

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0173b.3
(286 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
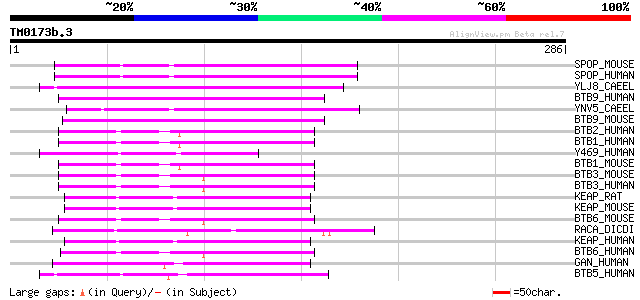
Score E
Sequences producing significant alignments: (bits) Value
SPOP_MOUSE (Q6ZWS8) Speckle-type POZ protein (PDX-1 C-terminal i... 66 1e-10
SPOP_HUMAN (O43791) Speckle-type POZ protein 66 1e-10
YLJ8_CAEEL (P34371) Hypothetical protein C50C3.8 in chromosome III 61 4e-09
BTB9_HUMAN (Q96Q07) BTB/POZ domain containing protein 9 60 6e-09
YNV5_CAEEL (P34568) Hypothetical protein T16H12.5 in chromosome III 58 3e-08
BTB9_MOUSE (Q8C726) BTB/POZ domain containing protein 9 57 7e-08
BTB2_HUMAN (Q9BX70) BTB/POZ domain containing protein 2 55 2e-07
BTB1_HUMAN (Q9H0C5) BTB/POZ domain containing protein 1 (Hepatit... 54 3e-07
Y469_HUMAN (Q9UJP4) Hypothetical protein KIAA0469 52 2e-06
BTB1_MOUSE (P58544) BTB/POZ domain containing protein 1 (Glucose... 50 6e-06
BTB3_MOUSE (P58545) BTB/POZ domain containing protein 3 48 3e-05
BTB3_HUMAN (Q9Y2F9) BTB/POZ domain containing protein 3 48 3e-05
KEAP_RAT (P57790) Kelch-like ECH-associated protein 1 (Cytosolic... 47 5e-05
KEAP_MOUSE (Q9Z2X8) Kelch-like ECH-associated protein 1 (Cytosol... 47 5e-05
BTB6_MOUSE (Q8K2J9) BTB/POZ domain containing protein 6 47 5e-05
RACA_DICDI (P34147) RAS-related protein racA 47 7e-05
KEAP_HUMAN (Q14145) Kelch-like ECH-associated protein 1 (Cytosol... 47 7e-05
BTB6_HUMAN (Q96KE9) BTB/POZ domain containing protein 6 (Lens BT... 45 3e-04
GAN_HUMAN (Q9H2C0) Gigaxonin (Kelch-like protein 16) 44 6e-04
BTB5_HUMAN (Q9NXS3) BTB/POZ domain containing protein 5 44 6e-04
>SPOP_MOUSE (Q6ZWS8) Speckle-type POZ protein (PDX-1 C-terminal
interacting factor 1) (PCIF1)
Length = 374
Score = 65.9 bits (159), Expect = 1e-10
Identities = 41/156 (26%), Positives = 71/156 (45%), Gaps = 3/156 (1%)
Query: 24 GTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTED 83
G AH ILA+ SPV +M + + R+ +I+ V + + F+Y+ +
Sbjct: 208 GQEFQAHKAILAARSPVFSAMFEHEMEESKKNRV-EINDVEPEVFKEMMCFIYTGKAPN- 265
Query: 84 EMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNH 143
+D+ LLA + Y + LK C L ++ EN ++L LA L A L + ++
Sbjct: 266 -LDKMADDLLAAADKYALERLKVMCEDALCSNLSVENAAEILILADLHSADQLKTQAVDF 324
Query: 144 LTRNFKTVEETEGWRFLTKHDPWLELEVLRFMDESE 179
+ + V ET GW+ + P L E R + ++
Sbjct: 325 INYHASDVLETSGWKSMVVSHPHLVAEAYRSLASAQ 360
>SPOP_HUMAN (O43791) Speckle-type POZ protein
Length = 374
Score = 65.9 bits (159), Expect = 1e-10
Identities = 41/156 (26%), Positives = 71/156 (45%), Gaps = 3/156 (1%)
Query: 24 GTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTED 83
G AH ILA+ SPV +M + + R+ +I+ V + + F+Y+ +
Sbjct: 208 GQEFQAHKAILAARSPVFSAMFEHEMEESKKNRV-EINDVEPEVFKEMMCFIYTGKAPN- 265
Query: 84 EMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNH 143
+D+ LLA + Y + LK C L ++ EN ++L LA L A L + ++
Sbjct: 266 -LDKMADDLLAAADKYALERLKVMCEDALCSNLSVENAAEILILADLHSADQLKTQAVDF 324
Query: 144 LTRNFKTVEETEGWRFLTKHDPWLELEVLRFMDESE 179
+ + V ET GW+ + P L E R + ++
Sbjct: 325 INYHASDVLETSGWKSMVVSHPHLVAEAYRSLASAQ 360
>YLJ8_CAEEL (P34371) Hypothetical protein C50C3.8 in chromosome III
Length = 410
Score = 60.8 bits (146), Expect = 4e-09
Identities = 45/157 (28%), Positives = 70/157 (43%), Gaps = 1/157 (0%)
Query: 16 DIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFL 75
D IH + I AH IL SPV +SM P + + I I D+V A + F+
Sbjct: 220 DCVIHVGN-KHIKAHRCILGQNSPVFKSMFSSPNMIEAQKGEIHIEDAKYDSVRAMVEFM 278
Query: 76 YSRRCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPD 135
Y+ E +LA++ Y V LK +C + ++Q +N +NV + + A
Sbjct: 279 YTGATESLESQGNIDEILAIADKYEVLMLKDQCERLIAQTINLKNVTQIAMFSDTYTADY 338
Query: 136 LHLRCMNHLTRNFKTVEETEGWRFLTKHDPWLELEVL 172
L + LT + + V +T+ W L K L E+L
Sbjct: 339 LKSAVIRFLTTHHRVVIKTQDWISLKKSRHELANELL 375
>BTB9_HUMAN (Q96Q07) BTB/POZ domain containing protein 9
Length = 612
Score = 60.1 bits (144), Expect = 6e-09
Identities = 39/138 (28%), Positives = 64/138 (46%), Gaps = 1/138 (0%)
Query: 26 RIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCT-EDE 84
R PAH ILA+ +++ + E I + +A T L ++Y+ R T DE
Sbjct: 46 RFPAHRVILAARCQYFRALLYGGMRESQPEAEIPLQDTTAEAFTMLLKYIYTGRATLTDE 105
Query: 85 MDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNHL 144
+ + L+L+H Y P L+ ++ L +N +NV +A L P L C +
Sbjct: 106 KEEVLLDFLSLAHKYGFPELEDSTSEYLCTILNIQNVCMTFDVASLYSLPKLTCMCCMFM 165
Query: 145 TRNFKTVEETEGWRFLTK 162
RN + V +EG+ L+K
Sbjct: 166 DRNAQEVLSSEGFLSLSK 183
>YNV5_CAEEL (P34568) Hypothetical protein T16H12.5 in chromosome III
Length = 451
Score = 57.8 bits (138), Expect = 3e-08
Identities = 38/151 (25%), Positives = 67/151 (44%), Gaps = 3/151 (1%)
Query: 30 HSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEMDRYG 89
H ILA+ S V +M + S + + + + + L ++Y+ + +++
Sbjct: 285 HKAILAARSRVFSAMFEH-HMQESDTNMTTVDDIEPEVMRELLVYMYTGQTKY--IEQMA 341
Query: 90 MHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNHLTRNFK 149
L+A + Y + LK C + L ++ T+N L LA + A L +N + N
Sbjct: 342 QSLIAAADKYQLDRLKVMCEQALCYQLTTDNASLTLMLADMYSASQLRAHSINFINVNAN 401
Query: 150 TVEETEGWRFLTKHDPWLELEVLRFMDESET 180
V +TEGW L + P L EV R + +T
Sbjct: 402 EVMDTEGWEDLVRDHPKLLEEVFRALATQQT 432
>BTB9_MOUSE (Q8C726) BTB/POZ domain containing protein 9
Length = 612
Score = 56.6 bits (135), Expect = 7e-08
Identities = 37/136 (27%), Positives = 63/136 (46%), Gaps = 1/136 (0%)
Query: 28 PAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCT-EDEMD 86
PAH ILA+ +++ + E I + +A T L ++Y+ R T DE +
Sbjct: 48 PAHRVILAARCQYFRALLYGGMRESQPEAEIPLQDTTAEAFTMLLRYIYTGRATLTDEKE 107
Query: 87 RYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNHLTR 146
+ L+L+H Y P L+ ++ L +N +NV +A L P L C + R
Sbjct: 108 EVLLDFLSLAHKYGFPELEDSTSEYLCTILNIQNVCMTFDVASLYSLPKLTCMCCMFMDR 167
Query: 147 NFKTVEETEGWRFLTK 162
N + V ++G+ L+K
Sbjct: 168 NAQEVLASDGFLSLSK 183
>BTB2_HUMAN (Q9BX70) BTB/POZ domain containing protein 2
Length = 525
Score = 55.1 bits (131), Expect = 2e-07
Identities = 43/135 (31%), Positives = 63/135 (45%), Gaps = 10/135 (7%)
Query: 26 RIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEM 85
RIPAH +LA S V ++M + S+E I++ V A A L FLYS DE+
Sbjct: 131 RIPAHRFVLAVGSAVFDAMFNGGMATTSTE--IELPDVEPAAFLALLKFLYS-----DEV 183
Query: 86 D---RYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMN 142
M L + Y VP L+ C + L + + +N +L ARL D P L C+
Sbjct: 184 QIGPETVMTTLYTAKKYAVPALEAHCVEFLKKNLRADNAFMLLTQARLFDEPQLASLCLE 243
Query: 143 HLTRNFKTVEETEGW 157
++ +N EG+
Sbjct: 244 NIDKNTADAITAEGF 258
>BTB1_HUMAN (Q9H0C5) BTB/POZ domain containing protein 1 (Hepatitis
C virus NS5A-transactivated protein 8) (NS5ATP8)
Length = 482
Score = 54.3 bits (129), Expect = 3e-07
Identities = 42/135 (31%), Positives = 65/135 (48%), Gaps = 10/135 (7%)
Query: 26 RIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEM 85
RIPAH +LA+ S V ++M + S+E I++ V A A L FLYS DE+
Sbjct: 89 RIPAHRFVLAAGSAVFDAMFNGGMATTSAE--IELPDVEPAAFLALLRFLYS-----DEV 141
Query: 86 D---RYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMN 142
M L + Y VP L+ C + L++ + +N +L ARL D P L C++
Sbjct: 142 QIGPETVMTTLYTAKKYAVPALEAHCVEFLTKHLRADNAFMLLTQARLFDEPQLASLCLD 201
Query: 143 HLTRNFKTVEETEGW 157
+ ++ EG+
Sbjct: 202 TIDKSTMDAISAEGF 216
>Y469_HUMAN (Q9UJP4) Hypothetical protein KIAA0469
Length = 539
Score = 52.0 bits (123), Expect = 2e-06
Identities = 32/113 (28%), Positives = 54/113 (47%), Gaps = 3/113 (2%)
Query: 16 DIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFL 75
D+ + + G PAH +LA+ SP +M + +ER +++HGVP D + L F
Sbjct: 36 DVTLEAAGGRDFPAHRAVLAAASPYFRAMFAGQLRESRAER-VRLHGVPPDMLQLLLDFS 94
Query: 76 YSRRCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLA 128
Y+ R + LL + + P +K+ C L Q+++ N +DM A
Sbjct: 95 YTGRVAVSGDN--AEPLLRAADLLQFPAVKEACGAFLQQQLDLANCLDMQDFA 145
>BTB1_MOUSE (P58544) BTB/POZ domain containing protein 1 (Glucose
signal repressing protein)
Length = 488
Score = 50.1 bits (118), Expect = 6e-06
Identities = 41/135 (30%), Positives = 64/135 (47%), Gaps = 10/135 (7%)
Query: 26 RIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEM 85
RIPA +LA+ S V ++M + S+E I++ V A A L FLYS DE+
Sbjct: 95 RIPAPRFVLAAGSAVFDAMFNGGMATTSAE--IELPDVEPAAFLALLRFLYS-----DEV 147
Query: 86 D---RYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMN 142
M L + Y VP L+ C + L++ + +N +L ARL D P L C++
Sbjct: 148 QIGPETVMTTLYTAKKYAVPALEAHCVEFLTKHLRADNAFMLLTQARLFDEPQLASLCLD 207
Query: 143 HLTRNFKTVEETEGW 157
+ ++ EG+
Sbjct: 208 TIDKSTVDAISAEGF 222
>BTB3_MOUSE (P58545) BTB/POZ domain containing protein 3
Length = 532
Score = 47.8 bits (112), Expect = 3e-05
Identities = 38/135 (28%), Positives = 60/135 (44%), Gaps = 10/135 (7%)
Query: 26 RIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEM 85
R+P H +LA S V +M E I+I V A A L ++Y DE+
Sbjct: 144 RLPGHKYVLAVGSSVFHAMFYGELAEDKDE--IRIPDVEPAAFLAMLKYIYC-----DEI 196
Query: 86 DRYGMHLLALSHV---YMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMN 142
D +LA + Y+VPHL + C L ++ +N +L + L + PDL RC
Sbjct: 197 DLAADTVLATLYAAKKYIVPHLARACVNFLETSLSAKNACVLLSQSCLFEEPDLTQRCWE 256
Query: 143 HLTRNFKTVEETEGW 157
+ + ++EG+
Sbjct: 257 VIDAQAELALKSEGF 271
>BTB3_HUMAN (Q9Y2F9) BTB/POZ domain containing protein 3
Length = 522
Score = 47.8 bits (112), Expect = 3e-05
Identities = 38/135 (28%), Positives = 60/135 (44%), Gaps = 10/135 (7%)
Query: 26 RIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEM 85
R+P H +LA S V +M E I+I V A A L ++Y DE+
Sbjct: 134 RLPGHKYVLAVGSSVFHAMFYGELAEDKDE--IRIPDVEPAAFLAMLKYIYC-----DEI 186
Query: 86 DRYGMHLLALSHV---YMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMN 142
D +LA + Y+VPHL + C L ++ +N +L + L + PDL RC
Sbjct: 187 DLAADTVLATLYAAKKYIVPHLARACVNFLETSLSAKNACVLLSQSCLFEEPDLTQRCWE 246
Query: 143 HLTRNFKTVEETEGW 157
+ + ++EG+
Sbjct: 247 VIDAQAELALKSEGF 261
>KEAP_RAT (P57790) Kelch-like ECH-associated protein 1 (Cytosolic
inhibitor of Nrf2) (INrf2)
Length = 624
Score = 47.0 bits (110), Expect = 5e-05
Identities = 28/127 (22%), Positives = 60/127 (47%), Gaps = 3/127 (2%)
Query: 29 AHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEMDRY 88
AH +LAS SPV ++M + + E ++ I G+ + + F Y+ + E +
Sbjct: 95 AHKVVLASSSPVFKAMFTNGLREQGME-VVSIEGIHPKVMERLIEFAYTASISVGE--KC 151
Query: 89 GMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNHLTRNF 148
+H++ + +Y + + + C+ L Q+++ N + + A +LH R ++ +F
Sbjct: 152 VLHVMNGAVMYQIDSVVRACSDFLVQQLDPSNAIGIANFAEQIGCTELHQRAREYIYMHF 211
Query: 149 KTVEETE 155
V + E
Sbjct: 212 GEVAKQE 218
>KEAP_MOUSE (Q9Z2X8) Kelch-like ECH-associated protein 1 (Cytosolic
inhibitor of Nrf2)
Length = 624
Score = 47.0 bits (110), Expect = 5e-05
Identities = 28/127 (22%), Positives = 60/127 (47%), Gaps = 3/127 (2%)
Query: 29 AHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEMDRY 88
AH +LAS SPV ++M + + E ++ I G+ + + F Y+ + E +
Sbjct: 95 AHKVVLASSSPVFKAMFTNGLREQGME-VVSIEGIHPKVMERLIEFAYTASISVGE--KC 151
Query: 89 GMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNHLTRNF 148
+H++ + +Y + + + C+ L Q+++ N + + A +LH R ++ +F
Sbjct: 152 VLHVMNGAVMYQIDSVVRACSDFLVQQLDPSNAIGIANFAEQIGCTELHQRAREYIYMHF 211
Query: 149 KTVEETE 155
V + E
Sbjct: 212 GEVAKQE 218
>BTB6_MOUSE (Q8K2J9) BTB/POZ domain containing protein 6
Length = 410
Score = 47.0 bits (110), Expect = 5e-05
Identities = 39/135 (28%), Positives = 59/135 (42%), Gaps = 10/135 (7%)
Query: 26 RIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEM 85
R+PAH +LA S V +M SE I I V A L ++YS DE+
Sbjct: 22 RVPAHKYVLAVGSSVFYAMFYGDLAEVKSE--IHIPDVEPAAFLVLLKYMYS-----DEI 74
Query: 86 DRYGMHLLALSHV---YMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMN 142
D +LA + Y+VP L + C L + +N +L +RL + P+L RC
Sbjct: 75 DLEADTVLATLYAAKKYIVPALAKACVNFLETSLEAKNACVLLSQSRLFEEPELTQRCWE 134
Query: 143 HLTRNFKTVEETEGW 157
+ + +EG+
Sbjct: 135 VIDAQAEMALRSEGF 149
>RACA_DICDI (P34147) RAS-related protein racA
Length = 598
Score = 46.6 bits (109), Expect = 7e-05
Identities = 41/183 (22%), Positives = 85/183 (46%), Gaps = 20/183 (10%)
Query: 23 DGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTE 82
+G I AH ++ S S V+ ++I R R ++ V D+ A L +LY+
Sbjct: 412 EGRSIFAHKALIFSRSNVVNALIGRNFIEREG-KVEMSDDVTFDSFMAILEYLYTAHAPI 470
Query: 83 DEMDRYGM----------HLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCD 132
+E D G+ L++L +Y+ +++ T G+ + + +++ +L A+ +
Sbjct: 471 EEGDSVGILVLGNRFDLRRLVSLCELYISKEIERATTIGIFR--SELDIIGLLICAQRHN 528
Query: 133 APDLHLRCMNHLTRNFKTVEETEGWRFL----TKH---DPWLELEVLRFMDESETRREKS 185
AP L C++ ++ N++ + + + L KH + W + LR +++ E EK
Sbjct: 529 APQLEKFCLHFISSNYQPMRRRKEFALLDDKTKKHIEDNQWPPIHYLRAVEDYEKEMEKL 588
Query: 186 RRS 188
+ S
Sbjct: 589 KSS 591
>KEAP_HUMAN (Q14145) Kelch-like ECH-associated protein 1 (Cytosolic
inhibitor of Nrf2)
Length = 624
Score = 46.6 bits (109), Expect = 7e-05
Identities = 28/127 (22%), Positives = 60/127 (47%), Gaps = 3/127 (2%)
Query: 29 AHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEMDRY 88
AH +LAS SPV ++M + + E ++ I G+ + + F Y+ + E +
Sbjct: 95 AHKVVLASSSPVFKAMFTNGLREQGME-VVSIEGIHPKVMERLIEFAYTASISMGE--KC 151
Query: 89 GMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNHLTRNF 148
+H++ + +Y + + + C+ L Q+++ N + + A +LH R ++ +F
Sbjct: 152 VLHVMNGAVMYQIDSVVRACSDFLVQQLDPSNAIGIANFAEQIGCVELHQRAREYIYMHF 211
Query: 149 KTVEETE 155
V + E
Sbjct: 212 GEVAKQE 218
>BTB6_HUMAN (Q96KE9) BTB/POZ domain containing protein 6 (Lens BTB
domain protein)
Length = 410
Score = 44.7 bits (104), Expect = 3e-04
Identities = 38/134 (28%), Positives = 58/134 (42%), Gaps = 10/134 (7%)
Query: 27 IPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEMD 86
+PAH +LA S V +M SE I I V A L ++YS DE+D
Sbjct: 23 VPAHKYVLAVGSSVFYAMFYGDLAEVKSE--IHIPDVEPAAFLILLKYMYS-----DEID 75
Query: 87 RYGMHLLALSHV---YMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNH 143
+LA + Y+VP L + C L + +N +L +RL + P+L RC
Sbjct: 76 LEADTVLATLYAAKKYIVPALAKACVNFLETSLEAKNACVLLSQSRLFEEPELTQRCWEV 135
Query: 144 LTRNFKTVEETEGW 157
+ + +EG+
Sbjct: 136 IDAQAEMALRSEGF 149
>GAN_HUMAN (Q9H2C0) Gigaxonin (Kelch-like protein 16)
Length = 597
Score = 43.5 bits (101), Expect = 6e-04
Identities = 33/136 (24%), Positives = 60/136 (43%), Gaps = 7/136 (5%)
Query: 23 DGTRIPAHSNILASMSPVLESMID-RPRKHRSSERIIQIHGVPGDAVTAFLTFLYSR--R 79
DG IP NILA+ SP + + ++ P K S I++ G+ + L +++S R
Sbjct: 37 DGEEIPVQKNILAAASPYIRTKLNYNPPKDDGSTYKIELEGISVMVMREILDYIFSGQIR 96
Query: 80 CTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLR 139
ED + ++ + + ++ LK C + L + EN + + A +H
Sbjct: 97 LNEDTI----QDVVQAADLLLLTDLKTLCCEFLEGCIAAENCIGIRDFALHYCLHHVHYL 152
Query: 140 CMNHLTRNFKTVEETE 155
+L +F+ V TE
Sbjct: 153 ATEYLETHFRDVSSTE 168
>BTB5_HUMAN (Q9NXS3) BTB/POZ domain containing protein 5
Length = 571
Score = 43.5 bits (101), Expect = 6e-04
Identities = 33/151 (21%), Positives = 71/151 (46%), Gaps = 8/151 (5%)
Query: 16 DIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFL 75
DI + D +I AH +LAS+SP ++M + + + + + A+ A + +
Sbjct: 36 DIILRVGD-VKIHAHKVVLASVSPYFKAMFTGNLSEKENSEV-EFQCIDETALQAIVEYA 93
Query: 76 YSRRC--TEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDA 133
Y+ ++D ++ LL +++ + + + C L +++ N + + + A
Sbjct: 94 YTGTVFISQDTVES----LLPAANLLQIKLVLKECCAFLESQLDPGNCIGISRFAETYGC 149
Query: 134 PDLHLRCMNHLTRNFKTVEETEGWRFLTKHD 164
DL+L ++ +NF+ V +TE + LT D
Sbjct: 150 RDLYLAATKYICQNFEAVCQTEEFFELTHAD 180
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.136 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,693,199
Number of Sequences: 164201
Number of extensions: 1333229
Number of successful extensions: 3941
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 41
Number of HSP's that attempted gapping in prelim test: 3887
Number of HSP's gapped (non-prelim): 75
length of query: 286
length of database: 59,974,054
effective HSP length: 109
effective length of query: 177
effective length of database: 42,076,145
effective search space: 7447477665
effective search space used: 7447477665
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0173b.3


