

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
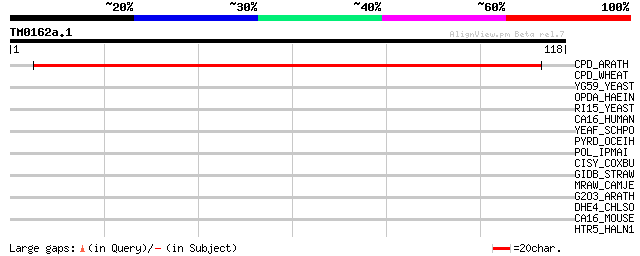
Query= TM0162a.1
(118 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CPD_ARATH (O04147) Cyclic phosphodiesterase (EC 3.1.4.-) (CPDase) 131 2e-31
CPD_WHEAT (P62809) Cyclic phosphodiesterase (EC 3.1.4.-) (CPDase... 32 0.26
YG59_YEAST (P53314) Hypothetical 26.7 kDa protein in TDS4-SOL4 i... 31 0.57
OPDA_HAEIN (P44573) Oligopeptidase A (EC 3.4.24.70) 30 0.75
RI15_YEAST (P43565) Serine/threonine-protein kinase RIM15 (EC 2.... 30 0.97
CA16_HUMAN (P12109) Collagen alpha 1(VI) chain precursor 29 1.7
YEAF_SCHPO (O14080) Hypothetical protein UNK4,15 in chromosome I 28 3.7
PYRD_OCEIH (Q8CWG1) Dihydroorotate dehydrogenase (EC 1.3.3.1) (D... 28 3.7
POL_IPMAI (P12894) Probable Pol polyprotein [Contains: Integrase... 28 3.7
CISY_COXBU (P18789) Citrate synthase (EC 2.3.3.1) 28 3.7
GIDB_STRAW (Q82FE3) Methyltransferase gidB (EC 2.1.-.-) (Glucose... 28 4.8
MRAW_CAMJE (Q9PPL4) S-adenosyl-methyltransferase mraW (EC 2.1.1.-) 27 6.3
G2O3_ARATH (O64692) Gibberellin 2-beta-dioxygenase 3 (EC 1.14.11... 27 6.3
DHE4_CHLSO (P28998) NADP-specific glutamate dehydrogenase (EC 1.... 27 6.3
CA16_MOUSE (Q04857) Collagen alpha 1(VI) chain precursor 27 6.3
HTR5_HALN1 (Q48318) Halobacterial transducer protein V 27 8.2
>CPD_ARATH (O04147) Cyclic phosphodiesterase (EC 3.1.4.-) (CPDase)
Length = 181
Score = 131 bits (330), Expect = 2e-31
Identities = 61/108 (56%), Positives = 75/108 (68%)
Query: 6 IETQKKEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKL 65
+E KK+VYSVWA+P + PR KLM LRSEF GP F PH+TV + LTAD+A
Sbjct: 1 MEEVKKDVYSVWALPDEESEPRFKKLMEALRSEFTGPRFVPHVTVAVSAYLTADEAKKMF 60
Query: 66 RSACEELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHF 113
SAC+ LKA+ TVDRV+ GTFF+QCV+LLL +P V+E HC NHF
Sbjct: 61 ESACDGLKAYTATVDRVSTGTFFFQCVFLLLQTTPEVMEAGEHCKNHF 108
>CPD_WHEAT (P62809) Cyclic phosphodiesterase (EC 3.1.4.-) (CPDase)
(Fragments)
Length = 51
Score = 32.0 bits (71), Expect = 0.26
Identities = 13/19 (68%), Positives = 14/19 (73%)
Query: 40 GGPHFEPHITVVGAITLTA 58
GGP FEPH TVVGA+ A
Sbjct: 4 GGPPFEPHATVVGAVAAAA 22
>YG59_YEAST (P53314) Hypothetical 26.7 kDa protein in TDS4-SOL4
intergenic region
Length = 239
Score = 30.8 bits (68), Expect = 0.57
Identities = 11/28 (39%), Positives = 18/28 (64%)
Query: 42 PHFEPHITVVGAITLTADDALNKLRSAC 69
P FEPH+TV + + D +NK+ ++C
Sbjct: 34 PVFEPHVTVTSHLVCNSKDDVNKILTSC 61
>OPDA_HAEIN (P44573) Oligopeptidase A (EC 3.4.24.70)
Length = 681
Score = 30.4 bits (67), Expect = 0.75
Identities = 18/58 (31%), Positives = 27/58 (46%), Gaps = 2/58 (3%)
Query: 19 IPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAI--TLTADDALNKLRSACEELKA 74
I P+H+RP + KL+ D R+ +PH T I +D LN+ S L +
Sbjct: 19 IKPEHIRPAVEKLIQDCRNTIEQVLKQPHFTWENFILPLTETNDRLNRAWSPVSHLNS 76
>RI15_YEAST (P43565) Serine/threonine-protein kinase RIM15 (EC
2.7.1.37)
Length = 1770
Score = 30.0 bits (66), Expect = 0.97
Identities = 18/50 (36%), Positives = 29/50 (58%), Gaps = 5/50 (10%)
Query: 18 AIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRS 67
AIPP H+R R + + D ++EFG +F + A+ DA+N+L+S
Sbjct: 1405 AIPP-HMRDRRSSKLNDSQTEFGSFNFRN----LSALDKANKDAINRLKS 1449
>CA16_HUMAN (P12109) Collagen alpha 1(VI) chain precursor
Length = 1028
Score = 29.3 bits (64), Expect = 1.7
Identities = 19/63 (30%), Positives = 34/63 (53%), Gaps = 3/63 (4%)
Query: 12 EVYSVWAIPPDHVRPRIAKLMTD--LRSEFGGPHFEPHITVVGAITLTADDALNKLRSAC 69
+V+SV AI PDH+ PR++ + TD R F + AI+ T D ++ +++
Sbjct: 185 KVFSV-AITPDHLEPRLSIIATDHTYRRNFTAADWGQSRDAEEAISQTIDTIVDMIKNNV 243
Query: 70 EEL 72
E++
Sbjct: 244 EQV 246
>YEAF_SCHPO (O14080) Hypothetical protein UNK4,15 in chromosome I
Length = 208
Score = 28.1 bits (61), Expect = 3.7
Identities = 16/61 (26%), Positives = 28/61 (45%), Gaps = 2/61 (3%)
Query: 46 PHITVVGAITLTADDALNKLRS--ACEELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVV 103
PHIT+ I L + + + A + + +VA G FY+ VY+ + +P +
Sbjct: 68 PHITLARGIKLKPNQSFKSVLQYIASKRKSPLNIKFGKVAIGDSFYKRVYIQVEKTPGLE 127
Query: 104 E 104
E
Sbjct: 128 E 128
>PYRD_OCEIH (Q8CWG1) Dihydroorotate dehydrogenase (EC 1.3.3.1)
(Dihydroorotate oxidase) (DHOdehase) (DHODase) (DHOD)
Length = 304
Score = 28.1 bits (61), Expect = 3.7
Identities = 15/52 (28%), Positives = 23/52 (43%)
Query: 22 DHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEELK 73
DH PR+AK T + + G E + V + T D +L +C +K
Sbjct: 82 DHEVPRLAKYDTSIIANIAGSSIEEYEFVAASFNQTTDVDALELNISCPNVK 133
>POL_IPMAI (P12894) Probable Pol polyprotein [Contains: Integrase
(IN); Reverse transcriptase/ribonuclease H (EC 2.7.7.49)
(EC 3.1.26.4) (RT)]
Length = 814
Score = 28.1 bits (61), Expect = 3.7
Identities = 19/70 (27%), Positives = 32/70 (45%), Gaps = 5/70 (7%)
Query: 24 VRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEELKAFQVTVDRVA 83
+RP + +LR F +PHI+ +TL A+ AL K+ A + + + R+
Sbjct: 192 LRPFLKIPSAELRPLFSILEGDPHISSPRTLTLAANQALQKVEKALQNAQ-----LQRIE 246
Query: 84 AGTFFYQCVY 93
F CV+
Sbjct: 247 DSQPFSLCVF 256
>CISY_COXBU (P18789) Citrate synthase (EC 2.3.3.1)
Length = 430
Score = 28.1 bits (61), Expect = 3.7
Identities = 18/67 (26%), Positives = 34/67 (49%), Gaps = 7/67 (10%)
Query: 39 FGGPHFEPH-----ITVVGAITLTADDALNKLRSACEELKAFQV--TVDRVAAGTFFYQC 91
F G H++ H ++ +GA++ DAL+ + A EL A ++ + +AA ++ Y
Sbjct: 121 FNGFHYDAHPMAMVLSTIGALSAFYHDALDITKPADRELSAIRLIAKMPTLAAMSYKYSI 180
Query: 92 VYLLLHP 98
+HP
Sbjct: 181 GQPFMHP 187
>GIDB_STRAW (Q82FE3) Methyltransferase gidB (EC 2.1.-.-) (Glucose
inhibited division protein B)
Length = 238
Score = 27.7 bits (60), Expect = 4.8
Identities = 23/82 (28%), Positives = 36/82 (43%), Gaps = 2/82 (2%)
Query: 4 TAIETQKKEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALN 63
T I + +EV IPP HV A D + +G P P+ ++ TA++ L
Sbjct: 120 TVIRGRAEEVMG--KIPPVHVVTARAVAPLDRLATWGIPLLRPYGEMLALKGDTAEEELK 177
Query: 64 KLRSACEELKAFQVTVDRVAAG 85
+A +L A + +V V G
Sbjct: 178 SAATALSKLGAVETSVLHVGEG 199
>MRAW_CAMJE (Q9PPL4) S-adenosyl-methyltransferase mraW (EC 2.1.1.-)
Length = 312
Score = 27.3 bits (59), Expect = 6.3
Identities = 13/41 (31%), Positives = 22/41 (52%)
Query: 35 LRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEELKAF 75
+R E G H I AI+ + ++ L RS+C +++AF
Sbjct: 264 MRCECGNNHSLGQIITKKAISASKEELLKNSRSSCAKMRAF 304
>G2O3_ARATH (O64692) Gibberellin 2-beta-dioxygenase 3 (EC
1.14.11.13) (Gibberellin 2-beta-hydroxylase 3)
(Gibberellin 2-oxidase 3) (GA 2-oxidase 3)
Length = 335
Score = 27.3 bits (59), Expect = 6.3
Identities = 13/36 (36%), Positives = 19/36 (52%)
Query: 42 PHFEPHITVVGAITLTADDALNKLRSACEELKAFQV 77
P +P ++ I LT DA ++ ACEE F+V
Sbjct: 18 PKCKPRPVLIPVIDLTDSDAKTQIVKACEEFGFFKV 53
>DHE4_CHLSO (P28998) NADP-specific glutamate dehydrogenase (EC
1.4.1.4) (NADP-GDH) (Fragment)
Length = 523
Score = 27.3 bits (59), Expect = 6.3
Identities = 22/76 (28%), Positives = 29/76 (37%), Gaps = 6/76 (7%)
Query: 31 LMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEELKAFQVTVDRVAAGTFFYQ 90
++T E+GG P T GA+ N L+ E LK + V AG
Sbjct: 267 VLTGKGQEYGGSEIRPEATGYGAVLFVE----NVLKDKGESLKGKRCLVS--GAGNVAQY 320
Query: 91 CVYLLLHPSPPVVETS 106
C LLL V+ S
Sbjct: 321 CAELLLEKGAIVLSLS 336
>CA16_MOUSE (Q04857) Collagen alpha 1(VI) chain precursor
Length = 1025
Score = 27.3 bits (59), Expect = 6.3
Identities = 18/63 (28%), Positives = 33/63 (51%), Gaps = 3/63 (4%)
Query: 12 EVYSVWAIPPDHVRPRIAKLMTD--LRSEFGGPHFEPHITVVGAITLTADDALNKLRSAC 69
+V+SV AI PDH+ PR++ + TD R F + I+ T D ++ +++
Sbjct: 184 KVFSV-AITPDHLEPRLSIIATDHTYRRNFTAADWGHSRDAEEVISQTIDTIVDMIKNNV 242
Query: 70 EEL 72
E++
Sbjct: 243 EQV 245
>HTR5_HALN1 (Q48318) Halobacterial transducer protein V
Length = 545
Score = 26.9 bits (58), Expect = 8.2
Identities = 13/34 (38%), Positives = 18/34 (52%)
Query: 51 VGAITLTADDALNKLRSACEELKAFQVTVDRVAA 84
V I DD +L S EE+ ++ TV+ VAA
Sbjct: 251 VQEIAAGTDDQREQLESVAEEMDSYSATVEEVAA 284
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.134 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,954,885
Number of Sequences: 164201
Number of extensions: 480537
Number of successful extensions: 1189
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1178
Number of HSP's gapped (non-prelim): 17
length of query: 118
length of database: 59,974,054
effective HSP length: 94
effective length of query: 24
effective length of database: 44,539,160
effective search space: 1068939840
effective search space used: 1068939840
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0162a.1


