

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0153b.6
(156 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
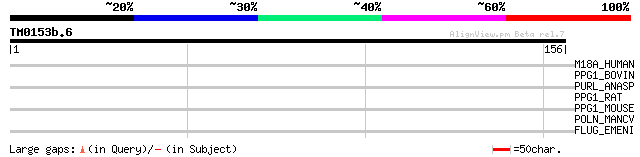
Score E
Sequences producing significant alignments: (bits) Value
M18A_HUMAN (Q92614) Myosin XVIIIA (Myosin 18A) (Myosin containin... 32 0.75
PPG1_BOVIN (Q865B7) Peroxisome proliferator activated receptor g... 30 2.8
PURL_ANASP (Q8YR06) Phosphoribosylformylglycinamidine synthase I... 28 8.3
PPG1_RAT (Q9QYK2) Peroxisome proliferator activated receptor gam... 28 8.3
PPG1_MOUSE (O70343) Peroxisome proliferator activated receptor g... 28 8.3
POLN_MANCV (Q69014) Non-structural polyprotein [Contains: p16; p... 28 8.3
FLUG_EMENI (P38094) Protein fluG 28 8.3
>M18A_HUMAN (Q92614) Myosin XVIIIA (Myosin 18A) (Myosin containing
PDZ domain)
Length = 2054
Score = 31.6 bits (70), Expect = 0.75
Identities = 35/136 (25%), Positives = 54/136 (38%), Gaps = 19/136 (13%)
Query: 15 LTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGL-------LPDTTVLGFAVNQT-SGDF 66
L + Q+P P P+ S EL FPV L LP T+ + + +GDF
Sbjct: 172 LEGQLVQHPGPGIPRPGHRSRAPELVTKKFPVDLRLPPVVPLPPPTLRELELQRRPTGDF 231
Query: 67 SVRL-----------GGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWW 115
L G AC+ + A + G + R+ E+NG V + +
Sbjct: 232 GFSLRRTTMLDRGPEGQACRRVVHFAEPGAGTKDLALGLVPGDRLVEINGHNVESKSRDE 291
Query: 116 SITGIRSSGDNIVFEV 131
+ IR SGD++ +V
Sbjct: 292 IVEMIRQSGDSVRLKV 307
>PPG1_BOVIN (Q865B7) Peroxisome proliferator activated receptor
gamma coactivator 1 alpha (PPAR gamma coactivator-1
alpha) (PPARGC-1 alpha) (PGC-1 alpha)
Length = 796
Score = 29.6 bits (65), Expect = 2.8
Identities = 39/149 (26%), Positives = 58/149 (38%), Gaps = 15/149 (10%)
Query: 11 LFLNLTSKSFQNPNPKP--PKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSV 68
L L LT +S +P P K+ + +EL+ GL P TT A S
Sbjct: 256 LSLPLTPESPNDPKGSPFENKTIERTLSVELSG---TAGLTPPTTPPHKANQDNPFRASP 312
Query: 69 RLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSGDNIV 128
+L +CK +PP + A YS + +G + G + S T + +SG
Sbjct: 313 KLKPSCKTVVPPPSKKARYSES---SCTQGSNSTKKGPEQSELYAQLSKTSVLTSGHE-- 367
Query: 129 FEVGMVTAKYPS-KNFDDSPACEGQRSSS 156
AK PS + F D C+ S +
Sbjct: 368 ----ERKAKRPSLRLFGDHDYCQSINSKT 392
>PURL_ANASP (Q8YR06) Phosphoribosylformylglycinamidine synthase II
(EC 6.3.5.3) (FGAM synthase II)
Length = 782
Score = 28.1 bits (61), Expect = 8.3
Identities = 24/71 (33%), Positives = 31/71 (42%), Gaps = 10/71 (14%)
Query: 24 NPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNY 83
NP+P TP VGL+ D T + Q GD + L GA TL Y
Sbjct: 576 NPQPIYPTPVVGM---------VGLIADLTKICGQGWQGVGDV-IYLLGASITTLGASEY 625
Query: 84 VATYSNTITGK 94
+AT NT+ G+
Sbjct: 626 LATIHNTVAGR 636
>PPG1_RAT (Q9QYK2) Peroxisome proliferator activated receptor gamma
coactivator 1 alpha (PPAR gamma coactivator-1 alpha)
(PPARGC-1 alpha) (PGC-1 alpha)
Length = 796
Score = 28.1 bits (61), Expect = 8.3
Identities = 25/80 (31%), Positives = 34/80 (42%), Gaps = 5/80 (6%)
Query: 11 LFLNLTSKSFQNPNPKP--PKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSV 68
L L LT +S +P P K+ + +EL+ GL P TT A S
Sbjct: 256 LSLPLTPESPNDPKGSPFENKTIERTLSVELSG---TAGLTPPTTPPHKANQDNPFKASP 312
Query: 69 RLGGACKITLPPDNYVATYS 88
+L +CK +PP A YS
Sbjct: 313 KLKPSCKTVVPPPTKRARYS 332
>PPG1_MOUSE (O70343) Peroxisome proliferator activated receptor
gamma coactivator 1 alpha (PPAR gamma coactivator-1
alpha) (PPARGC-1 alpha) (PGC-1 alpha)
Length = 797
Score = 28.1 bits (61), Expect = 8.3
Identities = 25/80 (31%), Positives = 34/80 (42%), Gaps = 5/80 (6%)
Query: 11 LFLNLTSKSFQNPNPKP--PKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSV 68
L L LT +S +P P K+ + +EL+ GL P TT A S
Sbjct: 257 LSLPLTPESPNDPKGSPFENKTIERTLSVELSG---TAGLTPPTTPPHKANQDNPFKASP 313
Query: 69 RLGGACKITLPPDNYVATYS 88
+L +CK +PP A YS
Sbjct: 314 KLKPSCKTVVPPPTKRARYS 333
>POLN_MANCV (Q69014) Non-structural polyprotein [Contains: p16; p23;
Helicase (2C-like protein) (P2C); 3A-like protein; Viral
genome-linked protein (VPg); Thiol protease P3C (EC
3.4.22.-) (3C-like protease) (3C-pro); RNA-directed RNA
polymerase (EC 2.7.7.
Length = 2208
Score = 28.1 bits (61), Expect = 8.3
Identities = 13/45 (28%), Positives = 26/45 (56%)
Query: 7 ILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPD 51
IL++ L T++ FQN P+++ + ++E + F +G+ PD
Sbjct: 2116 ILYSSQLERTAEYFQNDIVNIPENSMAVFNVETNSASFQIGIRPD 2160
>FLUG_EMENI (P38094) Protein fluG
Length = 865
Score = 28.1 bits (61), Expect = 8.3
Identities = 14/40 (35%), Positives = 19/40 (47%)
Query: 72 GACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAF 111
G + LPPDN VA I + V I E +G+R +
Sbjct: 630 GQFEFILPPDNPVAAVDTLIKSRQVIANIVEKHGLRATLY 669
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,355,485
Number of Sequences: 164201
Number of extensions: 776390
Number of successful extensions: 1709
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 1706
Number of HSP's gapped (non-prelim): 7
length of query: 156
length of database: 59,974,054
effective HSP length: 101
effective length of query: 55
effective length of database: 43,389,753
effective search space: 2386436415
effective search space used: 2386436415
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0153b.6


