

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.13
(433 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
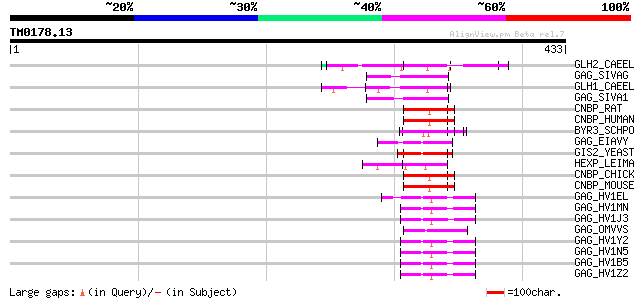
Score E
Sequences producing significant alignments: (bits) Value
GLH2_CAEEL (Q966L9) ATP-dependent RNA helicase glh-2 (EC 3.6.1.-... 54 8e-07
GAG_SIVAG (P27978) Gag polyprotein [Contains: Core protein p17; ... 52 4e-06
GLH1_CAEEL (P34689) ATP-dependent RNA helicase glh-1 (EC 3.6.1.-... 50 1e-05
GAG_SIVA1 (P27972) Gag polyprotein [Contains: Core protein p17; ... 49 3e-05
CNBP_RAT (P62634) Cellular nucleic acid binding protein (CNBP) (... 49 3e-05
CNBP_HUMAN (P62633) Cellular nucleic acid binding protein (CNBP)... 49 3e-05
BYR3_SCHPO (P36627) Cellular nucleic acid binding protein homolog 49 3e-05
GAG_EIAVY (P03351) Gag polyprotein [Contains: Core protein p15; ... 48 4e-05
GIS2_YEAST (P53849) Zinc-finger protein GIS2 48 6e-05
HEXP_LEIMA (Q04832) DNA-binding protein HEXBP (Hexamer-binding p... 47 1e-04
CNBP_CHICK (O42395) Cellular nucleic acid binding protein (CNBP) 47 1e-04
CNBP_MOUSE (P53996) Cellular nucleic acid binding protein (CNBP)... 46 2e-04
GAG_HV1EL (P04592) Gag polyprotein [Contains: Core protein p17 (... 46 2e-04
GAG_HV1MN (P05888) Gag polyprotein [Contains: Core protein p17 (... 45 3e-04
GAG_HV1J3 (P12494) Gag polyprotein [Contains: Core protein p17 (... 45 3e-04
GAG_OMVVS (P16900) Gag polyprotein [Contains: Core protein p16; ... 45 4e-04
GAG_HV1Y2 (P35962) Gag polyprotein [Contains: Core protein p17 (... 45 4e-04
GAG_HV1N5 (P12493) Gag polyprotein [Contains: Core protein p17 (... 45 4e-04
GAG_HV1B5 (P04593) Gag polyprotein [Contains: Core protein p17 (... 45 4e-04
GAG_HV1Z2 (P12495) Gag polyprotein [Contains: Core protein p17 (... 45 5e-04
>GLH2_CAEEL (Q966L9) ATP-dependent RNA helicase glh-2 (EC 3.6.1.-)
(Germline helicase-2)
Length = 974
Score = 53.9 bits (128), Expect = 8e-07
Identities = 48/150 (32%), Positives = 64/150 (42%), Gaps = 24/150 (16%)
Query: 248 NQGNG-GSAMTN---QGRNDKGRHYQKKPYVRPQGQGTASGSFYPTGGNAIAPRPLAVSL 303
N G+G GS +TN G N + K G G + GS + +GGN+
Sbjct: 193 NSGSGFGSGLTNGFGSGNNHESGFGGGKS--GGFGGGNSGGSGFGSGGNSNGFGSGGGGQ 250
Query: 304 D----GVTCFKCNQKGHYASHCPEG-----PLMCWNCNKPGHLARDC---RIPKVEDAAN 351
D CF C Q GH ++ CPE P +C+NC +PGH +RDC R P+ E
Sbjct: 251 DRGERNNNCFNCQQPGHRSNDCPEPKKEREPRVCYNCQQPGHNSRDCPEERKPR-EGRNG 309
Query: 352 VAGARRPTAGGRVYFISGTEVDEDEGLIHG 381
G GG +G D GL +G
Sbjct: 310 FTGGSSGFGGG-----NGGGTGFDSGLTNG 334
Score = 53.5 bits (127), Expect = 1e-06
Identities = 32/87 (36%), Positives = 39/87 (44%), Gaps = 17/87 (19%)
Query: 308 CFKCNQKGHYASHCPEG-----PLMCWNCNKPGHLARDCRIPKVEDAANVAGARRPTAGG 362
CF C Q GH ++ CPE P +C+NC +PGH +RDC R G
Sbjct: 373 CFNCQQPGHRSNDCPEPKKEREPRVCYNCQQPGHNSRDC------------PEERKPREG 420
Query: 363 RVYFISGTEVDEDEGLIHGNREIPGNS 389
R F SG D G GN E GN+
Sbjct: 421 RNGFTSGFGGGNDGGFGGGNAEGFGNN 447
Score = 49.3 bits (116), Expect = 2e-05
Identities = 31/101 (30%), Positives = 41/101 (39%), Gaps = 23/101 (22%)
Query: 244 QNKPNQGNGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGSFYPTGGNAIAPRPLAVSL 303
+ KP +G G G ND G G G A G GN P+
Sbjct: 414 ERKPREGRNGFTSGFGGGNDGGF-----------GGGNAEGF-----GNNEERGPMK--- 454
Query: 304 DGVTCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIP 344
CF C +GH ++ CPE P C+NC + GH + +C P
Sbjct: 455 ----CFNCKGEGHRSAECPEPPRGCFNCGEQGHRSNECPNP 491
>GAG_SIVAG (P27978) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 521
Score = 51.6 bits (122), Expect = 4e-06
Identities = 27/65 (41%), Positives = 31/65 (47%), Gaps = 7/65 (10%)
Query: 279 QGTASGSFYPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEGPLM-CWNCNKPGHL 337
Q S + GG PRP V C+ C + GH CPE M C C KPGHL
Sbjct: 381 QNMQSQNMMQQGGQRGRPRP------PVKCYNCGKFGHMQRQCPEPRKMRCLKCGKPGHL 434
Query: 338 ARDCR 342
A+DCR
Sbjct: 435 AKDCR 439
Score = 33.1 bits (74), Expect = 1.4
Identities = 16/56 (28%), Positives = 26/56 (45%), Gaps = 1/56 (1%)
Query: 290 GGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPK 345
GG + + +A + + Q+G P P+ C+NC K GH+ R C P+
Sbjct: 367 GGPSYKAKVMAEMMQNMQSQNMMQQGGQRGR-PRPPVKCYNCGKFGHMQRQCPEPR 421
>GLH1_CAEEL (P34689) ATP-dependent RNA helicase glh-1 (EC 3.6.1.-)
(Germline helicase-1)
Length = 763
Score = 50.1 bits (118), Expect = 1e-05
Identities = 27/75 (36%), Positives = 36/75 (48%), Gaps = 18/75 (24%)
Query: 278 GQGTASGSF------YPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEG-----PL 326
G G+ GSF + GG+ R CF C Q GH +S CPE P
Sbjct: 131 GFGSGGGSFGGGNSGFGEGGHGGGER-------NNNCFNCQQPGHRSSDCPEPRKEREPR 183
Query: 327 MCWNCNKPGHLARDC 341
+C+NC +PGH +R+C
Sbjct: 184 VCYNCQQPGHTSREC 198
Score = 47.8 bits (112), Expect = 6e-05
Identities = 32/106 (30%), Positives = 43/106 (40%), Gaps = 31/106 (29%)
Query: 244 QNKPNQGN-----GGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGSFYPTGGNAIAPRP 298
+ KP +G GG+ N G ND G G G F GG P
Sbjct: 201 ERKPREGRTGGFGGGAGFGNNGGND--------------GFG-GDGGF--GGGEERGP-- 241
Query: 299 LAVSLDGVTCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIP 344
+ CF C +GH ++ CPE P C+NC + GH + +C P
Sbjct: 242 -------MKCFNCKGEGHRSAECPEPPRGCFNCGEQGHRSNECPNP 280
Score = 30.4 bits (67), Expect = 9.2
Identities = 10/20 (50%), Positives = 14/20 (70%)
Query: 328 CWNCNKPGHLARDCRIPKVE 347
C+NC +PGH + DC P+ E
Sbjct: 160 CFNCQQPGHRSSDCPEPRKE 179
>GAG_SIVA1 (P27972) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 520
Score = 48.9 bits (115), Expect = 3e-05
Identities = 25/65 (38%), Positives = 30/65 (45%), Gaps = 7/65 (10%)
Query: 279 QGTASGSFYPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEG-PLMCWNCNKPGHL 337
Q S + GG PRP C+ C + GH CPE + C C KPGHL
Sbjct: 377 QNLQSQNMVQQGGGRGRPRPPP------KCYNCGKFGHMQRQCPEPRKIKCLKCGKPGHL 430
Query: 338 ARDCR 342
A+DCR
Sbjct: 431 AKDCR 435
Score = 30.8 bits (68), Expect = 7.1
Identities = 11/24 (45%), Positives = 14/24 (57%)
Query: 322 PEGPLMCWNCNKPGHLARDCRIPK 345
P P C+NC K GH+ R C P+
Sbjct: 394 PRPPPKCYNCGKFGHMQRQCPEPR 417
>CNBP_RAT (P62634) Cellular nucleic acid binding protein (CNBP)
(Zinc finger protein 9)
Length = 177
Score = 48.5 bits (114), Expect = 3e-05
Identities = 17/40 (42%), Positives = 25/40 (62%)
Query: 308 CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKVE 347
C++C + GH A C C+NC + GH+A+DC+ PK E
Sbjct: 54 CYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPKRE 93
Score = 46.2 bits (108), Expect = 2e-04
Identities = 19/38 (50%), Positives = 22/38 (57%), Gaps = 4/38 (10%)
Query: 308 CFKCNQKGHYASHCPEGPL----MCWNCNKPGHLARDC 341
C+ C + GH A C E C+NC KPGHLARDC
Sbjct: 74 CYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 111
Score = 42.7 bits (99), Expect = 0.002
Identities = 16/39 (41%), Positives = 25/39 (64%), Gaps = 1/39 (2%)
Query: 306 VTCFKCNQKGHYASHCPE-GPLMCWNCNKPGHLARDCRI 343
V C++C + GH A +C + + C+ C + GHLAR+C I
Sbjct: 135 VKCYRCGETGHVAINCSKTSEVNCYRCGESGHLARECTI 173
Score = 41.6 bits (96), Expect = 0.004
Identities = 23/75 (30%), Positives = 30/75 (39%), Gaps = 30/75 (40%)
Query: 308 CFKCNQKGHYASHCPEG----------------------------PLMCWNCNKPGHLAR 339
CFKC + GH+A CP G P +C+ C + GHLA+
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGRGGFTSDRGFQFVSSSLPDICYRCGESGHLAK 65
Query: 340 DCRIPKVEDAANVAG 354
DC + EDA G
Sbjct: 66 DCDLQ--EDACYNCG 78
Score = 34.7 bits (78), Expect = 0.49
Identities = 12/35 (34%), Positives = 19/35 (54%), Gaps = 1/35 (2%)
Query: 308 CFKCNQKGHYASHCPEGPLM-CWNCNKPGHLARDC 341
C+ C + GH A C C++C + GH+ +DC
Sbjct: 98 CYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDC 132
Score = 33.1 bits (74), Expect = 1.4
Identities = 12/38 (31%), Positives = 20/38 (52%), Gaps = 2/38 (5%)
Query: 304 DGVTCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDC 341
D C+ C + GH C + + C+ C + GH+A +C
Sbjct: 115 DEQKCYSCGEFGHIQKDCTK--VKCYRCGETGHVAINC 150
>CNBP_HUMAN (P62633) Cellular nucleic acid binding protein (CNBP)
(Zinc finger protein 9)
Length = 177
Score = 48.5 bits (114), Expect = 3e-05
Identities = 17/40 (42%), Positives = 25/40 (62%)
Query: 308 CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKVE 347
C++C + GH A C C+NC + GH+A+DC+ PK E
Sbjct: 54 CYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPKRE 93
Score = 46.2 bits (108), Expect = 2e-04
Identities = 19/38 (50%), Positives = 22/38 (57%), Gaps = 4/38 (10%)
Query: 308 CFKCNQKGHYASHCPEGPL----MCWNCNKPGHLARDC 341
C+ C + GH A C E C+NC KPGHLARDC
Sbjct: 74 CYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 111
Score = 42.7 bits (99), Expect = 0.002
Identities = 16/39 (41%), Positives = 25/39 (64%), Gaps = 1/39 (2%)
Query: 306 VTCFKCNQKGHYASHCPE-GPLMCWNCNKPGHLARDCRI 343
V C++C + GH A +C + + C+ C + GHLAR+C I
Sbjct: 135 VKCYRCGETGHVAINCSKTSEVNCYRCGESGHLARECTI 173
Score = 41.6 bits (96), Expect = 0.004
Identities = 23/75 (30%), Positives = 30/75 (39%), Gaps = 30/75 (40%)
Query: 308 CFKCNQKGHYASHCPEG----------------------------PLMCWNCNKPGHLAR 339
CFKC + GH+A CP G P +C+ C + GHLA+
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGRGGFTSDRGFQFVSSSLPDICYRCGESGHLAK 65
Query: 340 DCRIPKVEDAANVAG 354
DC + EDA G
Sbjct: 66 DCDLQ--EDACYNCG 78
Score = 34.7 bits (78), Expect = 0.49
Identities = 12/35 (34%), Positives = 19/35 (54%), Gaps = 1/35 (2%)
Query: 308 CFKCNQKGHYASHCPEGPLM-CWNCNKPGHLARDC 341
C+ C + GH A C C++C + GH+ +DC
Sbjct: 98 CYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDC 132
Score = 33.1 bits (74), Expect = 1.4
Identities = 12/38 (31%), Positives = 20/38 (52%), Gaps = 2/38 (5%)
Query: 304 DGVTCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDC 341
D C+ C + GH C + + C+ C + GH+A +C
Sbjct: 115 DEQKCYSCGEFGHIQKDCTK--VKCYRCGETGHVAINC 150
>BYR3_SCHPO (P36627) Cellular nucleic acid binding protein homolog
Length = 179
Score = 48.5 bits (114), Expect = 3e-05
Identities = 19/50 (38%), Positives = 27/50 (54%), Gaps = 1/50 (2%)
Query: 305 GVTCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKVEDAANVAG 354
G C+ C + GH A C +G + C+NCN+ GH A +C P+ E G
Sbjct: 16 GPRCYNCGENGHQARECTKGSI-CYNCNQTGHKASECTEPQQEKTCYACG 64
Score = 45.8 bits (107), Expect = 2e-04
Identities = 19/39 (48%), Positives = 22/39 (55%), Gaps = 2/39 (5%)
Query: 305 GVTCFKCNQKGHYASHC--PEGPLMCWNCNKPGHLARDC 341
G C+ CNQ GH AS C P+ C+ C GHL RDC
Sbjct: 35 GSICYNCNQTGHKASECTEPQQEKTCYACGTAGHLVRDC 73
Score = 45.4 bits (106), Expect = 3e-04
Identities = 17/45 (37%), Positives = 27/45 (59%), Gaps = 2/45 (4%)
Query: 305 GVTCFKCNQKGHYASHCPEGP--LMCWNCNKPGHLARDCRIPKVE 347
GV C+ C + GH + C + +C+ CN+PGH+A +C P +E
Sbjct: 134 GVKCYSCGKIGHRSFECQQASDGQLCYKCNQPGHIAVNCTSPVIE 178
Score = 45.4 bits (106), Expect = 3e-04
Identities = 19/55 (34%), Positives = 24/55 (43%), Gaps = 5/55 (9%)
Query: 307 TCFKCNQKGHYASHCPEGP-----LMCWNCNKPGHLARDCRIPKVEDAANVAGAR 356
TC+ C GH CP P C+ C + GH+ARDCR + G R
Sbjct: 59 TCYACGTAGHLVRDCPSSPNPRQGAECYKCGRVGHIARDCRTNGQQSGGRFGGHR 113
Score = 34.3 bits (77), Expect = 0.64
Identities = 16/50 (32%), Positives = 20/50 (40%), Gaps = 13/50 (26%)
Query: 305 GVTCFKCNQKGHYASHC-------------PEGPLMCWNCNKPGHLARDC 341
G C+KC + GH A C + C+ C GH ARDC
Sbjct: 82 GAECYKCGRVGHIARDCRTNGQQSGGRFGGHRSNMNCYACGSYGHQARDC 131
Score = 31.6 bits (70), Expect = 4.2
Identities = 11/18 (61%), Positives = 13/18 (72%)
Query: 304 DGVTCFKCNQKGHYASHC 321
DG C+KCNQ GH A +C
Sbjct: 155 DGQLCYKCNQPGHIAVNC 172
>GAG_EIAVY (P03351) Gag polyprotein [Contains: Core protein p15;
Major core protein p26 (Capsid protein p26); Core
protein p11; Core protein p9]
Length = 486
Score = 48.1 bits (113), Expect = 4e-05
Identities = 22/59 (37%), Positives = 33/59 (55%), Gaps = 5/59 (8%)
Query: 288 PTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCR-IPK 345
P G A+ PL + TC+ C + GH +S C P +C+ C +PGH ++ CR +PK
Sbjct: 366 PFKGGALKGGPLKAAQ---TCYNCGKPGHLSSQC-RAPKVCFKCKQPGHFSKQCRSVPK 420
Score = 37.7 bits (86), Expect = 0.058
Identities = 16/27 (59%), Positives = 18/27 (66%), Gaps = 4/27 (14%)
Query: 324 GPLM----CWNCNKPGHLARDCRIPKV 346
GPL C+NC KPGHL+ CR PKV
Sbjct: 375 GPLKAAQTCYNCGKPGHLSSQCRAPKV 401
>GIS2_YEAST (P53849) Zinc-finger protein GIS2
Length = 153
Score = 47.8 bits (112), Expect = 6e-05
Identities = 17/38 (44%), Positives = 25/38 (65%), Gaps = 1/38 (2%)
Query: 308 CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPK 345
C+ C + GH A C + +C+NCNKPGH+ DC +P+
Sbjct: 6 CYVCGKIGHLAEDC-DSERLCYNCNKPGHVQTDCTMPR 42
Score = 46.2 bits (108), Expect = 2e-04
Identities = 15/39 (38%), Positives = 26/39 (66%), Gaps = 1/39 (2%)
Query: 303 LDGVTCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDC 341
+ G+ C+ C Q GH + C + +C+NCN+ GH+++DC
Sbjct: 113 ISGLKCYTCGQAGHMSRDC-QNDRLCYNCNETGHISKDC 150
Score = 42.7 bits (99), Expect = 0.002
Identities = 16/38 (42%), Positives = 23/38 (60%), Gaps = 2/38 (5%)
Query: 308 CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPK 345
C+ C + GH S C C+NCN+ GH++R+C PK
Sbjct: 49 CYNCGETGHVRSECTVQ--RCFNCNQTGHISRECPEPK 84
Score = 39.7 bits (91), Expect = 0.015
Identities = 18/52 (34%), Positives = 26/52 (49%), Gaps = 5/52 (9%)
Query: 296 PRPLAVS-LDGVTCFKCNQKGHYASHCPE----GPLMCWNCNKPGHLARDCR 342
P P S V+C+KC H A C + L C+ C + GH++RDC+
Sbjct: 81 PEPKKTSRFSKVSCYKCGGPNHMAKDCMKEDGISGLKCYTCGQAGHMSRDCQ 132
>HEXP_LEIMA (Q04832) DNA-binding protein HEXBP (Hexamer-binding
protein)
Length = 271
Score = 46.6 bits (109), Expect = 1e-04
Identities = 26/82 (31%), Positives = 35/82 (41%), Gaps = 16/82 (19%)
Query: 276 PQGQGTASGS----FYPTGGNAIAPRPLAVSLDGVT------CFKCNQKGHYASHCPE-- 323
P GQG SG+ Y G R G + C+KC + GH + CP
Sbjct: 156 PNGQGGYSGAGDRTCYKCGDAGHISRDCPNGQGGYSGAGDRKCYKCGESGHMSRECPSAG 215
Query: 324 ----GPLMCWNCNKPGHLARDC 341
G C+ C KPGH++R+C
Sbjct: 216 STGSGDRACYKCGKPGHISREC 237
Score = 45.8 bits (107), Expect = 2e-04
Identities = 16/42 (38%), Positives = 25/42 (59%), Gaps = 7/42 (16%)
Query: 307 TCFKCNQKGHYASHCPE-------GPLMCWNCNKPGHLARDC 341
TCF+C ++GH + CP G + C+ C + GH++RDC
Sbjct: 44 TCFRCGEEGHMSRECPNEARSGAAGAMTCFRCGEAGHMSRDC 85
Score = 43.1 bits (100), Expect = 0.001
Identities = 28/85 (32%), Positives = 37/85 (42%), Gaps = 19/85 (22%)
Query: 276 PQGQGTASGS----FYPTGGNAIAPR--PLAVSLDGV--TCFKCNQKGHYASHCPE---- 323
P GQG SG+ Y G + R P A S C+KC + GH + CPE
Sbjct: 184 PNGQGGYSGAGDRKCYKCGESGHMSRECPSAGSTGSGDRACYKCGKPGHISRECPEAGGS 243
Query: 324 -------GPLMCWNCNKPGHLARDC 341
G C+ C + GH++RDC
Sbjct: 244 YGGSRGGGDRTCYKCGEAGHISRDC 268
Score = 42.7 bits (99), Expect = 0.002
Identities = 16/43 (37%), Positives = 23/43 (53%), Gaps = 7/43 (16%)
Query: 306 VTCFKCNQKGHYASHCPEGP-------LMCWNCNKPGHLARDC 341
+TCF+C + GH + CP C+ C + GHL+RDC
Sbjct: 70 MTCFRCGEAGHMSRDCPNSAKPGAAKGFECYKCGQEGHLSRDC 112
Score = 40.8 bits (94), Expect = 0.007
Identities = 15/42 (35%), Positives = 23/42 (54%), Gaps = 7/42 (16%)
Query: 307 TCFKCNQKGHYASHCPEG-------PLMCWNCNKPGHLARDC 341
+C C ++GHYA CPE C+ C + GH++R+C
Sbjct: 17 SCRNCGKEGHYARECPEADSKGDERSTTCFRCGEEGHMSREC 58
Score = 38.5 bits (88), Expect = 0.034
Identities = 21/75 (28%), Positives = 29/75 (38%), Gaps = 25/75 (33%)
Query: 305 GVTCFKCNQKGHYASHCP-----------------------EGPLMCWNCNKPGHLARDC 341
G C+KC Q+GH + CP G C+ C GH++RDC
Sbjct: 96 GFECYKCGQEGHLSRDCPSSQGGSRGGYGQKRGRSGAQGGYSGDRTCYKCGDAGHISRDC 155
Query: 342 RIPKVEDAANVAGAR 356
P + + AG R
Sbjct: 156 --PNGQGGYSGAGDR 168
>CNBP_CHICK (O42395) Cellular nucleic acid binding protein (CNBP)
Length = 172
Score = 46.6 bits (109), Expect = 1e-04
Identities = 17/41 (41%), Positives = 26/41 (62%), Gaps = 1/41 (2%)
Query: 308 CFKCNQKGHYASHCP-EGPLMCWNCNKPGHLARDCRIPKVE 347
C++C + GH A C + C+NC + GH+A+DC+ PK E
Sbjct: 48 CYRCGESGHLAKDCDLQEDKACYNCGRGGHIAKDCKEPKRE 88
Score = 46.2 bits (108), Expect = 2e-04
Identities = 19/38 (50%), Positives = 22/38 (57%), Gaps = 4/38 (10%)
Query: 308 CFKCNQKGHYASHCPEGPL----MCWNCNKPGHLARDC 341
C+ C + GH A C E C+NC KPGHLARDC
Sbjct: 69 CYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 106
Score = 42.7 bits (99), Expect = 0.002
Identities = 19/56 (33%), Positives = 25/56 (43%), Gaps = 22/56 (39%)
Query: 308 CFKCNQKGHYASHCPEG----------------------PLMCWNCNKPGHLARDC 341
CFKC + GH+A CP G P +C+ C + GHLA+DC
Sbjct: 6 CFKCGRTGHWARECPTGIGRGRGMRSRGRAGFQFMSSSLPDICYRCGESGHLAKDC 61
Score = 42.7 bits (99), Expect = 0.002
Identities = 16/39 (41%), Positives = 25/39 (64%), Gaps = 1/39 (2%)
Query: 306 VTCFKCNQKGHYASHCPE-GPLMCWNCNKPGHLARDCRI 343
V C++C + GH A +C + + C+ C + GHLAR+C I
Sbjct: 130 VKCYRCGETGHVAINCSKTSEVNCYRCGESGHLARECTI 168
Score = 34.7 bits (78), Expect = 0.49
Identities = 12/35 (34%), Positives = 19/35 (54%), Gaps = 1/35 (2%)
Query: 308 CFKCNQKGHYASHCPEGPLM-CWNCNKPGHLARDC 341
C+ C + GH A C C++C + GH+ +DC
Sbjct: 93 CYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDC 127
Score = 33.1 bits (74), Expect = 1.4
Identities = 12/38 (31%), Positives = 20/38 (52%), Gaps = 2/38 (5%)
Query: 304 DGVTCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDC 341
D C+ C + GH C + + C+ C + GH+A +C
Sbjct: 110 DEQKCYSCGEFGHIQKDCTK--VKCYRCGETGHVAINC 145
>CNBP_MOUSE (P53996) Cellular nucleic acid binding protein (CNBP)
(Zinc finger protein 9)
Length = 178
Score = 46.2 bits (108), Expect = 2e-04
Identities = 19/38 (50%), Positives = 22/38 (57%), Gaps = 4/38 (10%)
Query: 308 CFKCNQKGHYASHCPEGPL----MCWNCNKPGHLARDC 341
C+ C + GH A C E C+NC KPGHLARDC
Sbjct: 75 CYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 112
Score = 46.2 bits (108), Expect = 2e-04
Identities = 17/41 (41%), Positives = 26/41 (62%), Gaps = 1/41 (2%)
Query: 308 CFKCNQKGHYASHCP-EGPLMCWNCNKPGHLARDCRIPKVE 347
C++C + GH A C + C+NC + GH+A+DC+ PK E
Sbjct: 54 CYRCGESGHLAKDCDLQEDEACYNCGRGGHIAKDCKEPKRE 94
Score = 42.7 bits (99), Expect = 0.002
Identities = 16/39 (41%), Positives = 25/39 (64%), Gaps = 1/39 (2%)
Query: 306 VTCFKCNQKGHYASHCPE-GPLMCWNCNKPGHLARDCRI 343
V C++C + GH A +C + + C+ C + GHLAR+C I
Sbjct: 136 VKCYRCGETGHVAINCSKTSEVNCYRCGESGHLARECTI 174
Score = 42.4 bits (98), Expect = 0.002
Identities = 21/72 (29%), Positives = 29/72 (40%), Gaps = 28/72 (38%)
Query: 308 CFKCNQKGHYASHCPEG----------------------------PLMCWNCNKPGHLAR 339
CFKC + GH+A CP G P +C+ C + GHLA+
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGRGGFTSDRGFQFVSSSLPDICYRCGESGHLAK 65
Query: 340 DCRIPKVEDAAN 351
DC + + E N
Sbjct: 66 DCDLQEDEACYN 77
Score = 34.7 bits (78), Expect = 0.49
Identities = 12/35 (34%), Positives = 19/35 (54%), Gaps = 1/35 (2%)
Query: 308 CFKCNQKGHYASHCPEGPLM-CWNCNKPGHLARDC 341
C+ C + GH A C C++C + GH+ +DC
Sbjct: 99 CYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDC 133
Score = 33.1 bits (74), Expect = 1.4
Identities = 12/38 (31%), Positives = 20/38 (52%), Gaps = 2/38 (5%)
Query: 304 DGVTCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDC 341
D C+ C + GH C + + C+ C + GH+A +C
Sbjct: 116 DEQKCYSCGEFGHIQKDCTK--VKCYRCGETGHVAINC 151
>GAG_HV1EL (P04592) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 499
Score = 45.8 bits (107), Expect = 2e-04
Identities = 27/75 (36%), Positives = 34/75 (45%), Gaps = 13/75 (17%)
Query: 291 GNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEGPLM--CWNCNKPGHLARDCRIPKVED 348
GN PR + + CF C ++GH A +C P CW C K GH +DC E
Sbjct: 381 GNFKGPRKI------IKCFNCGKEGHIAKNC-RAPRKKGCWRCGKEGHQLKDC----TER 429
Query: 349 AANVAGARRPTAGGR 363
AN G P+ GR
Sbjct: 430 QANFLGRIWPSHKGR 444
Score = 35.4 bits (80), Expect = 0.29
Identities = 13/33 (39%), Positives = 22/33 (66%), Gaps = 2/33 (6%)
Query: 313 QKGHYASHCPEGPLMCWNCNKPGHLARDCRIPK 345
Q+G++ P + C+NC K GH+A++CR P+
Sbjct: 379 QRGNFKG--PRKIIKCFNCGKEGHIAKNCRAPR 409
>GAG_HV1MN (P05888) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 506
Score = 45.4 bits (106), Expect = 3e-04
Identities = 23/60 (38%), Positives = 29/60 (48%), Gaps = 7/60 (11%)
Query: 306 VTCFKCNQKGHYASHCPEGPLM--CWNCNKPGHLARDCRIPKVEDAANVAGARRPTAGGR 363
+ CF C ++GH A +C P CW C K GH +DC E AN G P+ GR
Sbjct: 392 IKCFNCGKEGHIAKNC-RAPRKRGCWKCGKEGHQMKDC----TERQANFLGKIWPSCKGR 446
Score = 34.3 bits (77), Expect = 0.64
Identities = 19/58 (32%), Positives = 31/58 (52%), Gaps = 4/58 (6%)
Query: 290 GGNAIAPRPLAVSLDGVT--CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPK 345
GG R LA ++ VT Q+G++ + + C+NC K GH+A++CR P+
Sbjct: 356 GGPGHKARVLAEAMSQVTNSATIMMQRGNFRNQ--RKIIKCFNCGKEGHIAKNCRAPR 411
>GAG_HV1J3 (P12494) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 499
Score = 45.4 bits (106), Expect = 3e-04
Identities = 23/60 (38%), Positives = 30/60 (49%), Gaps = 7/60 (11%)
Query: 306 VTCFKCNQKGHYASHCPEGPLM--CWNCNKPGHLARDCRIPKVEDAANVAGARRPTAGGR 363
+ CF C ++GH A +C P CW C K GH +DC E AN G P++ GR
Sbjct: 389 IKCFNCGKEGHLARNC-RAPRKKGCWKCGKEGHQMKDCN----ERQANFLGKIWPSSKGR 443
Score = 35.8 bits (81), Expect = 0.22
Identities = 21/58 (36%), Positives = 31/58 (53%), Gaps = 4/58 (6%)
Query: 290 GGNAIAPRPLAVSLDGVTCFKC--NQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPK 345
GG R LA ++ VT Q+G++ + + C+NC K GHLAR+CR P+
Sbjct: 353 GGPGHKARVLAEAMSQVTNSTTIMMQRGNFRNQ--RKIIKCFNCGKEGHLARNCRAPR 408
>GAG_OMVVS (P16900) Gag polyprotein [Contains: Core protein p16;
Core protein p25; Core protein p14]
Length = 446
Score = 45.1 bits (105), Expect = 4e-04
Identities = 18/50 (36%), Positives = 28/50 (56%), Gaps = 1/50 (2%)
Query: 308 CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKVEDAANVAGARR 357
C+ C + GH A C +G ++C +C K GH+ +DCR K + +RR
Sbjct: 386 CYYCGKPGHLARQCRQG-IICHHCGKRGHMQKDCRQKKGNPTSQQGNSRR 434
Score = 33.1 bits (74), Expect = 1.4
Identities = 12/20 (60%), Positives = 14/20 (70%)
Query: 323 EGPLMCWNCNKPGHLARDCR 342
+G C+ C KPGHLAR CR
Sbjct: 381 QGVQKCYYCGKPGHLARQCR 400
>GAG_HV1Y2 (P35962) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 499
Score = 45.1 bits (105), Expect = 4e-04
Identities = 24/60 (40%), Positives = 29/60 (48%), Gaps = 7/60 (11%)
Query: 306 VTCFKCNQKGHYASHCPEGPLM--CWNCNKPGHLARDCRIPKVEDAANVAGARRPTAGGR 363
V CF C ++GH A +C P CW C K GH +DC E AN G P+ GR
Sbjct: 389 VKCFNCGKEGHIAKNC-RAPRKKGCWKCGKEGHQMKDC----TERQANFLGKIWPSHKGR 443
Score = 34.7 bits (78), Expect = 0.49
Identities = 19/58 (32%), Positives = 31/58 (52%), Gaps = 4/58 (6%)
Query: 290 GGNAIAPRPLAVSLDGVT--CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPK 345
GG R LA ++ VT Q+G++ + + C+NC K GH+A++CR P+
Sbjct: 353 GGPGHKARVLAEAMSQVTNSATIMMQRGNFRNQ--RKTVKCFNCGKEGHIAKNCRAPR 408
>GAG_HV1N5 (P12493) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 499
Score = 45.1 bits (105), Expect = 4e-04
Identities = 24/60 (40%), Positives = 29/60 (48%), Gaps = 7/60 (11%)
Query: 306 VTCFKCNQKGHYASHCPEGPLM--CWNCNKPGHLARDCRIPKVEDAANVAGARRPTAGGR 363
V CF C ++GH A +C P CW C K GH +DC E AN G P+ GR
Sbjct: 389 VKCFNCGKEGHIAKNC-RAPRKKGCWKCGKEGHQMKDC----TERQANFLGKIWPSHKGR 443
Score = 35.4 bits (80), Expect = 0.29
Identities = 20/58 (34%), Positives = 31/58 (52%), Gaps = 4/58 (6%)
Query: 290 GGNAIAPRPLAVSLDGVT--CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPK 345
GG R LA ++ VT QKG++ + + C+NC K GH+A++CR P+
Sbjct: 353 GGPGHKARVLAEAMSQVTNPATIMIQKGNFRNQ--RKTVKCFNCGKEGHIAKNCRAPR 408
>GAG_HV1B5 (P04593) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 511
Score = 45.1 bits (105), Expect = 4e-04
Identities = 24/60 (40%), Positives = 30/60 (50%), Gaps = 7/60 (11%)
Query: 306 VTCFKCNQKGHYASHCPEGPLM--CWNCNKPGHLARDCRIPKVEDAANVAGARRPTAGGR 363
V CF C ++GH A +C + P CW C K GH +DC E AN G P+ GR
Sbjct: 389 VKCFNCGKEGHIARNC-KAPRKKGCWKCGKEGHQMKDC----TERQANFLGKIWPSYKGR 443
Score = 33.5 bits (75), Expect = 1.1
Identities = 19/58 (32%), Positives = 31/58 (52%), Gaps = 4/58 (6%)
Query: 290 GGNAIAPRPLAVSLDGVTCFKC--NQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPK 345
GG R LA ++ VT Q+G++ + + C+NC K GH+AR+C+ P+
Sbjct: 353 GGPGHKARVLAEAMSQVTNSTTIMMQRGNFRNQ--RKIVKCFNCGKEGHIARNCKAPR 408
>GAG_HV1Z2 (P12495) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 500
Score = 44.7 bits (104), Expect = 5e-04
Identities = 23/60 (38%), Positives = 29/60 (48%), Gaps = 7/60 (11%)
Query: 306 VTCFKCNQKGHYASHCPEGPLM--CWNCNKPGHLARDCRIPKVEDAANVAGARRPTAGGR 363
+ CF C ++GH A +C P CW C K GH +DC E AN G P+ GR
Sbjct: 391 IKCFNCGKEGHIAKNC-RAPRRKGCWKCGKEGHQLKDC----TERQANFLGKIWPSHKGR 445
Score = 36.2 bits (82), Expect = 0.17
Identities = 13/33 (39%), Positives = 22/33 (66%), Gaps = 2/33 (6%)
Query: 313 QKGHYASHCPEGPLMCWNCNKPGHLARDCRIPK 345
Q+G++ P + C+NC K GH+A++CR P+
Sbjct: 380 QRGNFKG--PRKTIKCFNCGKEGHIAKNCRAPR 410
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 51,178,841
Number of Sequences: 164201
Number of extensions: 2302833
Number of successful extensions: 7488
Number of sequences better than 10.0: 137
Number of HSP's better than 10.0 without gapping: 78
Number of HSP's successfully gapped in prelim test: 60
Number of HSP's that attempted gapping in prelim test: 7015
Number of HSP's gapped (non-prelim): 357
length of query: 433
length of database: 59,974,054
effective HSP length: 113
effective length of query: 320
effective length of database: 41,419,341
effective search space: 13254189120
effective search space used: 13254189120
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0178.13


