

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0159.10
(353 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
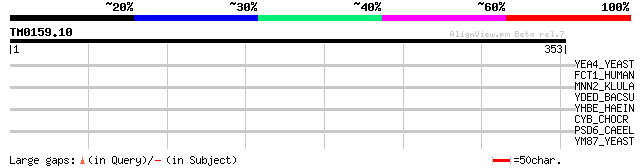
YEA4_YEAST (P40004) Hypothetical 39.3 kDa protein in GCN4-WBP1 i... 42 0.003
FCT1_HUMAN (Q96A29) GDP-fucose transporter 1 (Solute carrier fam... 40 0.009
MNN2_KLULA (Q00974) UDP-N-acetylglucosamine transporter (Golgi U... 39 0.020
YDED_BACSU (P96661) Hypothetical transport protein ydeD 38 0.044
YHBE_HAEIN (P71360) Hypothetical transport protein HI0878 35 0.29
CYB_CHOCR (P48875) Cytochrome b 32 3.2
PSD6_CAEEL (Q20585) 26S proteasome non-ATPase regulatory subunit... 30 7.0
YM87_YEAST (Q04835) Hypothetical 46.7 kDa protein in HOR7-COX7 i... 30 9.2
>YEA4_YEAST (P40004) Hypothetical 39.3 kDa protein in GCN4-WBP1
intergenic region
Length = 342
Score = 41.6 bits (96), Expect = 0.003
Identities = 49/266 (18%), Positives = 105/266 (39%), Gaps = 27/266 (10%)
Query: 91 PIYKYCLVSVSNILTTTCQYEALKY-VSFPVQTLAKCAKMIPVMIWGTIIMQKRYQGPDY 149
P++ Y + V +++T KY +S P+ + +C + M ++ ++Y
Sbjct: 69 PLHVYVITVVLFYISSTTNNNVFKYNISIPIHIVFRCFGTVITMFTCWLLNGRKYTKIQI 128
Query: 150 MLAFLVTLGCSVFILYPAGE------DISPYSRGRENTV-----WGVLLMTGYLGCDGFT 198
+ +T+G + L+ + + + G + +V +G+ ++
Sbjct: 129 LSTLFLTIGAIIASLFKDADFRYQDLKLQAWKIGSDQSVDLTFIFGICILVLSSFTSSLL 188
Query: 199 STFQDKLFRGYDMEIHNQIFYTTLCSCILSLTG---------LILQGHLIPAIEFVYRHH 249
S + ++ ++ Y IFY+ S L L ++ + I F +
Sbjct: 189 SAYNERTYQKYGKHWKENIFYSHFLSLPLFLFSRKQLIHEYRVMRKSERILCSNFGGKIL 248
Query: 250 DCFFDIALLSTVATASQFF----ISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLS 305
+ LL V T Q+F ++ ALT + + R+ +S++LS F + LS
Sbjct: 249 VPREETLLLFNVLT--QYFCVKGVNILASKTNALTLSITLLVRKFISLLLSVRLFDNNLS 306
Query: 306 WEQWIGAVIVFGSLYAKSFTRKAPQK 331
+ +IG +VF + S P++
Sbjct: 307 YTGYIGVYLVFFGAFIYSLGSIHPRQ 332
>FCT1_HUMAN (Q96A29) GDP-fucose transporter 1 (Solute carrier family
35 member C2)
Length = 364
Score = 40.0 bits (92), Expect = 0.009
Identities = 43/194 (22%), Positives = 75/194 (38%), Gaps = 11/194 (5%)
Query: 152 AFLVTLGCSVFILYPAGEDISPYSRGRENTVWGVLLMTGYLG--CDGFTSTFQDKLFRGY 209
+F L C + I G + G E T+ + + G L C + + K+
Sbjct: 167 SFYALLTCGIII---GGFWLGVDQEGAEGTLSWLGTVFGVLASLCVSLNAIYTTKVLPAV 223
Query: 210 DMEIHNQIFYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDCFFDIALLSTV-ATASQFF 268
D I FY + +CIL L L+L G L +F F+ + L + A +
Sbjct: 224 DGSIWRLTFYNNVNACILFLPLLLLLGELQALRDFAQLGSAHFWGMMTLGGLFGFAIGYV 283
Query: 269 ISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSLYAKSFTR-- 326
I+ LT T + +L+ +++ S+ W ++V G A ++ R
Sbjct: 284 TGLQIKFTSPLTHNVSGTAKACAQTVLAVLYYEETKSFLWWTSNMMVLGGSSAYTWVRGW 343
Query: 327 ---KAPQKTTSSDS 337
K P++ + DS
Sbjct: 344 EMKKTPEEPSPKDS 357
>MNN2_KLULA (Q00974) UDP-N-acetylglucosamine transporter (Golgi
UDP-GlcNAc transporter)
Length = 328
Score = 38.9 bits (89), Expect = 0.020
Identities = 47/231 (20%), Positives = 86/231 (36%), Gaps = 43/231 (18%)
Query: 116 VSFPVQTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGE------ 169
+S P+ + +C+ MI G + KRY A ++TLG V LY E
Sbjct: 94 ISVPIHIIIRCSGTTLTMIIGWAVCNKRYSKLQVQSAIIMTLGAIVASLYRDKEFSMDSL 153
Query: 170 DISPYSRG-RENTVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILS 228
++ S G + +++G+ ++ S + + +FY+ + L
Sbjct: 154 KLNTDSVGMTQKSMFGIFVVLVATALMSLLSLLNEWTYNKCGKHWKETLFYSHFLALPLF 213
Query: 229 LTGLILQGHLIPAIEFVYRHHDCFFDIALLS----------TVATASQFFISYTIRTF-- 276
+ G R D F D+ + S +AT I+ + F
Sbjct: 214 MLGYT-------------RLRDEFRDLLISSDSMDIPIVKLPIATKLFMLIANNVTQFIC 260
Query: 277 -----------GALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVF 316
ALT + ++ R+ VS++LS + + LS ++G + VF
Sbjct: 261 IKGVNMLASNTDALTLSVVLLVRKFVSLLLSVYIYKNVLSVTAYLGTITVF 311
>YDED_BACSU (P96661) Hypothetical transport protein ydeD
Length = 319
Score = 37.7 bits (86), Expect = 0.044
Identities = 56/265 (21%), Positives = 104/265 (39%), Gaps = 33/265 (12%)
Query: 69 ITTSAVSAGSLLAS----KKALDPVAPIYKY-------CLVSVSNILTTTCQYEA-LKYV 116
+TT ++AG LL +K V ++K+ + + +L Y A +++
Sbjct: 42 VTTRLLTAGMLLLMLQLFRKDRSQVIEVWKHKASAGQLLIFGLLGMLAVQYTYMASIQHG 101
Query: 117 SFPVQTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSR 176
+ V TL + + V+I+ + Q D + L +GC F L G +S S
Sbjct: 102 NAAVATLLQYLSPVIVIIYTLLRKQTVLTKQDIITVVLALVGC--FFLLTNGS-MSQLSV 158
Query: 177 GRENTVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQG 236
VWGVL GF + F + Y + + N+ + + + G L G
Sbjct: 159 PAAAVVWGVL--------SGFAAAF----YTLYAVGLLNKFDSLVVVGWAMIIGGFAL-G 205
Query: 237 HLIPAIEFVYRHHDC----FFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVS 292
+ P + ++ + +L A FFI ++ + + + + L +
Sbjct: 206 FIHPPWQLDFQRLTAEAYAYILFVILFGTMIAFWFFIK-SLESLSPKETSLLGSLEPLSA 264
Query: 293 IMLSCVWFSHPLSWEQWIGAVIVFG 317
++ + VW P QWIGA+ + G
Sbjct: 265 VVTTVVWLKAPFGSFQWIGAICIIG 289
>YHBE_HAEIN (P71360) Hypothetical transport protein HI0878
Length = 306
Score = 35.0 bits (79), Expect = 0.29
Identities = 30/130 (23%), Positives = 57/130 (43%), Gaps = 6/130 (4%)
Query: 68 RITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQY----EALKYVSFPVQTL 123
R +AVS +LLA KK L + + +Y + + ++ T + +L Y+ V +
Sbjct: 42 RFIIAAVSLLALLAYKKQLPELMKVRQYAWIMLIGVIGLTSNFLLFSSSLNYIEPSVAQI 101
Query: 124 AKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFI--LYPAGEDISPYSRGRENT 181
++I G +I +++ + FL+ +G +F + A ++ YS G
Sbjct: 102 FIHLSSFGMLICGVLIFKEKLGLHQKIGLFLLLIGLGLFFNDRFDAFAGLNQYSTGVILG 161
Query: 182 VWGVLLMTGY 191
V G L+ Y
Sbjct: 162 VGGALIWVAY 171
>CYB_CHOCR (P48875) Cytochrome b
Length = 381
Score = 31.6 bits (70), Expect = 3.2
Identities = 31/124 (25%), Positives = 53/124 (42%), Gaps = 18/124 (14%)
Query: 218 FYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDCFF-DIA---LLSTVAT--ASQFFISY 271
F +++C + LTG+ L H P ++ + + D+ LL + T AS FFI
Sbjct: 33 FLSSICLIVQILTGIFLAMHYTPHVDLAFASVEHIMRDVNYGWLLRYIHTNGASMFFIVV 92
Query: 272 TIRTFGALTFATIMTTRQ--------LVSIMLSCVWFSHPLSWEQ---WIGAVIVFGSLY 320
I F L F + + R ++ +M+ + + L W Q W GA ++ +
Sbjct: 93 YIHIFRGLYFGSYIKPRHWVWVIGVLILLLMILTAFIGYVLPWGQMSLW-GATVITNLVS 151
Query: 321 AKSF 324
A F
Sbjct: 152 AVPF 155
>PSD6_CAEEL (Q20585) 26S proteasome non-ATPase regulatory subunit 6
(26S proteasome regulatory subunit rpn-7)
Length = 410
Score = 30.4 bits (67), Expect = 7.0
Identities = 15/40 (37%), Positives = 22/40 (54%)
Query: 229 LTGLILQGHLIPAIEFVYRHHDCFFDIALLSTVATASQFF 268
LTG L G LIP E++ ++DC +D + A S+ F
Sbjct: 264 LTGGGLNGTLIPVREYLESYYDCHYDRFFIQLAALESERF 303
>YM87_YEAST (Q04835) Hypothetical 46.7 kDa protein in HOR7-COX7
intergenic region
Length = 414
Score = 30.0 bits (66), Expect = 9.2
Identities = 28/133 (21%), Positives = 58/133 (43%), Gaps = 7/133 (5%)
Query: 217 IFYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDCFFDI-ALLSTVATASQFFISYTIRT 275
+ Y +L + I+S G+ + IP+++F H + + L Q ++ I+
Sbjct: 275 VSYFSLITAIVSFIGI----NTIPSMKFQIPHSKKQWILFGNLGVSGFIFQLLLTMGIQR 330
Query: 276 FGALTFATIMTTRQLVSIMLSCVWFSH-PLSWEQWIGAVIVFGSLYAKSFTRKAPQKTTS 334
A + + T+ L ++ + H P W WIG +I+ + R A +TT+
Sbjct: 331 ERAGRGSLMTYTQLLYAVFWDVALYKHWPNIWS-WIGMIIIISATLWVIRIRAANNETTA 389
Query: 335 SDSIPLVQSGDSN 347
D P++ +++
Sbjct: 390 KDLTPIIDDEENS 402
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.137 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,172,585
Number of Sequences: 164201
Number of extensions: 1493760
Number of successful extensions: 4118
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 4114
Number of HSP's gapped (non-prelim): 9
length of query: 353
length of database: 59,974,054
effective HSP length: 111
effective length of query: 242
effective length of database: 41,747,743
effective search space: 10102953806
effective search space used: 10102953806
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0159.10


