

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
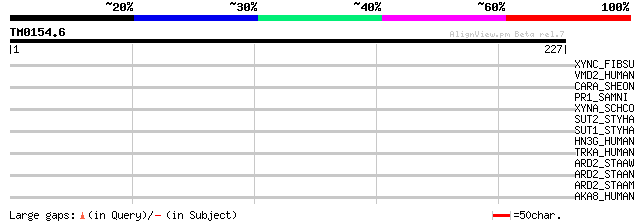
Query= TM0154.6
(227 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
XYNC_FIBSU (P35811) Endo-1,4-beta-xylanase C precursor (EC 3.2.1... 32 1.6
VMD2_HUMAN (O76090) Bestrophin 1 (Vitelliform macular dystrophy ... 32 1.6
CARA_SHEON (Q8EHS6) Carbamoyl-phosphate synthase small chain (EC... 32 1.6
PR1_SAMNI (Q41359) Pathogenesis-related protein PR-1 type precursor 30 3.6
XYNA_SCHCO (P35809) Endo-1,4-beta-xylanase A (EC 3.2.1.8) (Xylan... 30 4.7
SUT2_STYHA (P53392) High affinity sulphate transporter 2 30 4.7
SUT1_STYHA (P53391) High affinity sulphate transporter 1 30 4.7
HN3G_HUMAN (P55318) Hepatocyte nuclear factor 3-gamma (HNF-3G) (... 30 6.1
TRKA_HUMAN (P04629) High affinity nerve growth factor receptor p... 29 8.0
ARD2_STAAW (P60299) Acetylornithine aminotransferase 2 (EC 2.6.1... 29 8.0
ARD2_STAAN (P60298) Acetylornithine aminotransferase 2 (EC 2.6.1... 29 8.0
ARD2_STAAM (P60297) Acetylornithine aminotransferase 2 (EC 2.6.1... 29 8.0
AKA8_HUMAN (O43823) A-kinase anchor protein 8 (A-kinase anchor p... 29 8.0
>XYNC_FIBSU (P35811) Endo-1,4-beta-xylanase C precursor (EC 3.2.1.8)
(Xylanase C) (1,4-beta-D-xylan xylanohydrolase C)
Length = 608
Score = 31.6 bits (70), Expect = 1.6
Identities = 17/55 (30%), Positives = 30/55 (53%), Gaps = 5/55 (9%)
Query: 142 SKNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFEGGVAKLTKPGCFSGAND 196
++ Q ++VTG G SPY + + + GG + TF ++ G K + ++G ND
Sbjct: 309 TRTQGQNNSSVTGNVGSSPYHYEIWYQGGNNSMTF-YDNGTYKAS----WNGTND 358
>VMD2_HUMAN (O76090) Bestrophin 1 (Vitelliform macular dystrophy
protein 2) (TU15B)
Length = 585
Score = 31.6 bits (70), Expect = 1.6
Identities = 22/66 (33%), Positives = 29/66 (43%), Gaps = 3/66 (4%)
Query: 94 FAPNCGVKASSLAPITGRFLGCQTKYVHAPEAFSEVLIQNQRSLEILHS---KNHTQVGA 150
F PN + + A I GRFLG Q+ H P A S + + +LH KNH
Sbjct: 375 FQPNQEDEEDAHAGIIGRFLGLQSHDHHPPRANSRTKLLWPKRESLLHEGLPKNHKAAKQ 434
Query: 151 AVTGTD 156
V G +
Sbjct: 435 NVRGQE 440
>CARA_SHEON (Q8EHS6) Carbamoyl-phosphate synthase small chain (EC
6.3.5.5) (Carbamoyl-phosphate synthetase glutamine
chain)
Length = 386
Score = 31.6 bits (70), Expect = 1.6
Identities = 26/69 (37%), Positives = 32/69 (45%), Gaps = 5/69 (7%)
Query: 68 KAY---QGDCGAVGGPDAKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYVHAPE 124
KAY +G VGG A P E+ +V A + GVK + L + R GC V A
Sbjct: 170 KAYPWRKGSWRLVGGLPADTPAEALKYKVVAYDYGVKQNILRMLVDR--GCDVTVVPAKT 227
Query: 125 AFSEVLIQN 133
SEVL N
Sbjct: 228 PASEVLAMN 236
>PR1_SAMNI (Q41359) Pathogenesis-related protein PR-1 type
precursor
Length = 167
Score = 30.4 bits (67), Expect = 3.6
Identities = 21/74 (28%), Positives = 28/74 (37%)
Query: 21 AATNTQIKVTNNPADKLVAAINENRTAYKVSELYDNAGLACIALQYIKAYQGDCGAVGGP 80
A Q V N V A N R+A V + + +A A QY ++ GDC V
Sbjct: 19 ALVMVQYSVAQNSPQDYVDAHNAARSAVNVGPVTWDESVAAFARQYAQSRAGDCRLVHSG 78
Query: 81 DAKKPPESQFAEVF 94
D + F F
Sbjct: 79 DPRYGENLAFGSGF 92
>XYNA_SCHCO (P35809) Endo-1,4-beta-xylanase A (EC 3.2.1.8) (Xylanase
A) (1,4-beta-D-xylan xylanohydrolase A)
Length = 197
Score = 30.0 bits (66), Expect = 4.7
Identities = 18/54 (33%), Positives = 26/54 (47%), Gaps = 6/54 (11%)
Query: 153 TGTDGGSPYFWCVLFSGGKPNSTFAFEGGVAKLTKPGCFSGANDECSGAHDWSP 206
TGTDGG Y W ++ G ++T+ GG + +SG N G W+P
Sbjct: 7 TGTDGGYYYSW---WTDGAGDATYQNNGGGSYTL---TWSGNNGNLVGGKGWNP 54
>SUT2_STYHA (P53392) High affinity sulphate transporter 2
Length = 662
Score = 30.0 bits (66), Expect = 4.7
Identities = 18/60 (30%), Positives = 29/60 (48%)
Query: 96 PNCGVKASSLAPITGRFLGCQTKYVHAPEAFSEVLIQNQRSLEILHSKNHTQVGAAVTGT 155
P G+ +S +AP+ F+G P A +L+ S EI ++K+H + A T T
Sbjct: 130 PWYGLYSSFVAPLVYAFMGTSRDIAIGPVAVVSLLLGTLLSNEISNTKSHDYLRLAFTAT 189
>SUT1_STYHA (P53391) High affinity sulphate transporter 1
Length = 667
Score = 30.0 bits (66), Expect = 4.7
Identities = 18/60 (30%), Positives = 29/60 (48%)
Query: 96 PNCGVKASSLAPITGRFLGCQTKYVHAPEAFSEVLIQNQRSLEILHSKNHTQVGAAVTGT 155
P G+ +S +AP+ F+G P A +L+ S EI ++K+H + A T T
Sbjct: 133 PWYGLYSSFVAPLVYAFMGTSRDIAIGPVAVVSLLLGTLLSNEISNTKSHDYLRLAFTAT 192
>HN3G_HUMAN (P55318) Hepatocyte nuclear factor 3-gamma (HNF-3G)
(Forkhead box protein A3) (Fork head-related protein FKH
H3)
Length = 350
Score = 29.6 bits (65), Expect = 6.1
Identities = 25/81 (30%), Positives = 33/81 (39%), Gaps = 1/81 (1%)
Query: 80 PDAKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYVHAPEAFSEVLIQNQRSLEI 139
P A +P +V A +CG ASS TG L + K + AP F+ N E
Sbjct: 255 PPAPEPEAQGGEDVGALDCGSPASSTPYFTGLELPGELK-LDAPYNFNHPFSINNLMSEQ 313
Query: 140 LHSKNHTQVGAAVTGTDGGSP 160
+ VG G +GG P
Sbjct: 314 TPAPPKLDVGFGGYGAEGGEP 334
>TRKA_HUMAN (P04629) High affinity nerve growth factor receptor
precursor (EC 2.7.1.112) (TRK1 transforming tyrosine
kinase protein) (p140-TrkA) (Trk-A)
Length = 796
Score = 29.3 bits (64), Expect = 8.0
Identities = 23/103 (22%), Positives = 41/103 (39%), Gaps = 7/103 (6%)
Query: 43 ENRTAYKVSELYDNAGLACIALQYIKAYQGDCG-----AVGGPDAKKPPESQFAEVFAPN 97
++ T + VS A AC+ L + CG + P P + +
Sbjct: 411 KDETPFGVSVAVGLAVFACLFLSTLLLVLNKCGRRNKFGINRPAVLAPEDGLAMSLHFMT 470
Query: 98 CGVKASSLAPITGRFLGCQTKYVHAPEAFSEVLIQNQRSLEIL 140
G SSL+P G+ G Q + P+ FS+ + + + +I+
Sbjct: 471 LG--GSSLSPTEGKGSGLQGHIIENPQYFSDACVHHIKRRDIV 511
>ARD2_STAAW (P60299) Acetylornithine aminotransferase 2 (EC
2.6.1.11) (ACOAT 2)
Length = 396
Score = 29.3 bits (64), Expect = 8.0
Identities = 18/47 (38%), Positives = 22/47 (46%)
Query: 28 KVTNNPADKLVAAINENRTAYKVSELYDNAGLACIALQYIKAYQGDC 74
KV D L AAINEN A V + AG+ Y+KA + C
Sbjct: 170 KVDFGDVDALKAAINENTAAVLVEPIQGEAGINIPPEGYLKAIRELC 216
>ARD2_STAAN (P60298) Acetylornithine aminotransferase 2 (EC
2.6.1.11) (ACOAT 2)
Length = 396
Score = 29.3 bits (64), Expect = 8.0
Identities = 18/47 (38%), Positives = 22/47 (46%)
Query: 28 KVTNNPADKLVAAINENRTAYKVSELYDNAGLACIALQYIKAYQGDC 74
KV D L AAINEN A V + AG+ Y+KA + C
Sbjct: 170 KVDFGDVDALKAAINENTAAVLVEPIQGEAGINIPPEGYLKAIRELC 216
>ARD2_STAAM (P60297) Acetylornithine aminotransferase 2 (EC
2.6.1.11) (ACOAT 2)
Length = 396
Score = 29.3 bits (64), Expect = 8.0
Identities = 18/47 (38%), Positives = 22/47 (46%)
Query: 28 KVTNNPADKLVAAINENRTAYKVSELYDNAGLACIALQYIKAYQGDC 74
KV D L AAINEN A V + AG+ Y+KA + C
Sbjct: 170 KVDFGDVDALKAAINENTAAVLVEPIQGEAGINIPPEGYLKAIRELC 216
>AKA8_HUMAN (O43823) A-kinase anchor protein 8 (A-kinase anchor
protein 95 kDa) (AKAP 95)
Length = 692
Score = 29.3 bits (64), Expect = 8.0
Identities = 14/31 (45%), Positives = 19/31 (61%), Gaps = 1/31 (3%)
Query: 70 YQGDCGAVGGPDAKKPPESQFAEVFAPNCGV 100
Y G G +GGP +PP S F++ AP+ GV
Sbjct: 229 YVGGRG-LGGPSPSRPPPSLFSQSMAPDYGV 258
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,548,317
Number of Sequences: 164201
Number of extensions: 1170682
Number of successful extensions: 2275
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 2266
Number of HSP's gapped (non-prelim): 15
length of query: 227
length of database: 59,974,054
effective HSP length: 106
effective length of query: 121
effective length of database: 42,568,748
effective search space: 5150818508
effective search space used: 5150818508
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0154.6


