

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.5
(770 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
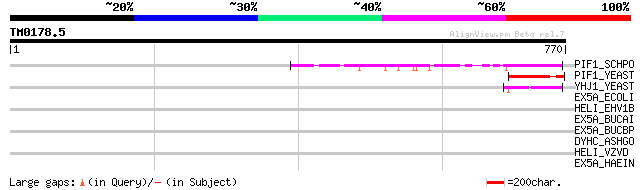
PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1, m... 63 3e-09
PIF1_YEAST (P07271) DNA repair and recombination protein PIF1, m... 54 2e-06
YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergeni... 50 2e-05
EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 37 0.20
HELI_EHV1B (P28934) Probable helicase 37 0.26
EX5A_BUCAI (P57530) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 35 0.98
EX5A_BUCBP (Q89AB2) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 34 1.3
DYHC_ASHGO (Q9C1M7) Dynein heavy chain, cytosolic (DYHC) 33 2.2
HELI_VZVD (P09303) Probable helicase 33 2.8
EX5A_HAEIN (P45158) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 32 8.3
>PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1,
mitochondrial precursor
Length = 805
Score = 62.8 bits (151), Expect = 3e-09
Identities = 94/411 (22%), Positives = 173/411 (41%), Gaps = 72/411 (17%)
Query: 390 KASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVIS 449
+ ++I DE M++ + L+ I K PFGGI +VL GDF Q+ PV
Sbjct: 410 RTRVLIIDEVSMVDAELMDKLEEVARVIRKDSK------PFGGIQLVLTGDFFQLPPVPE 463
Query: 450 KGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNA----TSTSSPAEIKEFADWLLQVGD 505
G S+ S WK + + + + + + + +D ++
Sbjct: 464 NGKESKFCFE----SQTWKSALDFTIGLTHVFRQKDEEFVKMLNELRLGKLSDESVRKFK 519
Query: 506 GTVKTIDEEETLIE---IPPNLLIEQCKEPLLELVN---FAYPKLAHNLQKNSFFQERAI 559
+TI+ E+ L+ P +E+ + ++ +N + + ++ F++R +
Sbjct: 520 VLNRTIEYEDGLLPTELFPTRYEVERSNDMRMQQINQNPVTFTAIDSGTVRDKEFRDRLL 579
Query: 560 ---LAPT------------LESVEE-INNFMLAMIPG---DETEYLSYDTLCKSDEDSGV 600
+AP ++++++ + N L + G DET Y K E G
Sbjct: 580 QGCMAPATLVLKVNAQVMLIKNIDDQLVNGSLGKVIGFIDDET----YQMEKKDAEMQGR 635
Query: 601 NAEWFTSEFLNDFKCSEIPNHAIKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIV 660
NA + S ++ F ++ KL + +R A K+ +V
Sbjct: 636 NAFEYDSLDISPFDLPDVKQKKYKL------IAMRKASSTA-------------IKWPLV 676
Query: 661 ATIL-NGNKMGETTFIPRMSLTPSNSDIP---FKFQRRQFPVTLCFAMTINKSQGQSLSH 716
L NG GE T + + N ++P + R Q P+ L +A++I+K+QGQ+L
Sbjct: 677 RFKLPNG---GERTIVVQRETW--NIELPNGEVQASRSQIPLILAYAISIHKAQGQTLDR 731
Query: 717 VGLYLPRPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQEV 767
V + L R VF GQ YVALSR +++GL++L V+++ Y+++
Sbjct: 732 VKVDLGR-VFEKGQAYVALSRATTQEGLQVLNFSPAKVMAHPKVVQFYKQL 781
>PIF1_YEAST (P07271) DNA repair and recombination protein PIF1,
mitochondrial precursor
Length = 857
Score = 53.5 bits (127), Expect = 2e-06
Identities = 35/77 (45%), Positives = 48/77 (61%), Gaps = 8/77 (10%)
Query: 693 RRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLIIDDE 752
R Q P+ L ++++I+KSQGQ+L V + L R VF GQ YVALSR SR+GL++L D
Sbjct: 691 RVQLPLMLAWSLSIHKSQGQTLPKVKVDLRR-VFEKGQAYVALSRAVSREGLQVLNFD-- 747
Query: 753 GVVSNTTRNVMYQEVFD 769
TR +Q+V D
Sbjct: 748 -----RTRIKAHQKVID 759
Score = 40.0 bits (92), Expect = 0.023
Identities = 54/245 (22%), Positives = 87/245 (35%), Gaps = 54/245 (22%)
Query: 394 IIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVISKGSR 453
++ DE ML+ + LD I K PFGGI ++ GDF Q+ PV +R
Sbjct: 338 LVVDEISMLDAELLDKLDFIARKIRKNHQ------PFGGIQLIFCGDFFQLPPVSKDPNR 391
Query: 454 SEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIKEFADWLLQVGDGTVKTIDE 513
S WK M + + + + +F + L ++ G + E
Sbjct: 392 PT---KFAFESKAWKEGVKMTIMLQKVFRQRGDV-------KFIEMLNRMRLGNIDDETE 441
Query: 514 EETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTLESVEEINNF 573
E + KL+ L + A L T VE NN
Sbjct: 442 RE-------------------------FKKLSRPLPDDEIIP--AELYSTRMEVERANNS 474
Query: 574 MLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVGVPIML 633
L+ +PG + + D DE+ L +F + + LKVG +M+
Sbjct: 475 RLSKLPGQVHIFNAIDGGALEDEE-------LKERLLQNF----LAPKELHLKVGAQVMM 523
Query: 634 IRNID 638
++N+D
Sbjct: 524 VKNLD 528
>YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergenic
region
Length = 723
Score = 50.4 bits (119), Expect = 2e-05
Identities = 30/85 (35%), Positives = 51/85 (59%), Gaps = 4/85 (4%)
Query: 686 DIPFK---FQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRK 742
DIP + +R Q P+ LC+A++I+K+QGQ++ + + L R +F GQ+YVALSR +
Sbjct: 636 DIPRENVGLERTQIPLMLCWALSIHKAQGQTIQRLKVDL-RRIFEAGQVYVALSRAVTMD 694
Query: 743 GLKMLIIDDEGVVSNTTRNVMYQEV 767
L++L D + +N Y+ +
Sbjct: 695 TLQVLNFDPGKIRTNERVKDFYKRL 719
Score = 40.0 bits (92), Expect = 0.023
Identities = 27/78 (34%), Positives = 39/78 (49%), Gaps = 8/78 (10%)
Query: 393 LIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVISKGS 452
++I DE M++ + + L++ I K D PFGGI +VL GDF Q+ PV K
Sbjct: 333 VLIIDEISMVDGNLLDKLEQIARRIRKN------DDPFGGIQLVLTGDFFQLPPVAKKDE 386
Query: 453 RSEIVGSAINSSYLWKHC 470
+ V S +WK C
Sbjct: 387 HN--VVKFCFESEMWKRC 402
>EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5) (Exodeoxyribonuclease V 67 kDa polypeptide)
Length = 608
Score = 37.0 bits (84), Expect = 0.20
Identities = 19/46 (41%), Positives = 26/46 (56%), Gaps = 3/46 (6%)
Query: 702 FAMTINKSQGQSLSHVGLYLP---RPVFTHGQLYVALSRVKSRKGL 744
+AMT++KSQG H L LP PV T +Y A++R + R L
Sbjct: 534 WAMTVHKSQGSEFDHAALILPSQRTPVVTRELVYTAVTRARRRLSL 579
>HELI_EHV1B (P28934) Probable helicase
Length = 881
Score = 36.6 bits (83), Expect = 0.26
Identities = 20/44 (45%), Positives = 26/44 (58%)
Query: 703 AMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKM 746
AMTI +SQG SL V + PR +YVA+SR S + L+M
Sbjct: 806 AMTIARSQGLSLDKVAICFPRNNLRINSVYVAMSRTVSSRFLRM 849
>EX5A_BUCAI (P57530) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5)
Length = 602
Score = 34.7 bits (78), Expect = 0.98
Identities = 37/135 (27%), Positives = 57/135 (41%), Gaps = 25/135 (18%)
Query: 627 VGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSD 686
+G PIM+I N ++A + N I N I+ + L N + ++
Sbjct: 480 IGKPIMIINN-NRALNVSNGNIGITNINKNGILQVSFLKENN--------------TINN 524
Query: 687 IPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPR---PVFTHGQLYVALSRVKSRKG 743
IP K R +A+T++KSQG + L LP + LY ++R SRK
Sbjct: 525 IPVKILRNY---KTAWAITVHKSQGSEFMNTALILPNFNSHILNKDTLYTGITR--SRKI 579
Query: 744 LKMLIIDDEGVVSNT 758
L I D+ + NT
Sbjct: 580 LS--IFSDKKIFLNT 592
>EX5A_BUCBP (Q89AB2) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5)
Length = 618
Score = 34.3 bits (77), Expect = 1.3
Identities = 29/107 (27%), Positives = 50/107 (46%), Gaps = 15/107 (14%)
Query: 662 TILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQFPVTLC-FAMTINKSQGQSLSHVGLY 720
T+L+ NK + F+P+ N + + P + MT++KSQG S V L
Sbjct: 509 TLLDANKKLKVFFLPK------NQEQIYSIPIHLVPEHQTNWTMTVHKSQGSEFSEVVLI 562
Query: 721 LP---RPVFTHGQLYVALSRVKSRKGLKMLIIDDEGV-VSNTTRNVM 763
LP + T +Y A++R K K+ I DE + + + +N++
Sbjct: 563 LPTIMTSILTKELIYTAVTRSKK----KLTIYSDENIFIKSLKKNII 605
>DYHC_ASHGO (Q9C1M7) Dynein heavy chain, cytosolic (DYHC)
Length = 4083
Score = 33.5 bits (75), Expect = 2.2
Identities = 20/74 (27%), Positives = 39/74 (52%), Gaps = 7/74 (9%)
Query: 648 RMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAMTIN 707
+M++ T+ + I++ + + ET+F+ RM+ +NSD+P F+ ++ L
Sbjct: 2795 QMLLRCGTESEKICLIIDESNILETSFLERMNTLLANSDVPGLFEADEYEALL------- 2847
Query: 708 KSQGQSLSHVGLYL 721
GQ +S +GL L
Sbjct: 2848 SKIGQRISQLGLLL 2861
>HELI_VZVD (P09303) Probable helicase
Length = 881
Score = 33.1 bits (74), Expect = 2.8
Identities = 19/44 (43%), Positives = 24/44 (54%)
Query: 703 AMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKM 746
AMTI +SQG SL V + +YVA+SR S + LKM
Sbjct: 806 AMTIARSQGLSLEKVAICFTADKLRLNSVYVAMSRTVSSRFLKM 849
>EX5A_HAEIN (P45158) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5)
Length = 640
Score = 31.6 bits (70), Expect = 8.3
Identities = 18/68 (26%), Positives = 32/68 (46%), Gaps = 7/68 (10%)
Query: 702 FAMTINKSQGQSLSHVGLYLP---RPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVVSNT 758
F MTI+KSQG H + LP PV + ++ ++R K ++ + DE +
Sbjct: 566 FMMTIHKSQGSEFKHTVMVLPTEVNPVLSRELVFTGVTRAKK----ELTVFADEKIWKTA 621
Query: 759 TRNVMYQE 766
R + ++
Sbjct: 622 IRQTVKRQ 629
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.347 0.154 0.513
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 78,062,328
Number of Sequences: 164201
Number of extensions: 3080349
Number of successful extensions: 12417
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 12403
Number of HSP's gapped (non-prelim): 15
length of query: 770
length of database: 59,974,054
effective HSP length: 118
effective length of query: 652
effective length of database: 40,598,336
effective search space: 26470115072
effective search space used: 26470115072
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0178.5


