

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0166.1
(916 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
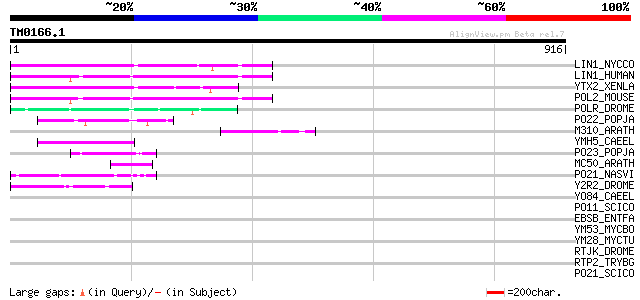
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 130 1e-29
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 119 5e-26
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 114 2e-24
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 106 3e-22
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 77 2e-13
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 69 4e-11
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 67 2e-10
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 67 3e-10
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 62 9e-09
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 56 5e-07
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 54 2e-06
Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I retro... 52 7e-06
YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III 44 0.002
PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type... 41 0.017
EBSB_ENTFA (P36921) Cell wall enzyme ebsB 39 0.048
YM53_MYCBO (P64956) Hypothetical protein Mb2253c 38 0.14
YM28_MYCTU (P64955) Hypothetical protein Rv2228c/MT2287 38 0.14
RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile elem... 37 0.31
RTP2_TRYBG (P15594) Retrotransposable element SLACS 132 kDa prot... 34 1.6
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 34 2.0
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 130 bits (328), Expect = 1e-29
Identities = 116/445 (26%), Positives = 205/445 (46%), Gaps = 24/445 (5%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKR-QTPEMVTDY 59
PGPDGF F++ F + ++ +F+ + G L I LIPK + P +Y
Sbjct: 470 PGPDGFTSEFYQTFKEELVPILLNLFQNIEKEGILPNTFYEANITLIPKPGKDPTRKENY 529
Query: 60 RLISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQ-IANCLLIGNEVVDTMLSS 118
R ISL+ K+L K+L R+Q + ++ +Q F G Q N N +
Sbjct: 530 RPISLMNIDAKILNKILTNRIQQHIKKIIHHDQVGFIPGSQGWFNIRKSINVIQHINKLK 589
Query: 119 GRGGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEF 178
+ +L +D KA+DNI+ F++ ++ +G + I + S T +++NG +
Sbjct: 590 NKDHMILSIDAEKAFDNIQHPFMIRTLKKIGIEGTFLKLIEAIYSKPTANIILNGVKLKS 649
Query: 179 FEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPC--GVVLGPGVCLNHLQFADDT 236
F + G RQG PLSPLLFN+ ++ L++ AIRE G+ +G L FADD
Sbjct: 650 FPLRSGTRQGCPLSPLLFNIVMEVLAI-----AIREEKAIKGIHIGSEEIKLSL-FADDM 703
Query: 237 LVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVGTL 296
+V+ + +L +V+ + SG K+N KS ++ + + V
Sbjct: 704 IVYLENTRDSTTKLLEVIKEYSNVSGYKINTHKSVAFIYTNNNQAEKTVKDSIPFTVVPK 763
Query: 297 PLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSN----SVSLSGRLVVLRS--VASAI 350
+ YLG L D + + E +R +A + S GR+ +++ + AI
Sbjct: 764 KMKYLGVYLTKDV--KDLYKENYETLRKEIAEDVNKWKNIPCSWLGRINIVKMSILPKAI 821
Query: 351 PIYHMAVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKF 410
++ KAPL D+EK++ F+W ++ +A + + + GG+ + L+
Sbjct: 822 YNFNAIPIKAPLSYFKDLEKIILHFIW-----NQKKPQIAKTLLSNKNKAGGITLPDLRL 876
Query: 411 RNKAMLLKWAWRFGQEPEA-LWRRV 434
K++++K AW + + E +W R+
Sbjct: 877 YYKSIVIKTAWYWHKNREVDVWNRI 901
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 119 bits (297), Expect = 5e-26
Identities = 110/447 (24%), Positives = 198/447 (43%), Gaps = 30/447 (6%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPK--RQTPEMVTD 58
PGP+GF F++ + + ++++F+ + G L I LIPK R T + +
Sbjct: 470 PGPEGFTAEFYQRYKEELVPFLLKLFQSIEKEGILPNSFYEASIILIPKPGRDTTKK-EN 528
Query: 59 YRLISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKG-------RQIANCLLIGNEV 111
+R ISL+ K+L K+LA ++Q + L+ +Q F R+ N + N
Sbjct: 529 FRPISLMNIDAKILNKILANQIQQHIKKLIHHDQVGFIPAMQGWFNIRKSINIIQHINRT 588
Query: 112 VDTMLSSGRGGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLV 171
DT ++ +D KA+D I+ F++ + LG + I + T +++
Sbjct: 589 KDT------NHMIISIDAEKAFDKIQQPFMLKPLNKLGIDGTYLKIIRAIYDKPTANIIL 642
Query: 172 NGSPTEFFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGPGVCLNHLQ 231
NG E ++ G RQG PLSPLL N+ L V+ + + G+ LG L
Sbjct: 643 NGQKLEAPPLKTGTRQGCPLSPLLPNIV---LEVLARAIRQEKEIKGIQLGKEEVKLSL- 698
Query: 232 FADDTLVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGV 291
FADD +V+ + + L +++ F SG K+N KS+ + +Q+ EL
Sbjct: 699 FADDMIVYLENPIVSAQNLLKLISNFSKVSGYKINVQKSQAFLYTNNRQTESQIMSELPF 758
Query: 292 VVGTLPLPYLGSVLGGDPRR--RGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASA 349
+ + + YLG L D + + + P++ +++ + S GR+ +++
Sbjct: 759 TIASKRIKYLGIQLTRDVKDLFKENYKPLLNEIKEDTNKWKNIPCSWVGRINIVKMAILP 818
Query: 350 IPIYHMAV--YKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEF 407
IY K P+ ++EK F+W ++ A +A + ++ + GG+ +
Sbjct: 819 KVIYRFNAIPIKLPMTFFTELEKTTLKFIW-----NQKRAHIAKSTLSQKNKAGGITLPD 873
Query: 408 LKFRNKAMLLKWAWRFGQEPEA-LWRR 433
K KA + K AW + Q + W R
Sbjct: 874 FKLYYKATVTKTAWYWYQNRDIDQWNR 900
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 114 bits (284), Expect = 2e-24
Identities = 91/386 (23%), Positives = 183/386 (46%), Gaps = 21/386 (5%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PG DG FF+ FW + + ++ E ++ G+L ++L+PK+ ++ ++R
Sbjct: 467 PGLDGLTIEFFQFFWDTLGPDFHRVLTEAFKKGELPLSCRRAVLSLLPKKGDLRLIKNWR 526
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGR 120
+SL+ + YK++AK ++ RL+ V+ ++ +Q GR I + + + +++ +G
Sbjct: 527 PVSLLSTDYKIVAKAISLRLKSVLAEVIHPDQSYTVPGRTIFDNVFLIRDLLHFARRTGL 586
Query: 121 GGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFE 180
L LD KA+D ++ +L+ ++ FGP++ ++ + ++A V +N S T
Sbjct: 587 SLAFLSLDQEKAFDRVDHQYLIGTLQAYSFGPQFVGYLKTMYASAECLVKINWSLTAPLA 646
Query: 181 IEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVL-GPGVCLNHLQFADDTLVF 239
+G+RQG PLS L++L ++ + +R+R G+VL P + + +ADD ++
Sbjct: 647 FGRGVRQGCPLSGQLYSLAIEPFLCL-----LRKRLTGLVLKEPDMRVVLSAYADDVILV 701
Query: 240 CDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVEL-GVVVGTLPL 298
+ +E + + S ++N++KS G S V+ L + + +
Sbjct: 702 AQ-DLVDLERAQECQEVYAAASSARINWSKSS--GLLEGSLKVDFLPPAFRDISWESKII 758
Query: 299 PYLGSVLGGD--PRRRGFWAPVIEKVRVALAS*G-----SNSVSLSGRLVVLRSVASAIP 351
YLG L + P + F IE L G + +S+ GR +V+ + ++
Sbjct: 759 KYLGVYLSAEEYPVSQNF----IELEECVLTRLGKWKGFAKVLSMRGRALVINQLVASQI 814
Query: 352 IYHMAVYKAPLGVVADIEKLMRCFLW 377
Y + +A I++ + FLW
Sbjct: 815 WYRLICLSPTQEFIAKIQRRLLDFLW 840
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 106 bits (264), Expect = 3e-22
Identities = 107/448 (23%), Positives = 198/448 (43%), Gaps = 30/448 (6%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQT-PEMVTDY 59
PGPDGF+ F++ F + + ++F + G L I LIPK Q P + ++
Sbjct: 497 PGPDGFSAEFYQTFKEDLIPILHKLFHKIEVEGTLPNSFYEATITLIPKPQKDPTKIENF 556
Query: 60 RLISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKG-------RQIANCLLIGNEVV 112
R ISL+ K+L K+LA R+Q + ++ +Q F G R+ N + N++
Sbjct: 557 RPISLMNIDAKILNKILANRIQEHIKAIIHPDQVGFIPGMQGWFNIRKSINVIHYINKLK 616
Query: 113 DTMLSSGRGGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVN 172
D + ++ LD KA+D I+ F++ ++E G + I + S + VN
Sbjct: 617 D------KNHMIISLDAEKAFDKIQHPFMIKVLERSGIQGPYLNMIKAIYSKPVANIKVN 670
Query: 173 GSPTEFFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGPGVCLNHLQF 232
G E ++ G RQG PLSP LFN+ L V+ + ++ G+ +G L
Sbjct: 671 GEKLEAIPLKSGTRQGCPLSPYLFNIV---LEVLARAIRQQKEIKGIQIGKEEVKISL-L 726
Query: 233 ADDTLVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVV 292
ADD +V+ EL +++ +F G K+N KS + ++
Sbjct: 727 ADDMIVYISDPKNSTRELLNLINSFGEVVGYKINSNKSMAFLYTKNKQAEKEIRETTPFS 786
Query: 293 VGTLPLPYLGSVLGGDPR---RRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRS--VA 347
+ T + YLG L + + + F + ++++ L S GR+ +++ +
Sbjct: 787 IVTNNIKYLGVTLTKEVKDLYDKNF-KSLKKEIKEDLRRWKDLPCSWIGRINIVKMAILP 845
Query: 348 SAIPIYHMAVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEF 407
AI ++ K P ++E + F+W + +A + ++ GG+ +
Sbjct: 846 KAIYRFNAIPIKIPTQFFNELEGAICKFVW-----NNKKPRIAKSLLKDKRTSGGITMPD 900
Query: 408 LKFRNKAMLLKWAWRFGQEPEA-LWRRV 434
LK +A+++K AW + ++ + W R+
Sbjct: 901 LKLYYRAIVIKTAWYWYRDRQVDQWNRI 928
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 77.0 bits (188), Expect = 2e-13
Identities = 97/385 (25%), Positives = 154/385 (39%), Gaps = 25/385 (6%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDG RE I IM + G L + IPK T + D+R
Sbjct: 360 PGPDGITPKSAREVPSGIMLRIMNLI---LWCGNLPHSIRLARTVFIPKTVTAKRPQDFR 416
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGR 120
IS+ + + L +LA RL + + + F T G A+ I + V+ R
Sbjct: 417 PISVPSVLVRQLNAILATRLNSSINWDPRQRGFLPTDG--CADNATIVDLVLRHSHKHFR 474
Query: 121 GGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFE 180
++ LD +KA+D++ + + + G + ++ + ++ +G +E F
Sbjct: 475 SCYIANLDVSKAFDSLSHASIYDTLRAYGAPKGFVDYVQNTYEGGGTSLNGDGWSSEEFV 534
Query: 181 IEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGPGVCLNHLQFADDTLVFC 240
+G++QGDPLSP+LFNL VM L G +G + N FADD ++F
Sbjct: 535 PARGVKQGDPLSPILFNL------VMDRLLRTLPSEIGAKVGNAI-TNAAAFADDLVLFA 587
Query: 241 DPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVGTLPLP- 299
+ ++ L D FL GLKLN K VG + VG+ +P
Sbjct: 588 ETRMG-LQVLLDKTLDFLSIVGLKLNADKCFTVGIKGQPKQKCTVLEAQSFYVGSSEIPS 646
Query: 300 --------YLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIP 351
YLG R R E + L + RL LR+V
Sbjct: 647 LKRTDEWKYLGINFTATGRVR---CNPAEDIGPKLQRLTKAPLKPQQRLFALRTVLIPQL 703
Query: 352 IYHMAVYKAPLGVVADIEKLMRCFL 376
+ +A+ +GV+ +KL+R ++
Sbjct: 704 YHKLALGSVAIGVLRKTDKLIRYYV 728
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 69.3 bits (168), Expect = 4e-11
Identities = 66/237 (27%), Positives = 106/237 (43%), Gaps = 31/237 (13%)
Query: 46 LIPKRQTPEMVTDYRLISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCL 105
LIPK E +++R I++ ++ +LL ++LA RL+ + ++ ++ G + + L
Sbjct: 63 LIPKDGDLENPSNWRPITIASALQRLLHRILAKRLEAAVELHPAQKGYARIDGTLVNSLL 122
Query: 106 LIGNEVVDTMLSSGRGGF----LLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSC 161
L DT +SS R ++ LD KA+D + + ++ LG +I
Sbjct: 123 L------DTYISSRREQRKTYNVVSLDVRKAFDTVSHSSICRALQRLGIDEGTSNYITGS 176
Query: 162 VSTATLAVLVN-GSPTEFFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVV 220
+S +T + V GS T I +G++QGDPLSP LFN + E C +
Sbjct: 177 LSDSTTTIRVGPGSQTRKICIRRGVKQGDPLSPFLFN------------AVLDELLCSLQ 224
Query: 221 LGPGV-------CLNHLQFADDTLVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKS 270
PG+ + L FADD L+ D + L V F G+ LN KS
Sbjct: 225 STPGIGGTIGEEKIPVLAFADDLLLLEDNDVLLPTTLATVANFFRL-RGMSLNAKKS 280
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 67.4 bits (163), Expect = 2e-10
Identities = 46/158 (29%), Positives = 75/158 (47%), Gaps = 12/158 (7%)
Query: 349 AIPIYHMAVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKE-IGGLGVEF 407
A+P+Y M+ ++ + + M F W R+I+WVAW+++CK KE GGLG
Sbjct: 2 ALPVYAMSCFRLSKLLCKKLTSAMTEFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRD 61
Query: 408 LKFRNKAMLLKWAWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVN 467
L + N+A+L K ++R +P L R+L ++Y +S + G R + I
Sbjct: 62 LGWFNQALLAKQSFRIIHQPHTLLSRLLRSRYFPHS---SMMECSVGTRPSYAWRSIIHG 118
Query: 468 VVLEDSVLGKTFRDQCFCVVGDGLRISFWRDPWVASKT 505
L L +T +GDG+ W D W+ +T
Sbjct: 119 RELLSRGLLRT--------IGDGIHTKVWLDRWIMDET 148
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 66.6 bits (161), Expect = 3e-10
Identities = 41/160 (25%), Positives = 75/160 (46%), Gaps = 1/160 (0%)
Query: 47 IPKRQTPEMVTDYRLISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLL 106
IPK+ P ++YR ISL +++ +++ +R++ +LLS +Q F R + L+
Sbjct: 673 IPKKGNPSSPSNYRPISLTDPFARIMERIICSRIRSEYSHLLSPHQHGFLNFRSCPSSLV 732
Query: 107 IGNEVVDTMLSSGRGGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTAT 166
+ ++L + + +L DFAKA+D + L+ + G W + T
Sbjct: 733 RSISLYHSILKNEKSLDILFFDFAKAFDKVSHPILLKKLALFGLDKLTCSWFKEFLHLRT 792
Query: 167 LAVLVNG-SPTEFFEIEKGLRQGDPLSPLLFNLCVQGLSV 205
+V +N + + I G+ QG PLLF L + L +
Sbjct: 793 FSVKINKFVSSNAYPISSGVPQGSVSGPLLFILFINDLLI 832
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 61.6 bits (148), Expect = 9e-09
Identities = 42/141 (29%), Positives = 70/141 (48%), Gaps = 6/141 (4%)
Query: 101 IANCLLIGNEVVDTMLSSGRGGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHS 160
+AN +++ + + L G+ ++ LD KA+D + ++ M G + +I S
Sbjct: 3 LANIIMLEHYIKLRRLK-GKTYNVVSLDIRKAFDTVSHPAILRAMRAFGIDDGMQDFIMS 61
Query: 161 CVSTATLAVLVNGSPTEFFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVV 220
++ A ++V G T I G++QGDPLSP+LFN+ + L N E+P G
Sbjct: 62 TITDAYTNIVVGGRTTNKIYIRNGVKQGDPLSPVLFNIVLDELVTRLN----DEQP-GAS 116
Query: 221 LGPGVCLNHLQFADDTLVFCD 241
+ P + L FADD L+ D
Sbjct: 117 MTPACKIASLAFADDLLLLED 137
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 55.8 bits (133), Expect = 5e-07
Identities = 32/71 (45%), Positives = 41/71 (57%), Gaps = 1/71 (1%)
Query: 167 LAVLVNGSPTEFFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGV-VLGPGV 225
L ++NG+P +GLRQGDPLSP LF LC + LS + R + R G+ V
Sbjct: 10 LLFIINGAPQGLVTPSRGLRQGDPLSPYLFILCTEVLSGLCRRAQEQGRLPGIRVSNNSP 69
Query: 226 CLNHLQFADDT 236
+NHL FADDT
Sbjct: 70 RINHLLFADDT 80
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 53.9 bits (128), Expect = 2e-06
Identities = 61/243 (25%), Positives = 101/243 (41%), Gaps = 19/243 (7%)
Query: 2 GPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYRL 61
GPDG T W+ I E I +F G+ + LIPK +R
Sbjct: 336 GPDGMTTTA----WNSIDECIKSLFNMIMYHGQCPRRYLDSRTVLIPKEPGTMDPACFRP 391
Query: 62 ISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGRG 121
+S+ + ++LA R+ LL Q +F +A + + ++ +G
Sbjct: 392 LSIASVALRHFHRILANRIGE--HGLLDTRQRAFIVADGVAENTSLLSAMIKEARMKIKG 449
Query: 122 GFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWR---CWIHSCVSTATLAVLVNGSPTEF 178
++ LD KA+D++E +++ + R W++ T V G +
Sbjct: 450 LYIAILDVKKAFDSVEHRSILDALRRKKLPLEMRNYIMWVYRNSKTRLEVVKTKG---RW 506
Query: 179 FEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGPGVCLNHLQFADDTLV 238
+G+RQGDPLSPLLFN CV + + RL G ++G + L FADD ++
Sbjct: 507 IRPARGVRQGDPLSPLLFN-CV--MDAVLRRL---PENTGFLMG-AEKIGALVFADDLVL 559
Query: 239 FCD 241
+
Sbjct: 560 LAE 562
>Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I
retrotransposable element R1DM (ORF 2)
Length = 1021
Score = 52.0 bits (123), Expect = 7e-06
Identities = 49/209 (23%), Positives = 88/209 (41%), Gaps = 16/209 (7%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMV--TD 58
PG DG N T + W I E + +F G + + K + +
Sbjct: 452 PGLDGINGTICKAVWRAIPEHLASLFSRCIRLGYFPAEWKCPRVVSLLKGPDKDKCEPSS 511
Query: 59 YRLISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSS 118
YR I L+ K+L ++ R++ V+P QF F +GR C+ V + + +
Sbjct: 512 YRGICLLPVFGKVLEAIMVNRVREVLPEGCRW-QFGFRQGR----CVEDAWRHVKSSVGA 566
Query: 119 GRGGFLLK--LDFAKAYDNIEWDFLMNMMEDLGFGPR--WRCWIHSCVSTATLAVLVNGS 174
++L +DF A+DN+EW ++ + DLG W+ + + AV+ + S
Sbjct: 567 SAAQYVLGTFVDFKGAFDNVEWSAALSRLADLGCREMGLWQSFF-----SGRRAVIRSSS 621
Query: 175 PTEFFEIEKGLRQGDPLSPLLFNLCVQGL 203
T + +G QG P ++++ + L
Sbjct: 622 GTVEVPVTRGCPQGSISGPFIWDILMDVL 650
>YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III
Length = 364
Score = 43.9 bits (102), Expect = 0.002
Identities = 43/178 (24%), Positives = 78/178 (43%), Gaps = 24/178 (13%)
Query: 127 LDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFEIEKGLR 186
LDF+KA+D + D L++ + + W+ ++ + V V + +E + G+
Sbjct: 23 LDFSKAFDKVSHDILLDKLTSIKINKHLIRWLDVFLTNRSFKVKVGNTLSEPKKTVCGVP 82
Query: 187 QGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGPGVCLNHLQFADDTLVFC----DP 242
QG +SP+LF + V +S P GV QFADD ++ +
Sbjct: 83 QGSVISPVLFGIFVNEISA--------NLPVGVYC--------KQFADDIKLYAAIPKNQ 126
Query: 243 ECAQMEELFDVMTAFLWGSGLKLNYAKS---ELVGCNVD-SAIVNQLAVELGVVVGTL 296
++ DV+T + + L LN +K+ L CN + + +N L ++ V+ L
Sbjct: 127 SQISLQLAIDVVTDWALNNKLLLNPSKTFHLTLGKCNTNYTYFLNGLPIKKEVLTRDL 184
>PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type I
retrotransposable element R1 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1004
Score = 40.8 bits (94), Expect = 0.017
Identities = 37/144 (25%), Positives = 62/144 (42%), Gaps = 11/144 (7%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPK-----RQTPEM 55
PGPDG R W I E + ++++ + + ++ K R P
Sbjct: 423 PGPDGIVGEMVRAVWGAIPEYMFCLYKQCLLESYFPQKWKIASLVILLKLLDRIRSDP-- 480
Query: 56 VTDYRLISLIGSIYKLLAKVLAARL-QGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDT 114
YR I L+ ++ K+L ++ RL Q +M +S QF+FT G+ + V+
Sbjct: 481 -GSYRPICLLDNLGKVLEGIMVKRLDQKLMDVEVSPYQFAFTYGKSTEDAWRCVQRHVE- 538
Query: 115 MLSSGRGGFLLKLDFAKAYDNIEW 138
S + L +DF A+DN+ W
Sbjct: 539 -CSEMKYVIGLNIDFQGAFDNLGW 561
>EBSB_ENTFA (P36921) Cell wall enzyme ebsB
Length = 135
Score = 39.3 bits (90), Expect = 0.048
Identities = 29/83 (34%), Positives = 44/83 (52%), Gaps = 4/83 (4%)
Query: 790 ILKWNVDGAARGAPGMGGIGGVVRDATGEIRGFFSMSLGVVFAYEAEVKAIRTALE--FS 847
+L+ VD A +G PG G GG+V T + R + LG+V +EAE K + AL+ +
Sbjct: 1 MLRIYVDAATKGNPGESG-GGIVY-LTDQSRQQLHVPLGIVSNHEAEFKVLIEALKKAIA 58
Query: 848 KEFGLSRVVVESDSTLAVGWVSK 870
E V++ SDS + V + K
Sbjct: 59 NEDNQQTVLLHSDSKIVVQTIEK 81
>YM53_MYCBO (P64956) Hypothetical protein Mb2253c
Length = 364
Score = 37.7 bits (86), Expect = 0.14
Identities = 32/111 (28%), Positives = 49/111 (43%), Gaps = 9/111 (8%)
Query: 796 DGAARGAPGMGGIGGVVRDAT-GEIRGFFSMSLGVVFAYEAEVKAIRTALEFSKEFGLSR 854
DG +RG PG G G VV A + ++G AE + + L+ + + G +
Sbjct: 8 DGGSRGNPGPAGYGAVVWTADHSTVLAESKQAIGRATNNVAEYRGLIAGLDDAVKLGATE 67
Query: 855 VVVESDSTLAVGWVSKRANRPWKLLNDLHMIDILRNEVECLRIYHVFREIN 905
V DS L V +S R WK+ + D+L+ V+ + FR IN
Sbjct: 68 AAVLMDSKLVVEQMSGR----WKVKHP----DLLKLYVQAQALASQFRRIN 110
>YM28_MYCTU (P64955) Hypothetical protein Rv2228c/MT2287
Length = 364
Score = 37.7 bits (86), Expect = 0.14
Identities = 32/111 (28%), Positives = 49/111 (43%), Gaps = 9/111 (8%)
Query: 796 DGAARGAPGMGGIGGVVRDAT-GEIRGFFSMSLGVVFAYEAEVKAIRTALEFSKEFGLSR 854
DG +RG PG G G VV A + ++G AE + + L+ + + G +
Sbjct: 8 DGGSRGNPGPAGYGAVVWTADHSTVLAESKQAIGRATNNVAEYRGLIAGLDDAVKLGATE 67
Query: 855 VVVESDSTLAVGWVSKRANRPWKLLNDLHMIDILRNEVECLRIYHVFREIN 905
V DS L V +S R WK+ + D+L+ V+ + FR IN
Sbjct: 68 AAVLMDSKLVVEQMSGR----WKVKHP----DLLKLYVQAQALASQFRRIN 110
>RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 36.6 bits (83), Expect = 0.31
Identities = 45/180 (25%), Positives = 78/180 (43%), Gaps = 10/180 (5%)
Query: 25 MFEEFYEGGKLVKGLNAGFIALIPKR-QTPEMVTDYRLISLIGSIYKLLAKVLAARLQGV 83
+F + G K + I +I K +TP V YR SL+ S+ K++ +++ RL
Sbjct: 480 IFNSVLDVGYFPKAWKSASIIMIHKTGKTPTDVDSYRPTSLLPSLGKIMERLILNRLLTC 539
Query: 84 --MPYLLSENQFSFTKGRQIANCL-LIGNEVVDTMLSS--GRGGFLLKLDFAKAYDNIEW 138
+ + + QF F L + N ++ M + G F LD +A+D +
Sbjct: 540 KDVTKAIPKFQFGFRLQHGTPEQLHRVVNFALEAMENKEYAVGAF---LDIQQAFDRVWH 596
Query: 139 DFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFEIEKGLRQGDPLSPLLFNL 198
L+ + L F P+ + S + T V V+G + I G+ QG L P L+++
Sbjct: 597 PGLLYKAKRL-FPPQLYLVVKSFLEERTFHVSVDGYKSSIKPIAAGVPQGSVLGPTLYSV 655
>RTP2_TRYBG (P15594) Retrotransposable element SLACS 132 kDa protein
(ORF2)
Length = 1182
Score = 34.3 bits (77), Expect = 1.6
Identities = 24/78 (30%), Positives = 36/78 (45%), Gaps = 1/78 (1%)
Query: 122 GFLLKLDFAKAYDNIEWDFLMNMME-DLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFE 180
G L LD AY+ I ++ + D + P WR +T + NG +E
Sbjct: 641 GSLAMLDGRNAYNAISRRAILEAVYGDSTWSPLWRLVSLLLGTTGEVGFYENGKLCHTWE 700
Query: 181 IEKGLRQGDPLSPLLFNL 198
+G+RQG L PLLF++
Sbjct: 701 STRGVRQGMVLGPLLFSI 718
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 33.9 bits (76), Expect = 2.0
Identities = 43/200 (21%), Positives = 76/200 (37%), Gaps = 6/200 (3%)
Query: 2 GPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYRL 61
GPDG + K + +F + +KG F I P + +R
Sbjct: 176 GPDGVTPRSWNALDDRYKRLLYNIFVFYGRVPSPIKGSRTVFTPKIEGGPDPGV---FRP 232
Query: 62 ISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGRG 121
+S+ I + K+LA R V Y E Q ++ + + + ++ +
Sbjct: 233 LSICSVILREFNKILARRF--VSCYTYDERQTAYLPIDGVCINVSMLTAIIAEAKRLRKE 290
Query: 122 GFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFEI 181
+ LD KA++++ L++ + + G P +I + + G E I
Sbjct: 291 LHIAILDLVKAFNSVYHSALIDAITEAGCPPGVVDYIADMYNNVITEMQFEGK-CELASI 349
Query: 182 EKGLRQGDPLSPLLFNLCVQ 201
G+ QGDPLS LF L +
Sbjct: 350 LAGVYQGDPLSGPLFTLAYE 369
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.142 0.462
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 112,804,884
Number of Sequences: 164201
Number of extensions: 4925536
Number of successful extensions: 12931
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 12877
Number of HSP's gapped (non-prelim): 45
length of query: 916
length of database: 59,974,054
effective HSP length: 119
effective length of query: 797
effective length of database: 40,434,135
effective search space: 32226005595
effective search space used: 32226005595
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0166.1


