

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0154.8
(247 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
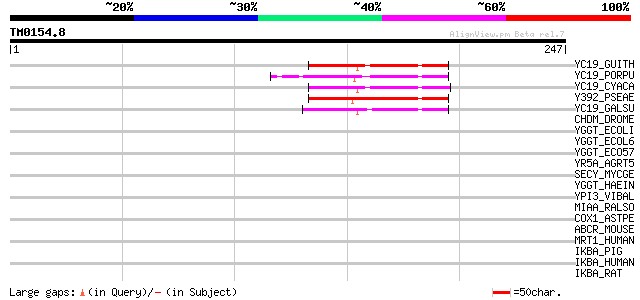
Score E
Sequences producing significant alignments: (bits) Value
YC19_GUITH (O78424) Hypothetical 10.4 kDa protein ycf19 64 3e-10
YC19_PORPU (P51353) Hypothetical 10.8 kDa protein ycf19 (ORF95) 56 7e-08
YC19_CYACA (Q9TM45) Hypothetical 10.5 kDa protein ycf19 51 3e-06
Y392_PSEAE (P25254) Hypothetical protein PA0392 49 1e-05
YC19_GALSU (P28255) Hypothetical 11.6 kDa protein ycf19 44 4e-04
CHDM_DROME (O97159) Chromodomain helicase-DNA-binding protein Mi... 36 0.075
YGGT_ECOLI (P64564) Hypothetical protein yggT 35 0.22
YGGT_ECOL6 (P64565) Hypothetical protein yggT 35 0.22
YGGT_ECO57 (P64566) Hypothetical protein yggT 35 0.22
YR5A_AGRT5 (Q8U530) Hypothetical protein Atu2659.1/AGR_C_4820 34 0.37
SECY_MYCGE (P47416) Preprotein translocase secY subunit 32 1.1
YGGT_HAEIN (P44097) Hypothetical protein HI1036 32 1.9
YPI3_VIBAL (P52059) Hypothetical protein in proC 3'region (ORF3) 31 3.2
MIAA_RALSO (Q8XWB0) tRNA delta(2)-isopentenylpyrophosphate trans... 31 3.2
COX1_ASTPE (Q33820) Cytochrome c oxidase polypeptide I (EC 1.9.3.1) 31 3.2
ABCR_MOUSE (O35600) Retinal-specific ATP-binding cassette transp... 31 3.2
MRT1_HUMAN (Q9H1U9) Mitochondrial carrier triple repeat 1 30 7.1
IKBA_PIG (Q08353) NF-kappaB inhibitor alpha (I-kappa-B-alpha) (I... 30 7.1
IKBA_HUMAN (P25963) NF-kappaB inhibitor alpha (Major histocompat... 30 7.1
IKBA_RAT (Q63746) NF-kappaB inhibitor alpha (I-kappa-B-alpha) (I... 29 9.2
>YC19_GUITH (O78424) Hypothetical 10.4 kDa protein ycf19
Length = 91
Score = 63.9 bits (154), Expect = 3e-10
Identities = 35/66 (53%), Positives = 45/66 (68%), Gaps = 7/66 (10%)
Query: 134 FLNIYNTLLVVRLVLTWFPN----SPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPI 189
FL IY LL++R+ LTWFPN P LS I DPYL +FRG++PPL G +D+SPI
Sbjct: 15 FLQIYLILLLIRVSLTWFPNVNWYGQPFY--SLSRITDPYLKMFRGIVPPLIG-IDISPI 71
Query: 190 LAFLVL 195
L F++L
Sbjct: 72 LGFILL 77
>YC19_PORPU (P51353) Hypothetical 10.8 kDa protein ycf19 (ORF95)
Length = 95
Score = 56.2 bits (134), Expect = 7e-08
Identities = 37/84 (44%), Positives = 50/84 (59%), Gaps = 12/84 (14%)
Query: 117 LPGDSVAGLVVGNGIQNFLNIYNTLLVVRLVLTWFP-----NSPPAIVGPLSTICDPYLN 171
LPG L++G+ I NF IY L++++L L WFP N P L+ I DPYL
Sbjct: 4 LPG--TLNLLLGS-IANFSEIYLILILLKLSLAWFPTVNWYNEPFC---SLNRITDPYLK 57
Query: 172 LFRGLIPPLGGTLDLSPILAFLVL 195
LFRG IPP+ G +D+SP+L + L
Sbjct: 58 LFRGSIPPMFG-MDMSPMLGIIFL 80
>YC19_CYACA (Q9TM45) Hypothetical 10.5 kDa protein ycf19
Length = 91
Score = 50.8 bits (120), Expect = 3e-06
Identities = 29/67 (43%), Positives = 40/67 (59%), Gaps = 7/67 (10%)
Query: 134 FLNIYNTLLVVRLVLTWFPN----SPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPI 189
FL IY L+++R+ L WFPN S P LS + DPYLNLF G+ P G +D SPI
Sbjct: 15 FLQIYIVLILLRMSLGWFPNINWYSQPFY--SLSQLSDPYLNLFHGVFPSFLG-IDFSPI 71
Query: 190 LAFLVLN 196
+ +++
Sbjct: 72 IGITLID 78
>Y392_PSEAE (P25254) Hypothetical protein PA0392
Length = 197
Score = 48.5 bits (114), Expect = 1e-05
Identities = 28/63 (44%), Positives = 39/63 (61%), Gaps = 2/63 (3%)
Query: 134 FLNIYNTLLVVRLVLTWF-PNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAF 192
FL ++ L++ ++L+W P S ++ IC+P L FR L+P LGG LDLSPI AF
Sbjct: 111 FLKVFFFALIISVILSWVAPGSYNPGAQLVNQICEPLLMPFRKLLPNLGG-LDLSPIFAF 169
Query: 193 LVL 195
L L
Sbjct: 170 LAL 172
>YC19_GALSU (P28255) Hypothetical 11.6 kDa protein ycf19
Length = 98
Score = 43.9 bits (102), Expect = 4e-04
Identities = 29/69 (42%), Positives = 37/69 (53%), Gaps = 7/69 (10%)
Query: 131 IQNFLNIYNTLLVVRLVLTWFPN----SPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDL 186
I F IY L +R+ L W + P IV L + DPYLNLFRG +P + G +D
Sbjct: 17 ITEFCRIYLFALSIRVFLAWIVTINWYTQPYIV--LKKLTDPYLNLFRGTLPLILG-MDF 73
Query: 187 SPILAFLVL 195
S +L FL L
Sbjct: 74 SSMLGFLFL 82
>CHDM_DROME (O97159) Chromodomain helicase-DNA-binding protein Mi-2
homolog (dMi-2)
Length = 1982
Score = 36.2 bits (82), Expect = 0.075
Identities = 21/70 (30%), Positives = 38/70 (54%), Gaps = 4/70 (5%)
Query: 178 PPLGGTLDLSPILAFLVLNAFTSASAALPAELPVTQQSEQGVAATLQSTDITSSQKKWMK 237
PP GG +D S + N+ T A++A P+ P +++ E + + D TS++K +K
Sbjct: 1538 PPTGGNVDKSATTS----NSVTPATSAAPSPAPASEKGEDKDKDSEKEKDKTSAEKSEVK 1593
Query: 238 RLQGNRENKE 247
+ Q E+K+
Sbjct: 1594 QEQEAEEDKK 1603
>YGGT_ECOLI (P64564) Hypothetical protein yggT
Length = 188
Score = 34.7 bits (78), Expect = 0.22
Identities = 18/55 (32%), Positives = 30/55 (53%), Gaps = 1/55 (1%)
Query: 141 LLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAFLVL 195
+L+V +++W I L + DP L R L+P +GG +D SP++ L+L
Sbjct: 109 VLLVMAIMSWVSQGRSPIEYVLIQLADPLLRPIRRLLPAMGG-IDFSPMILVLLL 162
>YGGT_ECOL6 (P64565) Hypothetical protein yggT
Length = 188
Score = 34.7 bits (78), Expect = 0.22
Identities = 18/55 (32%), Positives = 30/55 (53%), Gaps = 1/55 (1%)
Query: 141 LLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAFLVL 195
+L+V +++W I L + DP L R L+P +GG +D SP++ L+L
Sbjct: 109 VLLVMAIMSWVSQGRSPIEYVLIQLADPLLRPIRRLLPAMGG-IDFSPMILVLLL 162
>YGGT_ECO57 (P64566) Hypothetical protein yggT
Length = 188
Score = 34.7 bits (78), Expect = 0.22
Identities = 18/55 (32%), Positives = 30/55 (53%), Gaps = 1/55 (1%)
Query: 141 LLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAFLVL 195
+L+V +++W I L + DP L R L+P +GG +D SP++ L+L
Sbjct: 109 VLLVMAIMSWVSQGRSPIEYVLIQLADPLLRPIRRLLPAMGG-IDFSPMILVLLL 162
>YR5A_AGRT5 (Q8U530) Hypothetical protein Atu2659.1/AGR_C_4820
Length = 106
Score = 33.9 bits (76), Expect = 0.37
Identities = 20/70 (28%), Positives = 36/70 (50%), Gaps = 10/70 (14%)
Query: 135 LNIYNTLLVVRLVLTWFP-----NSPPAIVGPLST----ICDPYLNLFRGLIPPLGGTLD 185
LN+Y +L+ + +W NS V + + + +P L R ++P LGG +D
Sbjct: 22 LNLYTWVLIASAIFSWLYAFNVINSRNQFVNAIGSFLVNVTEPALRPIRRILPNLGG-ID 80
Query: 186 LSPILAFLVL 195
+SPI+ L++
Sbjct: 81 ISPIILLLII 90
>SECY_MYCGE (P47416) Preprotein translocase secY subunit
Length = 475
Score = 32.3 bits (72), Expect = 1.1
Identities = 34/119 (28%), Positives = 55/119 (45%), Gaps = 17/119 (14%)
Query: 108 LSSHNFAAVLPGDSVAGLVVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTICD 167
+++ + AV GD+++ VGNGI TLL++ +L+ P A LS I
Sbjct: 172 MTAGTYMAVFLGDTISKKGVGNGI--------TLLILSGILSQLPQGFIAAYNVLSGIVI 223
Query: 168 PYLNLFRGLIPPLGGTLD-LSPILAFLVLNAFTSASAALPAELPVTQQSEQGVAATLQS 225
L P L + LAFLVL T+ ++P+ QQS QG+ + +++
Sbjct: 224 T-------LTPQLTAAISFFIYFLAFLVLLFATTFITQATRKIPI-QQSGQGLVSEVKT 274
>YGGT_HAEIN (P44097) Hypothetical protein HI1036
Length = 181
Score = 31.6 bits (70), Expect = 1.9
Identities = 17/55 (30%), Positives = 28/55 (50%), Gaps = 1/55 (1%)
Query: 141 LLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAFLVL 195
+L + VL+WF +I + +P L R L+P L G +D SP++ +L
Sbjct: 109 VLFIGAVLSWFNRGNNSISYAFYQLSEPLLKPIRRLLPTL-GMIDFSPMVVMFIL 162
>YPI3_VIBAL (P52059) Hypothetical protein in proC 3'region (ORF3)
Length = 185
Score = 30.8 bits (68), Expect = 3.2
Identities = 17/55 (30%), Positives = 29/55 (51%), Gaps = 1/55 (1%)
Query: 141 LLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAFLVL 195
+L++R +L+W I + +P R +IP +GG LDLS ++ F+ L
Sbjct: 114 VLLIRAILSWVSQGRSPIEYVFHQLTEPMCAPIRRIIPAIGG-LDLSVLVLFIGL 167
>MIAA_RALSO (Q8XWB0) tRNA delta(2)-isopentenylpyrophosphate
transferase (EC 2.5.1.8) (IPP transferase)
(Isopentenyl-diphosphate:tRNA isopentenyltransferase)
(IPTase) (IPPT)
Length = 323
Score = 30.8 bits (68), Expect = 3.2
Identities = 18/53 (33%), Positives = 27/53 (49%), Gaps = 3/53 (5%)
Query: 185 DLSPILAFLVLNAFTSASAALPAELPVTQQSEQGVAATLQSTDITSSQKKWMK 237
DLSP+L + + A A L E+ + EQG+AAT Q + Q W++
Sbjct: 244 DLSPVLPSIRCVGYRQAWAYLDGEIDMATLREQGIAATRQ---LCKRQITWLR 293
>COX1_ASTPE (Q33820) Cytochrome c oxidase polypeptide I (EC 1.9.3.1)
Length = 517
Score = 30.8 bits (68), Expect = 3.2
Identities = 24/76 (31%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 136 NIYNTLLVVRLVLTWFPNSPPAIVGPLSTICD-PYLNLFRGLIPPLGGTLDLSPILAFLV 194
NI+ TL+ + + LT+FP + G D P + +G T+ L L FL
Sbjct: 411 NIHFTLMFIGVNLTFFPQHFLGLAGMPRRYSDYPDAYTLWNTVSSIGSTISLIATLVFLF 470
Query: 195 L--NAFTSASAALPAE 208
+ AFTS AL E
Sbjct: 471 ILWEAFTSQRTALQPE 486
>ABCR_MOUSE (O35600) Retinal-specific ATP-binding cassette
transporter (ATP-binding cassette, sub-family A, member
4) (RIM ABC transporter) (RIM protein) (RmP)
Length = 2310
Score = 30.8 bits (68), Expect = 3.2
Identities = 19/64 (29%), Positives = 32/64 (49%), Gaps = 3/64 (4%)
Query: 124 GLVVGNGIQNFLNIYNTL---LVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPL 180
GLV G ++N + ++ V RL +T++ N A +G L++ GL+PP
Sbjct: 924 GLVPGVCVKNLVKVFEPSGRPAVDRLNITFYENQITAFLGHNGAGKTTTLSILTGLLPPT 983
Query: 181 GGTL 184
GT+
Sbjct: 984 SGTV 987
>MRT1_HUMAN (Q9H1U9) Mitochondrial carrier triple repeat 1
Length = 297
Score = 29.6 bits (65), Expect = 7.1
Identities = 19/60 (31%), Positives = 30/60 (49%), Gaps = 4/60 (6%)
Query: 167 DPYLNLFRGLIPPLGGTLDLSPILAFLVLNAFTSASAALPAELPVTQQSEQGVAATLQST 226
D + NL+RG++PPL + + LA L+ + S L + + + GVAA L T
Sbjct: 75 DGFRNLYRGILPPL---MQKTTTLA-LMFGLYEDLSCLLHKHVSAPEFATSGVAAVLAGT 130
>IKBA_PIG (Q08353) NF-kappaB inhibitor alpha (I-kappa-B-alpha)
(IkappaBalpha) (IKB-alpha) (ECI-6)
Length = 314
Score = 29.6 bits (65), Expect = 7.1
Identities = 18/45 (40%), Positives = 18/45 (40%)
Query: 134 FLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIP 178
FLN N L L L N P L CDP L FRG P
Sbjct: 103 FLNFQNNLQQTPLHLAVITNQPEIAEALLEAGCDPELRDFRGNTP 147
>IKBA_HUMAN (P25963) NF-kappaB inhibitor alpha (Major
histocompatibility complex enhancer-binding protein
MAD3) (I-kappa-B-alpha) (IkappaBalpha) (IKB-alpha)
Length = 317
Score = 29.6 bits (65), Expect = 7.1
Identities = 18/45 (40%), Positives = 18/45 (40%)
Query: 134 FLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIP 178
FLN N L L L N P L CDP L FRG P
Sbjct: 103 FLNFQNNLQQTPLHLAVITNQPEIAEALLGAGCDPELRDFRGNTP 147
>IKBA_RAT (Q63746) NF-kappaB inhibitor alpha (I-kappa-B-alpha)
(IkappaBalpha) (IKB-alpha) (RL/IF-1)
Length = 314
Score = 29.3 bits (64), Expect = 9.2
Identities = 18/45 (40%), Positives = 18/45 (40%)
Query: 134 FLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIP 178
FLN N L L L N P L CDP L FRG P
Sbjct: 103 FLNFQNNLQQTPLHLAVITNQPGIAEALLKAGCDPELRDFRGNTP 147
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,783,899
Number of Sequences: 164201
Number of extensions: 1150770
Number of successful extensions: 2619
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 2605
Number of HSP's gapped (non-prelim): 25
length of query: 247
length of database: 59,974,054
effective HSP length: 107
effective length of query: 140
effective length of database: 42,404,547
effective search space: 5936636580
effective search space used: 5936636580
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0154.8


