

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0154.14
(906 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
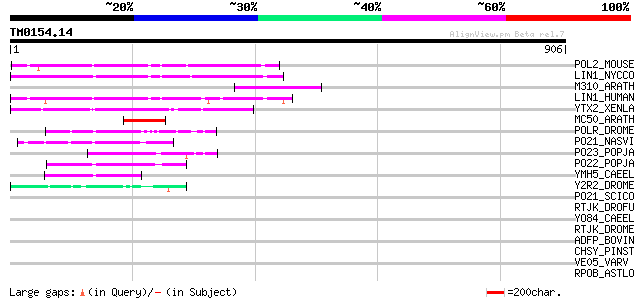
Sequences producing significant alignments: (bits) Value
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 149 3e-35
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 125 7e-28
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 121 7e-27
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 120 2e-26
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 118 6e-26
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 83 4e-15
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 71 1e-11
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 65 1e-09
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 60 3e-08
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 55 6e-07
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 49 6e-05
Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I retro... 45 7e-04
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 44 0.001
RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile elem... 38 0.11
YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III 34 1.5
RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile elem... 34 1.5
ADFP_BOVIN (Q9TUM6) Adipophilin (Adipose differentiation-related... 33 4.5
CHSY_PINST (O65872) Chalcone synthase (EC 2.3.1.74) (Naringenin-... 32 5.8
VE05_VARV (Q01483) Protein E5 32 7.6
RPOB_ASTLO (P27059) DNA-directed RNA polymerase beta chain (EC 2... 32 7.6
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 149 bits (376), Expect = 3e-35
Identities = 130/449 (28%), Positives = 205/449 (44%), Gaps = 31/449 (6%)
Query: 4 EALFQMHPTK-APGCDGLPALFYQKFWHIIGDDVSHFCLQVLH-----GDIS*SIINKTL 57
EA+ PTK +PG DG A FYQ F +D+ ++ H G + S T
Sbjct: 485 EAVINSLPTKKSPGPDGFSAEFYQTF----KEDLIPILHKLFHKIEVEGTLPNSFYEAT- 539
Query: 58 LVLIPKVKK-PMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLIT 116
+ LIPK +K P FR ISL N+ KI+ K LANR++ + +I Q F+PG
Sbjct: 540 ITLIPKPQKDPTKIENFRPISLMNIDAKILNKILANRIQEHIKAIIHPDQVGFIPGMQGW 599
Query: 117 DNALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIM 176
N + HY+ K M + LD KA+D+++ PF+ VL++ G ++++I
Sbjct: 600 FNIRKSINVIHYINKLKD--KNHMIISLDAEKAFDKIQHPFMIKVLERSGIQGPYLNMIK 657
Query: 177 TCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGI 236
S +I +NG E G RQG PLSPYLF + EV + I + + GI
Sbjct: 658 AIYSKPVANIKVNGEKLEAIPLKSGTRQGCPLSPYLFNIVLEVLARAIRQ---QKEIKGI 714
Query: 237 KIARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNV 296
+I + IS L ADD +++ + ++ S+ + G IN +KS ++
Sbjct: 715 QIGKEEVKIS--LLADDMIVYISDPKNSTRELLNLINSFGEVVGYKINSNKS-MAFLYTK 771
Query: 297 PDDCFHELRQLLTVKAVESYDKYLGLPTIIGKSK---TQIFNFVKERVWKKLKGWKERSL 353
E+R+ V + KYLG+ T+ + K + F +K+ + + L+ WK+
Sbjct: 772 NKQAEKEIRETTPFSIVTNNIKYLGV-TLTKEVKDLYDKNFKSLKKEIKEDLRRWKDLPC 830
Query: 354 YRAGREVLVK--AVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWK 411
GR +VK + AI + +P N++EG I +F W R L K
Sbjct: 831 SWIGRINIVKMAILPKAIYRFNAIPIKIPTQFFNELEGAICKFVWNNKKPRIAKSLLKDK 890
Query: 412 DLCKAKDNGRLGFRDFKMFNSALVAKNWW 440
+ +G + D K++ A+V K W
Sbjct: 891 -----RTSGGITMPDLKLYYRAIVIKTAW 914
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 125 bits (313), Expect = 7e-28
Identities = 112/452 (24%), Positives = 191/452 (41%), Gaps = 18/452 (3%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ + + K+PG DG + FYQ F + + + + I + + + L
Sbjct: 456 EIASTIQNLPKKKSPGPDGFTSEFYQTFKEELVPILLNLFQNIEKEGILPNTFYEANITL 515
Query: 61 IPKV-KKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNA 119
IPK K P +R ISL N+ KI+ K L NR++ + +I Q F+PG N
Sbjct: 516 IPKPGKDPTRKENYRPISLMNIDAKILNKILTNRIQQHIKKIIHHDQVGFIPGSQGWFNI 575
Query: 120 LVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCV 179
+ ++ K + + M L +D KA+D ++ PF+ L+++G +++ LI
Sbjct: 576 RKSINVIQHINKLKNKDH--MILSIDAEKAFDNIQHPFMIRTLKKIGIEGTFLKLIEAIY 633
Query: 180 STVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIA 239
S +I+LNG + F G RQG PLSP LF + EV + I + ++ GI I
Sbjct: 634 SKPTANIILNGVKLKSFPLRSGTRQGCPLSPLLFNIVMEVLAIAIRE---EKAIKGIHIG 690
Query: 240 RTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDD 299
+S LFADD +++ T + ++ Y SG IN KS N +
Sbjct: 691 SEEIKLS--LFADDMIVYLENTRDSTTKLLEVIKEYSNVSGYKINTHKSVAFIYTN-NNQ 747
Query: 300 CFHELRQLLTVKAVESYDKYLG--LPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAG 357
++ + V KYLG L + + + +++ + + + WK G
Sbjct: 748 AEKTVKDSIPFTVVPKKMKYLGVYLTKDVKDLYKENYETLRKEIAEDVNKWKNIPCSWLG 807
Query: 358 REVLVK--AVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCK 415
R +VK + AI ++ P +E +I F W + LS K+
Sbjct: 808 RINIVKMSILPKAIYNFNAIPIKAPLSYFKDLEKIILHFIWNQKKPQIAKTLLSNKNKA- 866
Query: 416 AKDNGRLGFRDFKMFNSALVAKNWWRIKSNPE 447
G + D +++ ++V K W N E
Sbjct: 867 ----GGITLPDLRLYYKSIVIKTAWYWHKNRE 894
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 121 bits (304), Expect = 7e-27
Identities = 57/143 (39%), Positives = 86/143 (59%), Gaps = 1/143 (0%)
Query: 368 AIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAK-DNGRLGFRD 426
A+P Y MSCF L LC ++ ++ F+W +R + ++W+ LCK+K D+G LGFRD
Sbjct: 2 ALPVYAMSCFRLSKLLCKKLTSAMTEFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRD 61
Query: 427 FKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSIMRSGEI 486
FN AL+AK +RI P +L+ ++ R+ YFP ++ G RPSYAW SI+ E+
Sbjct: 62 LGWFNQALLAKQSFRIIHQPHTLLSRLLRSRYFPHSSMMECSVGTRPSYAWRSIIHGREL 121
Query: 487 FATGGMWNIGNGKSVKAWSDKWV 509
+ G + IG+G K W D+W+
Sbjct: 122 LSRGLLRTIGDGIHTKVWLDRWI 144
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 120 bits (300), Expect = 2e-26
Identities = 116/487 (23%), Positives = 207/487 (41%), Gaps = 50/487 (10%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTL--- 57
++E + + K+PG +G A FYQ++ +++ F L++ I+ +
Sbjct: 456 EIEAIINSLPNKKSPGPEGFTAEFYQRY----KEELVPFLLKLFQSIEKEGILPNSFYEA 511
Query: 58 -LVLIPKVKKPMHANQ-FRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLI 115
++LIPK + + FR ISL N+ KI+ K LAN+++ + LI Q F+P
Sbjct: 512 SIILIPKPGRDTTKKENFRPISLMNIDAKILNKILANQIQQHIKKLIHHDQVGFIPAMQG 571
Query: 116 TDNALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLI 175
N + ++ + M + +D KA+D+++ PF+ L ++G +++ +I
Sbjct: 572 WFNIRKSINIIQHINRTKD--TNHMIISIDAEKAFDKIQQPFMLKPLNKLGIDGTYLKII 629
Query: 176 MTCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHG 235
+I+LNG E G RQG PLSP L + EV + I + + G
Sbjct: 630 RAIYDKPTANIILNGQKLEAPPLKTGTRQGCPLSPLLPNIVLEVLARAIRQ---EKEIKG 686
Query: 236 IKIARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSK---LSC 292
I++ + +S LFADD +++ + A ++ +++++ + SG IN+ KS+ +
Sbjct: 687 IQLGKEEVKLS--LFADDMIVYLENPIVSAQNLLKLISNFSKVSGYKINVQKSQAFLYTN 744
Query: 293 SRNVPDDCFHELRQLLTVKAVESYDKYLGL------PTIIGKSKTQIFNFVKERVWKKLK 346
+R EL + K + KYLG+ + ++ + N +KE K
Sbjct: 745 NRQTESQIMSELPFTIASKRI----KYLGIQLTRDVKDLFKENYKPLLNEIKEDTNK--- 797
Query: 347 GWKERSLYRAGREVLVK--AVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRG 404
WK GR +VK + I + LP ++E +F W
Sbjct: 798 -WKNIPCSWVGRINIVKMAILPKVIYRFNAIPIKLPMTFFTELEKTTLKFIWNQKRAHIA 856
Query: 405 MHCLSWKDLCKAKDNGRLGFRDFKMFNSALVAKNWWRIKSN----------PESLMGKVY 454
LS K+ G + DFK++ A V K W N P +M +Y
Sbjct: 857 KSTLSQKNKA-----GGITLPDFKLYYKATVTKTAWYWYQNRDIDQWNRTEPSEIMPHIY 911
Query: 455 RAVYFPR 461
+ F +
Sbjct: 912 NYLIFDK 918
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 118 bits (296), Expect = 6e-26
Identities = 110/404 (27%), Positives = 185/404 (45%), Gaps = 20/404 (4%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLH-GDIS*SIINKTLLV 59
++ +AL M K+PG DGL F+Q FW +G D + G++ S + +L
Sbjct: 453 ELSQALRLMPHNKSPGLDGLTIEFFQFFWDTLGPDFHRVLTEAFKKGELPLSC-RRAVLS 511
Query: 60 LIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNA 119
L+PK +R +SL + +KI+ K ++ RLK +L ++I Q+ VPGR I DN
Sbjct: 512 LLPKKGDLRLIKNWRPVSLLSTDYKIVAKAISLRLKSVLAEVIHPDQSYTVPGRTIFDNV 571
Query: 120 LVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCV 179
+ + H+ ++ +G + + L LD KA+DRV+ +L LQ F +V + T
Sbjct: 572 FLIRDLLHFARR--TGLS-LAFLSLDQEKAFDRVDHQYLIGTLQAYSFGPQFVGYLKTMY 628
Query: 180 STVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLH--GIK 237
++ + + +N + P RG+RQG PLS L+ L E F L+ K + L ++
Sbjct: 629 ASAECLVKINWSLTAPLAFGRGVRQGCPLSGQLYSLAIEPFLCLLRKRLTGLVLKEPDMR 688
Query: 238 IARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVP 297
+ +A +L A D V RA QE + Y AS IN KS ++
Sbjct: 689 VVLSAYADDVILVAQDLVDLERA--QECQEV------YAAASSARINWSKSSGLLEGSLK 740
Query: 298 DDCFHELRQLLTVKAVESYDKYLGLPTIIGK-SKTQIFNFVKERVWKKLKGWK--ERSLY 354
D + ++ ++ KYLG+ + +Q F ++E V +L WK + L
Sbjct: 741 VDFLPPAFRDISWES--KIIKYLGVYLSAEEYPVSQNFIELEECVLTRLGKWKGFAKVLS 798
Query: 355 RAGREVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGG 398
GR +++ + + Y + C +I+ + F W G
Sbjct: 799 MRGRALVINQLVASQIWYRLICLSPTQEFIAKIQRRLLDFLWIG 842
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 82.8 bits (203), Expect = 4e-15
Identities = 40/68 (58%), Positives = 49/68 (71%)
Query: 187 MLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIARTAPVIS 246
++NGAPQ P RGLRQGDPLSPYLFILC EV S L +A L GI+++ +P I+
Sbjct: 13 IINGAPQGLVTPSRGLRQGDPLSPYLFILCTEVLSGLCRRAQEQGRLPGIRVSNNSPRIN 72
Query: 247 HLLFADDS 254
HLLFADD+
Sbjct: 73 HLLFADDT 80
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 71.2 bits (173), Expect = 1e-11
Identities = 75/279 (26%), Positives = 115/279 (40%), Gaps = 23/279 (8%)
Query: 59 VLIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDN 118
V IPK FR IS+ +V+ + + LA RL + Q F+P DN
Sbjct: 401 VFIPKTVTAKRPQDFRPISVPSVLVRQLNAILATRLNSSIN--WDPRQRGFLPTDGCADN 458
Query: 119 ALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTC 178
A + + K C LD+SKA+D + + L+ G P +V +
Sbjct: 459 ATIVDLVLRHSHKHFRSC---YIANLDVSKAFDSLSHASIYDTLRAYGAPKGFVDYVQNT 515
Query: 179 VSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKI 238
S+ +G E F P RG++QGDPLSP LF L V L+ L S + G K+
Sbjct: 516 YEGGGTSLNGDGWSSEEFVPARGVKQGDPLSPILFNL---VMDRLLR--TLPSEI-GAKV 569
Query: 239 ARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPD 298
+ + FADD V+FA + + + L + G +N DK + P
Sbjct: 570 GNA--ITNAAAFADDLVLFAETRMGLQVLLDKTL-DFLSIVGLKLNADKCFTVGIKGQP- 625
Query: 299 DCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFV 337
+Q TV +S+ Y+G I +T + ++
Sbjct: 626 ------KQKCTVLEAQSF--YVGSSEIPSLKRTDEWKYL 656
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 64.7 bits (156), Expect = 1e-09
Identities = 72/257 (28%), Positives = 108/257 (42%), Gaps = 21/257 (8%)
Query: 14 APGCDGLPALFYQKFWHIIGDDV-SHFCLQVLHGDIS*SIINKTLLVLIPKVKKPMHANQ 72
A G DG+ W+ I + + S F + + HG ++ VLIPK M
Sbjct: 334 AAGPDGMTTTA----WNSIDECIKSLFNMIMYHGQCPRRYLDSRT-VLIPKEPGTMDPAC 388
Query: 73 FRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNALVAFECFHYMKKK 132
FR +S+ +V + + LANR+ L+ Q AF+ + +N + + K
Sbjct: 389 FRPLSIASVALRHFHRILANRIGE--HGLLDTRQRAFIVADGVAENTSLLSAMIKEARMK 446
Query: 133 ISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVSTVDFSIMLNGAP 192
I G + LD+ KA+D VE + L++ P + IM + +
Sbjct: 447 IKG---LYIAILDVKKAFDSVEHRSILDALRRKKLPLEMRNYIMWVYRNSKTRLEVVKTK 503
Query: 193 QEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIARTAPVISHLLFAD 252
+P RG+RQGDPLSP LF + AVL A I L+FAD
Sbjct: 504 GRWIRPARGVRQGDPLSPLLF--------NCVMDAVLRRLPENTGFLMGAEKIGALVFAD 555
Query: 253 DSVIFA--RATVQEALS 267
D V+ A R +Q +LS
Sbjct: 556 DLVLLAETREGLQASLS 572
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 59.7 bits (143), Expect = 3e-08
Identities = 57/223 (25%), Positives = 98/223 (43%), Gaps = 21/223 (9%)
Query: 127 HYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVSTVDFSI 186
HY+K + + LD+ KA+D V P + ++ G IM+ ++ +I
Sbjct: 11 HYIKLRRLKGKTYNVVSLDIRKAFDTVSHPAILRAMRAFGIDDGMQDFIMSTITDAYTNI 70
Query: 187 MLNGAPQEPFQPHRGLRQGDPLSPYLF-ILCGEVFSALINKAVLASSLHGIKIARTAPVI 245
++ G G++QGDPLSP LF I+ E+ + L ++ AS KIA
Sbjct: 71 VVGGRTTNKIYIRNGVKQGDPLSPVLFNIVLDELVTRLNDEQPGASMTPACKIA------ 124
Query: 246 SHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDK---------SKLSCSRNV 296
L FADD ++ + S+ A +Y + G +N +K S S R+
Sbjct: 125 -SLAFADDLLLLEDRDIDVPNSL-ATTCAYFRTRGMTLNPEKCASISAATVSGRSVPRSK 182
Query: 297 PDDCFHELRQLLTVKAVESYDKYLGLP-TIIGKSKTQIFNFVK 338
P + R + + + ++ KYLGL + G +K ++N +
Sbjct: 183 PSFTI-DGRYIKPLGGINTF-KYLGLTFSSTGVAKPTVYNLTR 223
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 55.5 bits (132), Expect = 6e-07
Identities = 57/231 (24%), Positives = 106/231 (45%), Gaps = 15/231 (6%)
Query: 60 LIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNA 119
LIPK + + +R I++ + + +++ + LA RL+ + + A + G L+
Sbjct: 63 LIPKDGDLENPSNWRPITIASALQRLLHRILAKRLEAAVELHPAQKGYARIDGTLVNSLL 122
Query: 120 LVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCV 179
L + ++K + LD+ KA+D V + LQ++G + I +
Sbjct: 123 LDTYISSRREQRKTYN-----VVSLDVRKAFDTVSHSSICRALQRLGIDEGTSNYITGSL 177
Query: 180 STVDFSIMLN-GAPQEPFQPHRGLRQGDPLSPYLF-ILCGEVFSALINKAVLASSLHGIK 237
S +I + G+ RG++QGDPLSP+LF + E+ +L + + ++ K
Sbjct: 178 SDSTTTIRVGPGSQTRKICIRRGVKQGDPLSPFLFNAVLDELLCSLQSTPGIGGTIGEEK 237
Query: 238 IARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKS 288
I PV L FADD ++ V ++ A +A++ + G +N KS
Sbjct: 238 I----PV---LAFADDLLLLEDNDVLLPTTL-ATVANFFRLRGMSLNAKKS 280
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 48.9 bits (115), Expect = 6e-05
Identities = 39/160 (24%), Positives = 67/160 (41%), Gaps = 4/160 (2%)
Query: 57 LLVLIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLIT 116
+++ IPK P + +R ISL + +I+ + + +R++ L+ Q F+ R
Sbjct: 669 VIIPIPKKGNPSSPSNYRPISLTDPFARIMERIICSRIRSEYSHLLSPHQHGFLNFRSCP 728
Query: 117 DNALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIM 176
+ + + +H + K + L D +KA+D+V P L L G S
Sbjct: 729 SSLVRSISLYHSILKNEKSLD---ILFFDFAKAFDKVSHPILLKKLALFGLDKLTCSWFK 785
Query: 177 TCVSTVDFSIMLNGAPQEPFQP-HRGLRQGDPLSPYLFIL 215
+ FS+ +N P G+ QG P LFIL
Sbjct: 786 EFLHLRTFSVKINKFVSSNAYPISSGVPQGSVSGPLLFIL 825
>Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I
retrotransposable element R1DM (ORF 2)
Length = 1021
Score = 45.4 bits (106), Expect = 7e-04
Identities = 60/300 (20%), Positives = 119/300 (39%), Gaps = 43/300 (14%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHF---CLQVLHGDIS*SIINKTL 57
+V+ + ++ ++PG DG+ + W I + ++ C+++ +
Sbjct: 438 EVDTCVARLKSRRSPGLDGINGTICKAVWRAIPEHLASLFSRCIRLGYFPAEWKCPRVVS 497
Query: 58 LVLIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITD 117
L+ P K + +R I L V K++ + NR++ +LP+ C Q F GR + D
Sbjct: 498 LLKGPD-KDKCEPSSYRGICLLPVFGKVLEAIMVNRVREVLPEG-CRWQFGFRQGRCVED 555
Query: 118 NALVAFECFHYMKKKI--SGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGF--PASWVS 173
+ ++K + S V+ +D A+D VEW S L +G W S
Sbjct: 556 -------AWRHVKSSVGASAAQYVLGTFVDFKGAFDNVEWSAALSRLADLGCREMGLWQS 608
Query: 174 LIMTCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSL 233
+ + S +G + P RG QG P+++ + L
Sbjct: 609 FFSGRRAVIRSS---SGTVEVPVT--RGCPQGSISGPFIWDI-----------------L 646
Query: 234 HGIKIARTAPVISHLLFADDSVIF----ARATVQE-ALSIKAILASYEQASGQVINLDKS 288
+ + R P +ADD ++ +RA ++E + +I+ ++ G ++ K+
Sbjct: 647 MDVLLQRLQPYCQLSAYADDLLLLVEGNSRAVLEEKGAQLMSIVETWGAEVGDCLSTSKT 706
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 44.3 bits (103), Expect = 0.001
Identities = 57/210 (27%), Positives = 86/210 (40%), Gaps = 17/210 (8%)
Query: 13 KAPGCDGLPALFYQKFWHIIGDDVSH--FCLQVLHGDIS*SIINKTLLVLIPKVKKPMHA 70
K G DG+ + W+ + D + + V +G + S I + V PK++
Sbjct: 173 KGAGPDGVTP----RSWNALDDRYKRLLYNIFVFYGRVP-SPIKGSRTVFTPKIEGGPDP 227
Query: 71 NQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVP--GRLITDNALVAFECFHY 128
FR +S+C+VI + K LA R + E QTA++P G I + L A
Sbjct: 228 GVFRPLSICSVILREFNKILARR--FVSCYTYDERQTAYLPIDGVCINVSMLTAIIAEAK 285
Query: 129 MKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVSTVDFSIML 188
+K + LD+ KA++ V L + + G P V I + V +
Sbjct: 286 RLRK-----ELHIAILDLVKAFNSVYHSALIDAITEAGCPPGVVDYIADMYNNVITEMQF 340
Query: 189 NGAPQEPFQPHRGLRQGDPLSPYLFILCGE 218
G E G+ QGDPLS LF L E
Sbjct: 341 EG-KCELASILAGVYQGDPLSGPLFTLAYE 369
>RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 38.1 bits (87), Expect = 0.11
Identities = 58/277 (20%), Positives = 114/277 (40%), Gaps = 44/277 (15%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINK----- 55
+V+E + ++ P KAPG D L + ++ D + + + + + K
Sbjct: 442 EVKELVSKLKPKKAPGED----LLDNRTIRLLPDQALLYLVLIFNSILRVGYFPKARPTA 497
Query: 56 TLLVLIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVP---- 111
++++++ K+P+ + +R SL + K++ + + NR+ L E T +P
Sbjct: 498 SIIMILKPGKQPLDVDSYRPTSLLPSLGKMLERLILNRI------LTSEEVTRAIPKFQF 551
Query: 112 ------GRLITDNALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQM 165
G + +V F KK+ +G + LD+ +A+DRV P L + +
Sbjct: 552 GFRLQHGTPEQLHRVVNFALEALEKKEYAG-----SCFLDIQQAFDRVWHPGLLYKAKSL 606
Query: 166 GFPASWVSLIMTCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSA-LI 224
P + LI + FS+ +G G+ QG L P L+ +F+A +
Sbjct: 607 LSPQLF-QLIKSFWEGRKFSVTADGCRSSVKFIEAGVPQGSVLGPTLY----SIFTADMP 661
Query: 225 NKAVLASSLHGIKIARTAPVISHLLFADDSVIFARAT 261
N+ + G + T +ADD + ++T
Sbjct: 662 NQNAVTGLAEGEVLIAT--------YADDIAVLTKST 690
>YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III
Length = 364
Score = 34.3 bits (77), Expect = 1.5
Identities = 23/90 (25%), Positives = 36/90 (39%)
Query: 124 ECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVSTVD 183
EC + K I LD SKA+D+V L L + + + ++
Sbjct: 3 ECMNNWTKNIDEGAQTDVKYLDFSKAFDKVSHDILLDKLTSIKINKHLIRWLDVFLTNRS 62
Query: 184 FSIMLNGAPQEPFQPHRGLRQGDPLSPYLF 213
F + + EP + G+ QG +SP LF
Sbjct: 63 FKVKVGNTLSEPKKTVCGVPQGSVISPVLF 92
>RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 34.3 bits (77), Expect = 1.5
Identities = 46/223 (20%), Positives = 88/223 (38%), Gaps = 21/223 (9%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTL--- 57
+V+ + ++ KAPG D L + ++ D F + + + K
Sbjct: 442 EVKNLIAKLPLKKAPGED----LLDNRTIRLLPDQALQFLALIFNSVLDVGYFPKAWKSA 497
Query: 58 -LVLIPKV-KKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPG--- 112
+++I K K P + +R SL + KI+ + + NRL L C+ T +P
Sbjct: 498 SIIMIHKTGKTPTDVDSYRPTSLLPSLGKIMERLILNRL------LTCKDVTKAIPKFQF 551
Query: 113 --RLITDNALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPAS 170
RL ++ + + + LD+ +A+DRV P L +++ FP
Sbjct: 552 GFRLQHGTPEQLHRVVNFALEAMENKEYAVGAFLDIQQAFDRVWHPGLLYKAKRL-FPPQ 610
Query: 171 WVSLIMTCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLF 213
++ + + F + ++G G+ QG L P L+
Sbjct: 611 LYLVVKSFLEERTFHVSVDGYKSSIKPIAAGVPQGSVLGPTLY 653
>ADFP_BOVIN (Q9TUM6) Adipophilin (Adipose differentiation-related
protein) (ADRP)
Length = 450
Score = 32.7 bits (73), Expect = 4.5
Identities = 25/89 (28%), Positives = 36/89 (40%), Gaps = 29/89 (32%)
Query: 695 LGAFLTLLYAIWMARNELCFKEKAS---------------TLEQILTHAASLTPLPPLEG 739
LG +Y+++ RN FKE + +L+ ++ + + TPL L G
Sbjct: 346 LGVMAGDIYSVF--RNAASFKEVSDGLLASSKGQLQKMKESLDDVMDYLVNNTPLNWLVG 403
Query: 740 PMVPQV------------HRPNTWSRPHP 756
P PQV RP WSR HP
Sbjct: 404 PFYPQVTESESAQAPGTTRRPGRWSRKHP 432
>CHSY_PINST (O65872) Chalcone synthase (EC 2.3.1.74)
(Naringenin-chalcone synthase)
Length = 395
Score = 32.3 bits (72), Expect = 5.8
Identities = 14/47 (29%), Positives = 22/47 (46%)
Query: 578 SDGHYSTKSGYQFLRHGAEAVVASSSQVPSLPRADWRKLWKADALLP 624
SD H + G GA A++ + VP + + + LW A +LP
Sbjct: 207 SDTHLDSMVGQALFGDGARALIVGADPVPEVEKPCFEMLWTAQTILP 253
>VE05_VARV (Q01483) Protein E5
Length = 341
Score = 32.0 bits (71), Expect = 7.6
Identities = 19/58 (32%), Positives = 29/58 (49%), Gaps = 1/58 (1%)
Query: 283 INLDKSKLSCSRNVPDDCFHELRQLLTVKAVESYDKYLGLPT-IIGKSKTQIFNFVKE 339
I LDK KL+ ++ + C R + + E +KY P IIGK+ + +FV E
Sbjct: 284 IGLDKLKLNIVHDIVEPCMPVRRPVAKILCKEMVNKYFENPLHIIGKNLQECIDFVSE 341
>RPOB_ASTLO (P27059) DNA-directed RNA polymerase beta chain (EC
2.7.7.6) (PEP) (Plastid-encoded RNA polymerase beta
subunit) (RNA polymerase beta subunit)
Length = 1076
Score = 32.0 bits (71), Expect = 7.6
Identities = 24/118 (20%), Positives = 54/118 (45%), Gaps = 2/118 (1%)
Query: 262 VQEALSIKAILASY--EQASGQVINLDKSKLSCSRNVPDDCFHELRQLLTVKAVESYDKY 319
+ E L + IL S ++ +IN + K ++N+PD ++ L ++ K
Sbjct: 697 ISERLRNEDILTSIHVKKCKSFLINHKEKKEEITKNIPDIKLKNVKNLNNNGIIKIGSKI 756
Query: 320 LGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREVLVKAVAHAIPSYVMSCF 377
G +IG K ++ N+ E +++ L ++ +++ + LV V + + +C+
Sbjct: 757 NGKEILIGIIKKRLINYEVELIYEILSEYERKNISVLSPKNLVGIVTNTKIHKIKNCY 814
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.139 0.444
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 108,592,364
Number of Sequences: 164201
Number of extensions: 4582678
Number of successful extensions: 12025
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 11981
Number of HSP's gapped (non-prelim): 32
length of query: 906
length of database: 59,974,054
effective HSP length: 119
effective length of query: 787
effective length of database: 40,434,135
effective search space: 31821664245
effective search space used: 31821664245
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0154.14


