

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
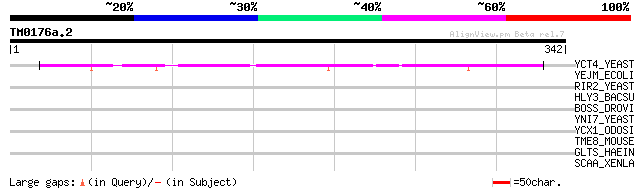
Query= TM0176a.2
(342 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
YCT4_YEAST (P25625) Hypothetical 42.5 kDa protein in TSM1-ARE1 i... 118 2e-26
YEJM_ECOLI (P33922) Hypothetical protein yejM 32 3.0
RIR2_YEAST (P09938) Ribonucleoside-diphosphate reductase small c... 32 3.0
HLY3_BACSU (P54175) Hemolysin III homolog 32 3.0
BOSS_DROVI (Q24738) Bride of sevenless protein precursor 31 3.9
YNI7_YEAST (P48231) Hypothetical 132.5 kDa protein in TOP2-MKT1 ... 31 5.2
YCX1_ODOSI (P49827) Hypothetical 41.6 kDa protein in psaA-ycf32.... 30 6.7
TME8_MOUSE (Q9ESN3) Transmembrane protein 8 precursor (M83 protein) 30 6.7
GLTS_HAEIN (P45240) Sodium/glutamate symport carrier protein (Gl... 30 6.7
SCAA_XENLA (P51167) Amiloride-sensitive sodium channel alpha-sub... 30 8.8
>YCT4_YEAST (P25625) Hypothetical 42.5 kDa protein in TSM1-ARE1
intergenic region
Length = 357
Score = 118 bits (296), Expect = 2e-26
Identities = 97/350 (27%), Positives = 155/350 (43%), Gaps = 56/350 (16%)
Query: 19 VIDASAGDVDPRYRDCIRQCQETGCVAQSCF----PHCKFPSDGEHFDRPWYMQEPLYLQ 74
++ S GD + DC C+ S P D E FD P PLY +
Sbjct: 15 LVTCSPGDNLDEFIDCTYACEYNRRCPNSQINYIDPETNMFHDIEFFDTP-----PLYSK 69
Query: 75 WKKWDCQNDCRYHCM-------LDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVAFS 127
WDC +DC Y C +D E+E ++HGKWPF R+ G QE S FS
Sbjct: 70 LLFWDCISDCDYQCQHIITRWRIDEEEEI-------YQFHGKWPFLRVLGTQEFFSTIFS 122
Query: 128 ALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLW-HIY-GLLSLNSWFWSAVFHSR 185
N H+ G+V F ++ + D + +W ++Y + + +W S+VFH R
Sbjct: 123 IGNFIPHYKGFVKFSRIIREE---GDRRRKNSRSILIWNYLYVTVAGMLAWTASSVFHCR 179
Query: 186 DVDLTETLDY--SSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFY 243
D+ +TE LDY + +L G+ I A + S + + + +A + A H++ + +
Sbjct: 180 DLIITEKLDYFFAGLTVLTGFHAIFARMTSMFLYPKIAQAF-TASVAAIFALHILRL-YV 237
Query: 244 KLDYGWNMIVCVVMAVVQLVIWAVWAGLSGHP-SRWKLW--------------------- 281
Y +NM + V+Q ++ + + + H + KL
Sbjct: 238 DWSYTYNMRFNIFFGVLQYILLIMLSCQNYHALQKQKLMGEFKKTAYSSFKRQIFKLCVI 297
Query: 282 --LVVIAGGLAMLLEIYDFPPYEGLLDAHALWHATTIPLTYIWWSFIRDD 329
L+VI +AM LE++DF YE +DAHALWH TI +++ + F +D
Sbjct: 298 PILLVIVTTMAMSLELFDFFSYEWQIDAHALWHLCTIWPSWVLYDFFLED 347
>YEJM_ECOLI (P33922) Hypothetical protein yejM
Length = 586
Score = 31.6 bits (70), Expect = 3.0
Identities = 25/85 (29%), Positives = 39/85 (45%), Gaps = 11/85 (12%)
Query: 171 LSLNSWFWSAVFHSRDVDLTET--LDYSSAVILLGYSLILAI-----LRSFNVRDEATRV 223
L LN W V + + ++ L + S ++L L+ A LRS R R
Sbjct: 113 LHLNPIVWQLVINPDENEMARDWQLMFISVPVILLLELVFATWSWQKLRSLTRRRRFARP 172
Query: 224 MVSAPLIAFVITHVMYI----NFYK 244
+ + IAF+ +HV+YI NFY+
Sbjct: 173 LAAFLFIAFIASHVVYIWADANFYR 197
>RIR2_YEAST (P09938) Ribonucleoside-diphosphate reductase small
chain 1 (EC 1.17.4.1) (Ribonucleotide reductase small
subunit)
Length = 399
Score = 31.6 bits (70), Expect = 3.0
Identities = 19/59 (32%), Positives = 27/59 (45%), Gaps = 10/59 (16%)
Query: 87 HCMLDREKERKLLN-------LVPVKYHGKWPFARIYGMQEAASVAFSALNLAMHFHGW 138
H + + EKE LLN L P+KYH W + Y EA+ ++L+ H W
Sbjct: 67 HKLKEMEKEEPLLNEDKERTVLFPIKYHEIW---QAYKRAEASFWTAEEIDLSKDIHDW 122
>HLY3_BACSU (P54175) Hemolysin III homolog
Length = 213
Score = 31.6 bits (70), Expect = 3.0
Identities = 36/152 (23%), Positives = 63/152 (40%), Gaps = 23/152 (15%)
Query: 170 LLSLNSWFWSAVFHSRDVDLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPL 229
LL L+S ++ H + D+ E +D+S+ +L+ + +L T +++ L
Sbjct: 53 LLYLSSTLLHSITHKKTKDILEIIDHSAIYVLIAGTYTPFLLGPLKGTLGFTLLVIVWSL 112
Query: 230 --------IAFVITHVMYINFYKLDYGWNMIVCVVMAVVQLVIWAVWAGLSGHPSRWKLW 281
I FV ++ F L GW MI+ V ++A LSG L+
Sbjct: 113 ALGGIVFKIFFVKRFILLSTFVYLVMGWLMIIAVK---------PLYASLSG-AGFSLLF 162
Query: 282 LVVIAGGLAMLLEIYDFPPYEGLLDAHALWHA 313
L I + + I+ P+ HA+WH+
Sbjct: 163 LGGILYSVGTIFYIWKKIPFH-----HAIWHS 189
>BOSS_DROVI (Q24738) Bride of sevenless protein precursor
Length = 893
Score = 31.2 bits (69), Expect = 3.9
Identities = 18/47 (38%), Positives = 28/47 (59%), Gaps = 3/47 (6%)
Query: 252 IVCVVMAVVQLVIWAVWAGLSGHPSRWK---LWLVVIAGGLAMLLEI 295
I+ V+ AV+ L+IW+VW LS W+ + L + A G A+L+ I
Sbjct: 723 ILIVIGAVLILIIWSVWIALSMFGDEWRDAAIPLGMQASGWAVLVGI 769
>YNI7_YEAST (P48231) Hypothetical 132.5 kDa protein in TOP2-MKT1
intergenic region
Length = 1178
Score = 30.8 bits (68), Expect = 5.2
Identities = 16/47 (34%), Positives = 28/47 (59%), Gaps = 4/47 (8%)
Query: 68 QEPLYLQWKK---WDCQNDCRYHCML-DREKERKLLNLVPVKYHGKW 110
Q+P Y+ WK+ W+ +++ +L D K+ L NL+P K++G W
Sbjct: 53 QDPSYVGWKQVGGWEEKDELTSEDLLVDVNKDTFLGNLLPDKFYGDW 99
>YCX1_ODOSI (P49827) Hypothetical 41.6 kDa protein in psaA-ycf32.1
and ycf32.2-acpP intergenic regions (ORF355)
Length = 355
Score = 30.4 bits (67), Expect = 6.7
Identities = 27/116 (23%), Positives = 52/116 (44%), Gaps = 23/116 (19%)
Query: 117 GMQEAASVAFSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSW 176
G++ + FS L ++F+ + I+L Y + + Y W +Y L+SL W
Sbjct: 128 GLETIRNTLFSML---VYFYFFAELRIMLSYFISINP-------YTRPW-VY-LISLTDW 175
Query: 177 FWSAVFHSRDVDLTETLDYSSAVILLGYSLILAILRSF--NVRDEATRVMVSAPLI 230
+ +FH L S V+LLG+ L+ ++ + N+ D ++ + P +
Sbjct: 176 IYDILFH---------LGISKRVVLLGFPLLPILIHAALGNLIDSLNHLVFTMPFL 222
>TME8_MOUSE (Q9ESN3) Transmembrane protein 8 precursor (M83 protein)
Length = 769
Score = 30.4 bits (67), Expect = 6.7
Identities = 17/71 (23%), Positives = 33/71 (45%), Gaps = 7/71 (9%)
Query: 249 WNMI-VCVVMAVVQLVIWAVWAGLSG--HPSRWKLWLVVIAGGLAML---LEIY-DFPPY 301
WN++ CV V+ +W G G +P+ W+ W+ + G++M + +Y
Sbjct: 654 WNLMGPCVFAFVIMASMWIYRCGHRGQCYPTSWQRWVFYLLPGISMASVGIAMYTSMMTS 713
Query: 302 EGLLDAHALWH 312
+ H++WH
Sbjct: 714 DNYYYTHSIWH 724
>GLTS_HAEIN (P45240) Sodium/glutamate symport carrier protein
(Glutamate permease)
Length = 404
Score = 30.4 bits (67), Expect = 6.7
Identities = 26/108 (24%), Positives = 55/108 (50%), Gaps = 18/108 (16%)
Query: 191 ETLDYSSAVILLGYSLI--LAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKLDYG 248
ETL +S V+LLGY L+ + +L++FN+ + P++ I + + ++K+D G
Sbjct: 9 ETLALASLVLLLGYFLVKRINVLKTFNIPE---------PVVGGFIVAIGLLIWHKID-G 58
Query: 249 WNMIVCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIY 296
+ + ++++ GLS + SR +I GG +++ ++
Sbjct: 59 TSFNFDKNLQTTMMLVFFTSIGLSANFSR------LIKGGKPLVVFLF 100
>SCAA_XENLA (P51167) Amiloride-sensitive sodium channel
alpha-subunit (Epithelial Na+ channel alpha subunit)
(Alpha ENaC) (Nonvoltage-gated sodium channel 1 alpha
subunit) (SCNEA) (Alpha NaCH)
Length = 632
Score = 30.0 bits (66), Expect = 8.8
Identities = 48/221 (21%), Positives = 85/221 (37%), Gaps = 34/221 (15%)
Query: 34 CIRQCQETGCVAQSCFPHCKFP-SDGEHF-----DRPW---YMQEPLYLQWKKWDCQNDC 84
C+R C + VA+ + +P S G+ + + W Y + + K C C
Sbjct: 377 CVRSCFQAAMVARCGCGYAFYPLSPGDQYCDYNKHKSWGHCYYKLIIEFTSNKLGCFTKC 436
Query: 85 RYHCMLDREKERKLLNLVPVKYHGKW---PFARIYGMQEAASVAFSALNLAMHFHGWVSF 141
R C++ + + P + W +R Y + + +A LN+ +
Sbjct: 437 RKPCLVSEYQLTAGYSKWPNRVSQDWVLHTLSRQYNLTDRNGIA--KLNI---------Y 485
Query: 142 FILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRDVDLTETLDYSSAVIL 201
F L YK L+ + ++ LLSL WS F S + + E L+ ++
Sbjct: 486 FEELNYKTILE---------SPTINMAMLLSLLGSQWSLWFGSSVLSVVEMLELVIDFVI 536
Query: 202 LGYSLIL--AILRSFNVRDEATRVMVSAPLIAFVITHVMYI 240
+G ++L + N +E T V AP A + V +I
Sbjct: 537 IGVMILLHRYYYKKANEGEETTVVPTPAPAFADLEQQVPHI 577
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.328 0.142 0.485
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,709,739
Number of Sequences: 164201
Number of extensions: 1808256
Number of successful extensions: 4609
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 4604
Number of HSP's gapped (non-prelim): 11
length of query: 342
length of database: 59,974,054
effective HSP length: 111
effective length of query: 231
effective length of database: 41,747,743
effective search space: 9643728633
effective search space used: 9643728633
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0176a.2


