADPDNT---DVNMQHVMMPRLTEAERHKAEAEVIKEEV 892
D + I+ Y++ L+A +N D + +PRL + E + E+
Sbjct: 398 DITTDPTEIQTTIREYYKHLYANKLENLEEMDKFLDTYTLPRLNQEEVESLNRPITSSEI 457
Query: 893 FQALLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIP 952
+ + + + GP+GF F+++Y E + L KL + + G++ +S E I+LIP
Sbjct: 458 EAIINSLPNKKSPGPEGFTAEFYQRYKEELVPFLLKLFQSIEKEGILPNSFYEASIILIP 517
Query: 953 KVDSPSTVKE-FRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFL 1011
K +T KE FRPISL N+ K++ K++ N+I+ + ++I Q F+P M N
Sbjct: 518 KPGRDTTKKENFRPISLMNIDAKILNKILANQIQQHIKKLIHHDQVGFIPAMQGWFNIRK 577
Query: 1012 AQEIIHHMGASMAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGS 1071
+ II H+ + + ID EKA+D + F+ L + G +++I
Sbjct: 578 SINIIQHINRT-KDTNHMIISIDAEKAFDKIQQPFMLKPLNKLGIDGTYLKIIRAIYDKP 636
Query: 1072 NLSMLWNGSRLPSFKPGRGLRKGDPLSPYL 1101
+++ NG +L + G R+G PLSP L
Sbjct: 637 TANIILNGQKLEAPPLKTGTRQGCPLSPLL 666
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 97.1 bits (240), Expect = 4e-19
Identities = 73/260 (28%), Positives = 119/260 (45%), Gaps = 23/260 (8%)
Query: 861 DNTDVNMQHVMMPR----LTEAERHKAEAEVIK-----EEVFQALL-------RMRSYSA 904
D + + Q +M+P +T+ EVI+ E V+ A+ R+ S+
Sbjct: 300 DESVMPSQEIMVPYWREVMTQPSPSSCSGEVIQMDHSLERVWSAITEQDLRASRVSLSSS 359
Query: 905 LGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFR 964
GPDG P K E+ + +++ G + S+ V IPK + ++FR
Sbjct: 360 PGPDGITP---KSAREVPSGIMLRIMNLILWCGNLPHSIRLARTVFIPKTVTAKRPQDFR 416
Query: 965 PISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGASMA 1024
PIS+ +V+ + + ++ R+ +N P Q FLP G DNA + ++ H
Sbjct: 417 PISVPSVLVRQLNAILATRLNSSINW--DPRQRGFLPTDGCADNATIVDLVLRHSHKHF- 473
Query: 1025 KKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPS 1084
+ +D+ KA+DS+S A + DTL +G P V + G S+ +G
Sbjct: 474 -RSCYIANLDVSKAFDSLSHASIYDTLRAYGAPKGFVDYVQNTYEGGGTSLNGDGWSSEE 532
Query: 1085 FKPGRGLRKGDPLSPYLFVL 1104
F P RG+++GDPLSP LF L
Sbjct: 533 FVPARGVKQGDPLSPILFNL 552
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 72.8 bits (177), Expect = 8e-12
Identities = 56/200 (28%), Positives = 90/200 (45%), Gaps = 8/200 (4%)
Query: 903 SALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKE 962
+A GPDG W + + + L G L++ VLIPK
Sbjct: 333 TAAGPDGMTT----TAWNSIDECIKSLFNMIMYHGQCPRRYLDSRTVLIPKEPGTMDPAC 388
Query: 963 FRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGAS 1022
FRP+S+ +V + +++ NRI + ++ Q +F+ G+ +N L +I A
Sbjct: 389 FRPLSIASVALRHFHRILANRIGE--HGLLDTRQRAFIVADGVAENTSLLSAMIKE--AR 444
Query: 1023 MAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRL 1082
M KG +D++KA+DSV + D L + P + IM S + ++
Sbjct: 445 MKIKGLYIAILDVKKAFDSVEHRSILDALRRKKLPLEMRNYIMWVYRNSKTRLEVVKTKG 504
Query: 1083 PSFKPGRGLRKGDPLSPYLF 1102
+P RG+R+GDPLSP LF
Sbjct: 505 RWIRPARGVRQGDPLSPLLF 524
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 68.2 bits (165), Expect = 2e-10
Identities = 42/142 (29%), Positives = 64/142 (44%), Gaps = 2/142 (1%)
Query: 1256 AIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQRGWHRVNWKTLTQPKEH-GGLAIKD 1314
A+P Y M F L + + K+ M F W +R V W+ L + KE GGL +D
Sbjct: 2 ALPVYAMSCFRLSKLLCKKLTSAMTEFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRD 61
Query: 1315 MGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHGESIFSVHRRANASPIWQGILKARDQ 1374
+G FN +LL K + + + L ++L +Y S+ S W+ I+ R+
Sbjct: 62 LGWFNQALLAKQSFRIIHQPHTLLSRLLRSRYFPHSSMMECSVGTRPSYAWRSIIHGREL 121
Query: 1375 LQEGFQFRMGDG-SSSVWYHDW 1395
L G +GDG + VW W
Sbjct: 122 LSRGLLRTIGDGIHTKVWLDRW 143
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 60.8 bits (146), Expect = 3e-08
Identities = 56/209 (26%), Positives = 90/209 (42%), Gaps = 12/209 (5%)
Query: 898 RMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFET-GMIDDSLLETLIVLIPKVDS 956
R + GPDG P + W + D +L+ + F G + + + V PK++
Sbjct: 168 RASNEKGAGPDGVTP----RSWNALDDRYKRLLYNIFVFYGRVPSPIKGSRTVFTPKIEG 223
Query: 957 PSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEII 1016
FRP+S+C+V+ + K++ R Q ++LP G+ N + II
Sbjct: 224 GPDPGVFRPLSICSVILREFNKILARRFVSCYT--YDERQTAYLPIDGVCINVSMLTAII 281
Query: 1017 HHMGASMAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSML 1076
A +K +DL KA++SV + L D + + G P +V I + M
Sbjct: 282 AE--AKRLRKELHIAILDLVKAFNSVYHSALIDAITEAGCPPGVVDYIADMYNNVITEMQ 339
Query: 1077 WNGS-RLPSFKPGRGLRKGDPLSPYLFVL 1104
+ G L S G+ +GDPLS LF L
Sbjct: 340 FEGKCELASIL--AGVYQGDPLSGPLFTL 366
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 54.3 bits (129), Expect = 3e-06
Identities = 43/165 (26%), Positives = 77/165 (46%), Gaps = 11/165 (6%)
Query: 950 LIPKVDSPSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNA 1009
LIPK +RPI++ + + +L+ +++ R+ + + P Q + G + N+
Sbjct: 63 LIPKDGDLENPSNWRPITIASALQRLLHRILAKRLEAAVE--LHPAQKGYARIDGTLVNS 120
Query: 1010 FLAQEIIHHMGASMAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVS 1069
L I +K +D+ KA+D+VS + + L + G I +S
Sbjct: 121 LLLDTYISSRREQ--RKTYNVVSLDVRKAFDTVSHSSICRALQRLGIDEGTSNYITGSLS 178
Query: 1070 GSNLSM-LWNGSRLPSFKPGRGLRKGDPLSPYLF------VLCSL 1107
S ++ + GS+ RG+++GDPLSP+LF +LCSL
Sbjct: 179 DSTTTIRVGPGSQTRKICIRRGVKQGDPLSPFLFNAVLDELLCSL 223
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 52.8 bits (125), Expect = 9e-06
Identities = 42/173 (24%), Positives = 73/173 (41%), Gaps = 3/173 (1%)
Query: 933 AFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRII 992
+F I + +I+ IPK +PS+ +RPISL + +++ +++ +RIR + ++
Sbjct: 655 SFSDSAIPNRWKHAVIIPIPKKGNPSSPSNYRPISLTDPFARIMERIICSRIRSEYSHLL 714
Query: 993 GPMQNSFLPGMGIMDNAFLAQEIIHHMGASMAKKGTLAFKIDLEKAYDSVSWAFLEDTLV 1052
P Q+ FL + + + H + + L F D KA+D VS L L
Sbjct: 715 SPHQHGFLNFRSCPSSLVRSISLYHSILKNEKSLDILFF--DFAKAFDKVSHPILLKKLA 772
Query: 1053 QFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKP-GRGLRKGDPLSPYLFVL 1104
FG + S+ N + P G+ +G P LF+L
Sbjct: 773 LFGLDKLTCSWFKEFLHLRTFSVKINKFVSSNAYPISSGVPQGSVSGPLLFIL 825
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 43.1 bits (100), Expect = 0.007
Identities = 19/34 (55%), Positives = 24/34 (69%)
Query: 1073 LSMLWNGSRLPSFKPGRGLRKGDPLSPYLFVLCS 1106
L + NG+ P RGLR+GDPLSPYLF+LC+
Sbjct: 10 LLFIINGAPQGLVTPSRGLRQGDPLSPYLFILCT 43
>PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type I
retrotransposable element R1 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1004
Score = 41.2 bits (95), Expect = 0.026
Identities = 41/177 (23%), Positives = 81/177 (45%), Gaps = 12/177 (6%)
Query: 876 TEAERHKAEAEVIK-----EEVFQALLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLV 930
+E +R KA A + +EV ++ R + + GPDG + W + + ++ L
Sbjct: 389 SEIDRLKAIARPLPPDLEMDEVSDSVRRCKVRKSPGPDGIVGEMVRAVWGAIPEYMFCLY 448
Query: 931 RDAF-ETGMIDDSLLETLIVLIPKVDS-PSTVKEFRPISLCNVVYKLITKVVVNRI-RPL 987
+ E+ + +L++L+ +D S +RPI L + + K++ ++V R+ + L
Sbjct: 449 KQCLLESYFPQKWKIASLVILLKLLDRIRSDPGSYRPICLLDNLGKVLEGIMVKRLDQKL 508
Query: 988 LNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGASMAKKGTLAFKIDLEKAYDSVSW 1044
++ + P Q +F G D Q H+ S K + ID + A+D++ W
Sbjct: 509 MDVEVSPYQFAFTYGKSTEDAWRCVQR---HVECSEMKY-VIGLNIDFQGAFDNLGW 561
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 40.0 bits (92), Expect = 0.059
Identities = 20/70 (28%), Positives = 37/70 (52%)
Query: 1033 IDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKPGRGLR 1092
+D+ KA+D+VS + + FG + + IM ++ + +++ G G++
Sbjct: 28 LDIRKAFDTVSHPAILRAMRAFGIDDGMQDFIMSTITDAYTNIVVGGRTTNKIYIRNGVK 87
Query: 1093 KGDPLSPYLF 1102
+GDPLSP LF
Sbjct: 88 QGDPLSPVLF 97
>Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I
retrotransposable element R1DM (ORF 2)
Length = 1021
Score = 37.7 bits (86), Expect = 0.29
Identities = 46/188 (24%), Positives = 77/188 (40%), Gaps = 12/188 (6%)
Query: 875 LTEAERHKAEAEVIKE--EVFQA---LLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKL 929
+ E+E A AE + EVF+ + R++S + G DG + K W + + L L
Sbjct: 417 VAESEAPTAIAEEVPPALEVFEVDTCVARLKSRRSPGLDGINGTICKAVWRAIPEHLASL 476
Query: 930 VRDAFETGMIDDSLLETLIVLIPKVDSPSTVK--EFRPISLCNVVYKLITKVVVNRIRPL 987
G +V + K + +R I L V K++ ++VNR+R +
Sbjct: 477 FSRCIRLGYFPAEWKCPRVVSLLKGPDKDKCEPSSYRGICLLPVFGKVLEAIMVNRVREV 536
Query: 988 LNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGASMAKKGTLAFKIDLEKAYDSVSWAFL 1047
L Q F G + D + + +GAS A+ L +D + A+D+V W+
Sbjct: 537 LPEGC-RWQFGFRQGRCVED---AWRHVKSSVGASAAQY-VLGTFVDFKGAFDNVEWSAA 591
Query: 1048 EDTLVQFG 1055
L G
Sbjct: 592 LSRLADLG 599
>PLE1_HUMAN (Q15149) Plectin 1 (PLTN) (PCN) (Hemidesmosomal protein
1) (HD1)
Length = 4684
Score = 34.3 bits (77), Expect = 3.2
Identities = 35/125 (28%), Positives = 54/125 (43%), Gaps = 17/125 (13%)
Query: 701 LASWVEHPQYKEVVQSAWTGETETVAGKLNKVRTKSCKFNKEVYGSIFQRKRWVEGRL-- 758
L +WVE Q++ V + W + +V +L R + E + + +R R EG+L
Sbjct: 665 LLAWVEENQHR-VDGAEWGVDLPSVEAQLGSHR--GLHQSIEEFQAKIERARSDEGQLSP 721
Query: 759 --RGVQRELNYRVTFDMTRLETELQAEYRFL--------LRQEELLWFQKARENRVCFD- 807
RG R+ R+ +L +A R L +EL+W + E V FD
Sbjct: 722 ATRGAYRDCLGRLDLQYAKLLNSSKARLRSLESLHSFVAAATKELMWLNEKEEEEVGFDW 781
Query: 808 -DRNT 811
DRNT
Sbjct: 782 SDRNT 786
>IDI2_THEAC (Q9HLX2) Isopentenyl-diphosphate delta-isomerase (EC
5.3.3.2) (IPP isomerase) (Isopentenyl pyrophosphate
isomerase)
Length = 348
Score = 33.9 bits (76), Expect = 4.2
Identities = 22/74 (29%), Positives = 35/74 (46%), Gaps = 2/74 (2%)
Query: 625 MVDLGMSGGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHR-IHSDHCPLL 683
M+ M+GG N + ++R MG G+ R A + S+E + I+ H PL
Sbjct: 58 MIISSMTGGAEIAKNINRNLAVAAERFGIGMGVGSMRAAIVDRSIEDTYSVINESHVPLK 117
Query: 684 I-NCSGPEIVHHDR 696
I N P++V D+
Sbjct: 118 IANIGAPQLVRQDK 131
>RTP2_TRYBG (P15594) Retrotransposable element SLACS 132 kDa protein
(ORF2)
Length = 1182
Score = 33.1 bits (74), Expect = 7.2
Identities = 38/149 (25%), Positives = 68/149 (45%), Gaps = 15/149 (10%)
Query: 962 EFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGA 1021
++RPI +V KL + + ++R+ + +Q G+ + +E I +
Sbjct: 585 KYRPIGAESVWAKLASHIAISRVMKTAEKKFSGIQ------FGVGGHI---EEAIAKIRK 635
Query: 1022 SMAKKGTLAFKIDLEKAYDSVSW-AFLEDTLVQFGFPN--RLVQLIMRCVSGSNLSMLWN 1078
A KG+LA +D AY+++S A LE + RLV L++ + N
Sbjct: 636 DFATKGSLAM-LDGRNAYNAISRRAILEAVYGDSTWSPLWRLVSLLLGTTG--EVGFYEN 692
Query: 1079 GSRLPSFKPGRGLRKGDPLSPYLFVLCSL 1107
G +++ RG+R+G L P LF + +L
Sbjct: 693 GKLCHTWESTRGVRQGMVLGPLLFSIGTL 721
>ARFP_ARATH (Q93YR9) Auxin response factor 16
Length = 670
Score = 33.1 bits (74), Expect = 7.2
Identities = 20/47 (42%), Positives = 25/47 (52%), Gaps = 7/47 (14%)
Query: 1603 VLDPLPWTLDGAWQKLRHDHDELLQFSSGDLLDEHHFLINLWKPPPP 1649
VLD +P L GA RH+ + SS DL HH+ +N PPPP
Sbjct: 437 VLDNVPVGLQGA----RHNAHQYYGLSSSDL---HHYYLNRPPPPPP 476
>ALBU_PETMA (Q91274) Serum albumin SDS-1 precursor
Length = 1423
Score = 32.7 bits (73), Expect = 9.4
Identities = 28/92 (30%), Positives = 39/92 (41%), Gaps = 10/92 (10%)
Query: 1686 SHTSIGNPFLAEVLALRDGLCL--AWVQGHRRIIREVDCAGLSDILRFEDRCRLHDHAGV 1743
SH G+ FL E AL D +C AW+ G + R G + IL F++ L H
Sbjct: 486 SHRKNGSCFLEERYALHDAICRDEAWLSGLAEVSRCCAMDGRARILCFDE---LSSHLNA 542
Query: 1744 LQEIRELMERPWRCSVHWTNRDSNASADFLAR 1775
E ERP CS ++ + +F R
Sbjct: 543 SVE-----ERPELCSTSLCSKYHDLGFEFKQR 569
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.333 0.144 0.485
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 214,626,845
Number of Sequences: 164201
Number of extensions: 9139828
Number of successful extensions: 20970
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 20928
Number of HSP's gapped (non-prelim): 33
length of query: 1813
length of database: 59,974,054
effective HSP length: 125
effective length of query: 1688
effective length of database: 39,448,929
effective search space: 66589792152
effective search space used: 66589792152
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0178.7


