

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0153b.2
(402 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
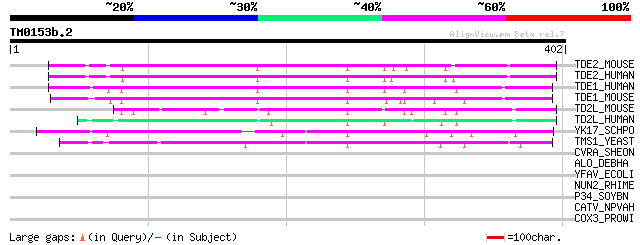
TDE2_MOUSE (Q9QZI8) Tumor differentially expressed protein 2 (Tu... 141 3e-33
TDE2_HUMAN (Q9NRX5) Tumor differentially expressed protein 2 (Tu... 140 8e-33
TDE1_HUMAN (Q13530) Tumor differentially expressed protein 1 (Tr... 132 1e-30
TDE1_MOUSE (Q9QZI9) Tumor differentially expressed protein 1 (Me... 120 6e-27
TD2L_MOUSE (Q8K0E7) Tumor differentially expressed 2-like 120 6e-27
TD2L_HUMAN (Q96SA4) Tumor differentially expressed 2-like (FKSG8... 117 7e-26
YK17_SCHPO (Q9HDY3) Membrane protein PB1A10.07c 113 8e-25
TMS1_YEAST (Q12116) Membrane protein TMS1 95 3e-19
CVRA_SHEON (Q8EAZ0) Cell volume regulation protein A homolog 37 0.090
ALO_DEBHA (Q6BZA0) D-arabinono-1,4-lactone oxidase (EC 1.1.3.37)... 35 0.34
YFAV_ECOLI (P76470) Hypothetical transport protein yfaV 33 1.7
NUN2_RHIME (P56911) NADH-quinone oxidoreductase chain N 2 (EC 1.... 33 1.7
P34_SOYBN (P22895) P34 probable thiol protease precursor (EC 3.4... 32 2.2
CATV_NPVAH (Q80LP4) Viral cathepsin (EC 3.4.22.50) (V-cath) (Cys... 32 3.8
COX3_PROWI (Q37620) Cytochrome c oxidase polypeptide III (EC 1.9... 31 4.9
>TDE2_MOUSE (Q9QZI8) Tumor differentially expressed protein 2 (Tumor
differentially expressed 1 protein like) (Membrane
protein TMS-2) (Axotomy induced glyco/Golgi protein 2)
Length = 453
Score = 141 bits (355), Expect = 3e-33
Identities = 109/427 (25%), Positives = 193/427 (44%), Gaps = 67/427 (15%)
Query: 29 NASNPKMARYVYALIFLVCNLLAWASRDELPG-RSVLTKLKGLKTCKDPKDCL------G 81
+ +N + R +YAL LV +A +PG L K+ G C++ K + G
Sbjct: 31 SGNNSTVTRLIYALFLLVGVCVACVML--IPGMEEQLNKIPGF--CENEKGVVPCNILVG 86
Query: 82 TNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPS- 140
V R+ G +F++++ + ++ R H+G+W K VA+ I F +P
Sbjct: 87 YKAVYRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFATAVAIIIGAFFIPEG 146
Query: 141 ELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFAS--EKYAERCQIHVMLFATA-SY 197
++ V GA F+ IQL+ +I F N+ + E+ RC +L ATA +Y
Sbjct: 147 TFTTVWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNY 206
Query: 198 FICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-----NAGILSPGL 252
+ +V V+L +++Y SC N +FI+ ++L + +S+ PK+ +G+L +
Sbjct: 207 LLSLVAVVLFFVYYTHPASCAENKAFISVNMLLCIGASVMSILPKIQESQPRSGLLQSSV 266
Query: 253 MGLYVVFLCWCAIRSEPE-----------GYECIR--KSDSPNKTDW--QNIISLVVGIL 297
+ +Y ++L W A+ +EPE G+ R D + W Q II LV+ +L
Sbjct: 267 ITVYTMYLTWSAMTNEPETNCNPSLLSIIGFNTTRPIPKDGQSVQWWHPQGIIGLVLFLL 326
Query: 298 ALVIATFSTGIDSKCFQ----------------------------YRKGDKPAEEDDVPY 329
+ ++ T +S+ + +R D E D V Y
Sbjct: 327 CVFYSSIRTSNNSQVNKLTLTSDESTLIEDGNGRSDGSLDDGDGIHRAVDN--ERDGVTY 384
Query: 330 GYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLI 389
Y FFHF+ ++Y M L W + ++ + WT+ WV+I + W+ + +Y+W L+
Sbjct: 385 SYSFFHFMLFLASLYIMMTLTNWYRYEPSRE--MKSQWTAVWVKISSSWIGLVLYVWTLV 442
Query: 390 APIIWKN 396
AP++ N
Sbjct: 443 APLVLTN 449
>TDE2_HUMAN (Q9NRX5) Tumor differentially expressed protein 2 (Tumor
differentially expressed 1 protein like) (UNQ396/PRO732)
Length = 453
Score = 140 bits (352), Expect = 8e-33
Identities = 105/425 (24%), Positives = 191/425 (44%), Gaps = 63/425 (14%)
Query: 29 NASNPKMARYVYALIFLVCNLLAWASRDELPG-RSVLTKLKGLKTCKDPKDCL------G 81
+ +N + R +YAL LV +A +PG L K+ G C++ K + G
Sbjct: 31 SGNNSTVTRLIYALFLLVGVCVACVML--IPGMEEQLNKIPGF--CENEKGVVPCNILVG 86
Query: 82 TNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPS- 140
V R+ G +F++++ + ++ R H+G+W K +A+ I F +P
Sbjct: 87 YKAVYRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFAAAIAIIIGAFFIPEG 146
Query: 141 ELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFAS--EKYAERCQIHVMLFATA-SY 197
++ V GA F+ IQL+ +I F N+ + E+ RC +L ATA +Y
Sbjct: 147 TFTTVWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNY 206
Query: 198 FICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-----NAGILSPGL 252
+ +V ++L +++Y SC N +FI+ ++L + +S+ PK+ +G+L +
Sbjct: 207 LLSLVAIVLFFVYYTHPASCSENKAFISVNMLLCVGASVMSILPKIQESQPRSGLLQSSV 266
Query: 253 MGLYVVFLCWCAIRSEPE-----------GYECI----RKSDSPNKTDWQNIISLVVGIL 297
+ +Y ++L W A+ +EPE GY ++ S Q II L++ +L
Sbjct: 267 ITVYTMYLTWSAMTNEPETNCNPSLLSIIGYNTTSTVPKEGQSVQWWHAQGIIGLILFLL 326
Query: 298 ALVIATFSTGIDSKCFQ---------------------YRKGDK-----PAEEDDVPYGY 331
+ ++ T +S+ + GD E D V Y Y
Sbjct: 327 CVFYSSIRTSNNSQVNKLTLTSDESTLIEDGGARSDGSLEDGDDVHRAVDNERDGVTYSY 386
Query: 332 GFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAP 391
FFHF+ ++Y M L W + ++ + WT+ WV+I + W+ + +Y+W L+AP
Sbjct: 387 SFFHFMLFLASLYIMMTLTNWYRYEPSRE--MKSQWTAVWVKISSSWIGIVLYVWTLVAP 444
Query: 392 IIWKN 396
++ N
Sbjct: 445 LVLTN 449
>TDE1_HUMAN (Q13530) Tumor differentially expressed protein 1
(Transmembrane protein SBBI99)
Length = 473
Score = 132 bits (333), Expect = 1e-30
Identities = 101/439 (23%), Positives = 186/439 (42%), Gaps = 77/439 (17%)
Query: 29 NASNPKMARYVYALIFLVCNLLAWA-SRDELPGRSVLTKLKGL---------KTCKDPKD 78
N+ N + R +YA I L+ ++++ R E+ + L K+ G KD
Sbjct: 31 NSKNSTVTRLIYAFILLLSTVVSYIMQRKEM--ETYLKKIPGFCEGGFKIHEADINADKD 88
Query: 79 C---LGTNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFP 135
C +G V R+S +FF + + + R H+G+W KI + + +
Sbjct: 89 CDVLVGYKAVYRISFAMAIFFFVFSLLMFKVKTSKDLRAAVHNGFWFFKIAALIGIMVGS 148
Query: 136 FLLPSELID-LYGEVAHFGAGVFLFIQLISIISFITWLNDFFAS--EKYAERCQIHVML- 191
F +P ++ V GA +F+ IQL+ ++ F N+ + + E+ R +L
Sbjct: 149 FYIPGGYFSSVWFVVGMIGAALFILIQLVLLVDFAHSWNESWVNRMEEGNPRLWYAALLS 208
Query: 192 FATASYFICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-----NAG 246
F +A Y + ++ V L+Y +Y C N FI+ L+L + + +S+HPK+ +G
Sbjct: 209 FTSAFYILSIICVGLLYTYYTKPDGCTENKFFISINLILCVVASIISIHPKIQEHQPRSG 268
Query: 247 ILSPGLMGLYVVFLCWCAIRSEPE-----------------------GYECIRKSDSPNK 283
+L L+ LY ++L W A+ +EP+ + P+K
Sbjct: 269 LLQSSLITLYTMYLTWSAMSNEPDRSCNPNLMSFITRITAPTLAPGNSTAVVPTPTPPSK 328
Query: 284 T----DWQNIISLVVGILALVIATFSTGIDSKCFQYRKGDKPA----------------- 322
+ D N I L V +L L+ ++ T +S+ + +
Sbjct: 329 SGSLLDSDNFIGLFVFVLCLLYSSIRTSTNSQVDKLTLSGSDSVILGDTTTSGASDEEDG 388
Query: 323 --------EEDDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRI 374
E++ V Y Y FH + ++Y M L W S + K ++ W + WV+I
Sbjct: 389 QPRRAVDNEKEGVQYSYSLFHLMLCLASLYIMMTLTSWYSPDA-KFQSMTSKWPAVWVKI 447
Query: 375 VNEWLAVCVYLWMLIAPII 393
+ W+ + +Y+W L+AP++
Sbjct: 448 SSSWVCLLLYVWTLVAPLV 466
>TDE1_MOUSE (Q9QZI9) Tumor differentially expressed protein 1
(Membrane protein TMS-1) (Axotomy induced glycoprotein
1) (Axotomy induced glyco/Golgi protein 1) (AIGP-1)
Length = 472
Score = 120 bits (301), Expect = 6e-27
Identities = 101/439 (23%), Positives = 176/439 (40%), Gaps = 80/439 (18%)
Query: 30 ASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLK------------TCKDPK 77
+ N + R +YA I + +++ E + T+LK + K K
Sbjct: 32 SKNSTVTRLIYAFILFLGTIVSCIMMTE----GIQTQLKKIPGFCEGGFQIKMVDTKAEK 87
Query: 78 DC---LGTNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIF 134
DC +G V R++ +FF F + + R H+G+W KI + + I
Sbjct: 88 DCDVLVGFKAVYRINFAVAIFFFAFFLLMLKVKTSKDPRAAVHNGFWFFKIAAIIGIMIG 147
Query: 135 PFLLP-SELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFAS--EKYAERCQIHVML 191
F +P +++ GA F+ IQL+ ++ N+ + + E+ R +L
Sbjct: 148 SFYIPGGSFTEVWFVAGMLGASFFIIIQLVLLVDMAHSWNELWVNRMEEGNPRLWYAALL 207
Query: 192 -FATASYFICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-----NA 245
F + Y + +V L+Y++Y C N FI+ L+ ++ VS+ PKV +
Sbjct: 208 SFTSLFYILSIVFAALLYVFYTKPDDCTENKVFISLNLIFCVAVSIVSILPKVQEHQPRS 267
Query: 246 GILSPGLMGLYVVFLCWCAIRSEPEG------YECIRKSDSP-----NKT---------- 284
G+L ++ LY ++L W A+ +EPE I SP N T
Sbjct: 268 GLLQSSIITLYTLYLTWSAMTNEPERSCNPSLMSIITHLTSPTVSPANSTTLAPAYAPPS 327
Query: 285 ------DWQNIISLVVGILALVIATFST------------GIDSKCFQYRKGDKPAEEDD 326
+ +I L++ + L+ ++F T G DS EED
Sbjct: 328 QSGHFMNLDDIWGLIIFVFCLIYSSFRTSSNSQVNKLTLSGSDSVILGDTTNGANDEEDG 387
Query: 327 VP------------YGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRI 374
P Y Y FFH + ++Y M + W S + K + W + W ++
Sbjct: 388 QPRRAVDNEKEGVQYSYSFFHLMLCCASLYIMMTITSWYSPDA-KFQKVSSKWLAVWFKM 446
Query: 375 VNEWLAVCVYLWMLIAPII 393
+ WL + +YLW L+AP++
Sbjct: 447 GSSWLCLLLYLWTLVAPLV 465
>TD2L_MOUSE (Q8K0E7) Tumor differentially expressed 2-like
Length = 450
Score = 120 bits (301), Expect = 6e-27
Identities = 95/372 (25%), Positives = 162/372 (43%), Gaps = 61/372 (16%)
Query: 76 PKDC---LGTNGVLRV--SMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVA 130
P DC LG V R+ + F FF M+ R+S+ + R +G+W K ++ V
Sbjct: 85 PLDCGSLLGFRAVYRMCFATAAFFFFFMLLMICVRSSR--DPRAAIQNGFWFFKFLILVG 142
Query: 131 VTIFPFLLPS----ELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFASEKYAERCQ 186
+T+ F +P ++ +G V F +F+ IQLI + F N + + AE C
Sbjct: 143 ITVGAFYIPDGSFPKIWFYFGVVGSF---LFILIQLILFVDFAHSWNQRWLCK--AEECD 197
Query: 187 -----IHVMLFATASYFICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHP 241
+ F Y + + V LM+++Y +C FI+ L ++ +++ P
Sbjct: 198 SPAWYAGLFFFTFLFYLLSIAAVALMFVYYTESGACHEGKVFISLNLTFCVCVSIIAVLP 257
Query: 242 KV-----NAGILSPGLMGLYVVFLCWCAIRSEPEGYECIRKSDSPNKT------DWQNI- 289
KV N+G+L ++ LY +F+ W A+ + P+ +C + N T D+ +
Sbjct: 258 KVQDAQPNSGLLQASVITLYTMFVTWSALSNVPD-QKCNPHLPTKNGTGQVDLEDYSTVW 316
Query: 290 --ISLVVGILALVIATFSTGIDSKCFQ-----YRKGDKPA------------------EE 324
+VG++ ++ TF + S + + + PA E+
Sbjct: 317 WDAPSIVGLVIFILCTFFISLRSSDHRQVNSLMQTEECPAEMVQQQQVAVSDGRAYDNEQ 376
Query: 325 DDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVY 384
D V Y Y FFHF +++ M L W S +K WTS WV+I W + +Y
Sbjct: 377 DGVTYSYSFFHFCLVLASLHVMMTLTNWYSPGETRKMIST--WTSVWVKICASWAGLFLY 434
Query: 385 LWMLIAPIIWKN 396
LW L+AP++ N
Sbjct: 435 LWTLVAPLLLPN 446
>TD2L_HUMAN (Q96SA4) Tumor differentially expressed 2-like (FKSG84
protein) (UNQ263/PRO300)
Length = 456
Score = 117 bits (292), Expect = 7e-26
Identities = 94/396 (23%), Positives = 158/396 (39%), Gaps = 60/396 (15%)
Query: 50 LAWASRDELPGRSVLTKLKGLKTCKDPKDCLGTNGVLRVSMGCFLFFMMMFWSTARASKL 109
L W + G + T L+G C LG V R+ FF S
Sbjct: 68 LPWVCEE---GAGIPTVLQGHIDCGS---LLGYRAVYRMCFATAAFFFFFTLLMLCVSSS 121
Query: 110 NETRDTWHSGWWSIKIVLWVAVTIFPFLLPS-ELIDLYGEVAHFGAGVFLFIQLISIISF 168
+ R +G+W K ++ V +T+ F +P +++ G+ +F+ IQL+ +I F
Sbjct: 122 RDPRAAIQNGFWFFKFLILVGLTVGAFYIPDGSFTNIWFYFGVVGSFLFILIQLVLLIDF 181
Query: 169 ITWLNDFFASEKYAERCQIH-----VMLFATASYFICMVGVILMYIWYAPQPSCLLNISF 223
N + + AE C + F Y + + V LM+++Y C F
Sbjct: 182 AHSWNQRWLGK--AEECDSRAWYAGLFFFTLLFYLLSIAAVALMFMYYTEPSGCHEGKVF 239
Query: 224 ITFTLVLLQIMTSVSLHPKV-----NAGILSPGLMGLYVVFLCWCAIRSEPE-------- 270
I+ L ++ ++ PKV N+G+L ++ LY +F+ W A+ S PE
Sbjct: 240 ISLNLTFCVCVSIAAVLPKVQDAQPNSGLLQASVITLYTMFVTWSALSSIPEQKCNPHLP 299
Query: 271 ---GYECIRKSDSPNKTDWQNIISLVVGILALVIATFSTGIDSK---------------- 311
G E + +T W + S +VG++ ++ T + S
Sbjct: 300 TQLGNETVVAGPEGYETQWWDAPS-IVGLIIFLLCTLFISLRSSDHRQVNSLMQTEECPP 358
Query: 312 ---CFQYRKGDKPA--------EEDDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKK 360
Q ++ A E+D V Y Y FFHF +++ M L W +K
Sbjct: 359 MLDATQQQQQQVAACEGRAFDNEQDGVTYSYSFFHFCLVLASLHVMMTLTNWYKPGETRK 418
Query: 361 WTIDVGWTSAWVRIVNEWLAVCVYLWMLIAPIIWKN 396
WT+ WV+I W + +YLW L+AP++ +N
Sbjct: 419 MIST--WTAVWVKICASWAGLLLYLWTLVAPLLLRN 452
>YK17_SCHPO (Q9HDY3) Membrane protein PB1A10.07c
Length = 441
Score = 113 bits (283), Expect = 8e-25
Identities = 97/420 (23%), Positives = 179/420 (42%), Gaps = 56/420 (13%)
Query: 20 VSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLTKLKG---LKTCKDP 76
V+S LS + + A YA+++ V +LL+W S L+KL C++
Sbjct: 27 VTSVLSLTNSIQSNVGAVISYAVLYFVNSLLSWCMLSSW-FNSKLSKLSAGYLQFDCQND 85
Query: 77 KDCLGTNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPF 136
C V R+S +F + + + + + + +G W KIVLW + IF F
Sbjct: 86 GKCYSVIAVHRLSFTLVMFHLFLAFILSLCNTRSRVAIKIQNGLWPFKIVLWFVLGIFSF 145
Query: 137 LLPSELIDLYGE-VAHFGAGVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLFATA 195
+P++ + +G ++ G+ +F+ L+ ++ F + +AERC V+ ++
Sbjct: 146 FIPTKFLSFWGNIISVMGSALFIVYGLMLLVDF---------AHTWAERCVDRVLTSDSS 196
Query: 196 S------------YFICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV 243
S Y + +V IL Y+++ SC N + T L+L ++ +S+HP +
Sbjct: 197 SSKFYLIGSTVGMYVVGLVLTILTYVFFCAS-SCSFNQAINTINLLLCIAVSCLSVHPTI 255
Query: 244 -----NAGILSPGLMGLYVVFLCWCAIRSEPEGYECIRKSDSPNKT-DWQNIISLVVGIL 297
+G+ ++ Y +L A+ + P+ +C +S + T ++ +I
Sbjct: 256 QEYNPRSGLAQSSMVMCYTCYLILSALANRPDEGQCNPWGNSASGTREFSKVIGAAFTFF 315
Query: 298 ALV-----IATFSTGIDSKCFQYRKG----------DKPAEEDDVPYGYGF--FHFVFAT 340
++ A+ DS + Y D +EED Y F FH VF
Sbjct: 316 TILYSAVRAASSRESDDSYSYLYADSHDMGVSTPLEDGSSEEDKHQSDYNFIWFHIVFVL 375
Query: 341 GAMYFAMLLVGWNSHHSMKKWTIDV------GWTSAWVRIVNEWLAVCVYLWMLIAPIIW 394
A Y A LL WN+ + DV + + WV+I+ W+ +Y+W +AP+ +
Sbjct: 376 AAFYTASLLTNWNTTSVYENQKNDVFVRIGFSYAAVWVKIITSWVCHGLYVWSCLAPVFF 435
>TMS1_YEAST (Q12116) Membrane protein TMS1
Length = 473
Score = 95.1 bits (235), Expect = 3e-19
Identities = 87/429 (20%), Positives = 177/429 (40%), Gaps = 81/429 (18%)
Query: 37 RYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLKTCKDPKDC-LGTNGVLRVSMGCFLF 95
R +YA+ L+ +L++W S +S+L K TC +C T L ++GC
Sbjct: 43 RLLYAVWLLLNSLISWVSYSA--NKSILWPGK---TCTGTGECGFFTVHRLNFALGCLHL 97
Query: 96 FMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPSELIDLYGEVAHFGAG 155
+ + + +++ N+ R + WWS+K +L++ + + F++P++ + + +G
Sbjct: 98 ILALVLTGVKST--NDVRAALQNSWWSLKFILYLCLIVLSFVIPNDFYIFFSKWVSVPSG 155
Query: 156 -VFLFIQLISIISFI-----TWLNDFFASEKYAERCQIHVMLFATASYFICMVGVILMYI 209
+F+ + LI ++ F T ++ + ++ + Q ++L T+ Y ++ ++MY+
Sbjct: 156 AIFILVGLILLVDFAHEWAETCISHVESEDEDSSFWQRFLVLGTTSMYTASIIMTVVMYV 215
Query: 210 WYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-----NAGILSPGLMGLYVVFLCWCA 264
+ Q C +N + +T L+L I +S++PK+ +G+ ++ +Y +L A
Sbjct: 216 MFCHQ-QCNMNQTAVTVNLILTVITLVLSVNPKIQEANPKSGLAQSSMVSVYCTYLTMSA 274
Query: 265 IRSEPEGYECIRKSDSPNKTDWQNIISLVVGILALVIATFSTGIDS-------------- 310
+ SEP+ C S + I+ + +A+ T +S
Sbjct: 275 MSSEPDDKMCNPLVRSSGTRKFSIILGSLFTFIAIAYTTTRAAANSAFQGTNTNGAIYLG 334
Query: 311 -------------KCFQYRKGDKPAEEDDVP----------------------------- 328
+Y + EE +P
Sbjct: 335 NDIEYEGLGGQTRNQLRYEAIKQAVEEGSLPESALYDTAWLGTSSPTGAMDNQNDDERTG 394
Query: 329 --YGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWT--SAWVRIVNEWLAVCVY 384
Y Y FH +F + A+LL + + + I VG T +WV+IV+ W+ +Y
Sbjct: 395 TKYNYTLFHVIFFLATQWIAILLTINVTQDDVGDF-IPVGRTYFYSWVKIVSAWICYALY 453
Query: 385 LWMLIAPII 393
W ++AP I
Sbjct: 454 GWTVVAPAI 462
>CVRA_SHEON (Q8EAZ0) Cell volume regulation protein A homolog
Length = 574
Score = 37.0 bits (84), Expect = 0.090
Identities = 44/192 (22%), Positives = 77/192 (39%), Gaps = 39/192 (20%)
Query: 11 QQGCGVLLKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLT-KLKG 69
Q G GVLL + ++ + K+A +Y+++ L L+ +A+ ++L G +L+ L G
Sbjct: 195 QFGLGVLLGLGGGWLLWKLVNVSKLAEGLYSILVLSGGLMIYATSNKLGGSGILSIYLVG 254
Query: 70 LKTCKDPKDCLGTNGVLRV--------SMGCFL----------------------FFMMM 99
L P G + +L V +G FL F M++
Sbjct: 255 LFLGNKP--TRGRHAILNVLDGMTWVSQIGMFLVLGLLLTPSDLLDIWLPGLALAFGMIL 312
Query: 100 F------WSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPSELIDLYGEVAHFG 153
F W + K +RD W W ++ + + + +FP + LY +A F
Sbjct: 313 FARPLAVWLSLLPFKSFGSRDRWFISWVGLRGAVPIILAVFPMMAGLPGAQLYFNLAFFV 372
Query: 154 AGVFLFIQLISI 165
V L +Q S+
Sbjct: 373 VIVSLLVQGASL 384
>ALO_DEBHA (Q6BZA0) D-arabinono-1,4-lactone oxidase (EC 1.1.3.37)
(ALO) (L-galactono-gamma-lactone oxidase)
Length = 557
Score = 35.0 bits (79), Expect = 0.34
Identities = 22/65 (33%), Positives = 36/65 (54%), Gaps = 12/65 (18%)
Query: 101 WSTARASK-LNETRDTWHSGWWSIKI---VLWVAVTIFPFLLPSELID------LYGEVA 150
W +++S+ L++ RD+W+ W+ K +LW++V I P L P LI+ YG+V
Sbjct: 252 WRASKSSEPLSKPRDSWYGTWFGRKFYESLLWISVHICPHLTP--LIEKFVFKNQYGDVE 309
Query: 151 HFGAG 155
G G
Sbjct: 310 TLGHG 314
>YFAV_ECOLI (P76470) Hypothetical transport protein yfaV
Length = 429
Score = 32.7 bits (73), Expect = 1.7
Identities = 37/137 (27%), Positives = 58/137 (42%), Gaps = 7/137 (5%)
Query: 121 WSIKIV---LWVAVTIFPFLLPSELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFA 177
W + I+ + VAV F LP+++ L G F A V I ++ + F TWL +
Sbjct: 241 WQLAIIYLTIQVAVYGLIFFLPTQVAALLGTKVGFTASVVTAIPWVAAL-FGTWLIPRY- 298
Query: 178 SEKYAERCQIHVMLFATASYFICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSV 237
S+K ER + + A I + G++ + A C+ I FI V + T +
Sbjct: 299 SDKTGERRNVAALTLLAAGIGIGLSGLLSPVM--AIVALCVAAIGFIAVQPVFWTMPTQL 356
Query: 238 SLHPKVNAGILSPGLMG 254
+ AGI L G
Sbjct: 357 LSGTALAAGIGFVNLFG 373
>NUN2_RHIME (P56911) NADH-quinone oxidoreductase chain N 2 (EC
1.6.99.5) (NADH dehydrogenase I, chain N 2) (NDH-1,
chain N 2)
Length = 479
Score = 32.7 bits (73), Expect = 1.7
Identities = 27/95 (28%), Positives = 46/95 (48%), Gaps = 10/95 (10%)
Query: 125 IVLWVAVTIFPFLLPSELIDLYGEVAHFGAGVFL---FIQLISIISFITWLNDFFASEKY 181
++LW +V I + L+ L GE+ AG+F+ F ++ ++ + FF S KY
Sbjct: 40 LLLWASVAIVLLAAVATLM-LAGEMRPAYAGMFISDRFAVFFKLVFYLATILTFFLSRKY 98
Query: 182 AE-----RCQIHV-MLFATASYFICMVGVILMYIW 210
AE R + +V +LFA I + L+ I+
Sbjct: 99 AEIEGIGRSEYYVLLLFALVGMMIMASAIDLLSIY 133
>P34_SOYBN (P22895) P34 probable thiol protease precursor (EC
3.4.22.-)
Length = 379
Score = 32.3 bits (72), Expect = 2.2
Identities = 28/103 (27%), Positives = 40/103 (38%), Gaps = 18/103 (17%)
Query: 270 EGYECIRKSDSPNKTDWQNIISLVVGILALVIATFSTGIDSKCFQ-YRKGDKPAEEDDVP 328
+GYE + SD +++ + + A++ S ID+K F Y G E P
Sbjct: 242 DGYETLIMSDESTESETEQAF-----LSAILEQPISVSIDAKDFHLYTGGIYDGENCTSP 296
Query: 329 YGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAW 371
YG HFV LLVG+ S + W W W
Sbjct: 297 YGIN--HFV----------LLVGYGSADGVDYWIAKNSWGFDW 327
>CATV_NPVAH (Q80LP4) Viral cathepsin (EC 3.4.22.50) (V-cath)
(Cysteine proteinase) (CP)
Length = 337
Score = 31.6 bits (70), Expect = 3.8
Identities = 15/43 (34%), Positives = 21/43 (47%), Gaps = 1/43 (2%)
Query: 329 YGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAW 371
Y G HF G + A+LLVG+ + + WT+ W S W
Sbjct: 268 YSKGIIHFCENLGLNH-AVLLVGYGTEGGVSYWTLKNSWGSDW 309
>COX3_PROWI (Q37620) Cytochrome c oxidase polypeptide III (EC
1.9.3.1)
Length = 262
Score = 31.2 bits (69), Expect = 4.9
Identities = 30/109 (27%), Positives = 42/109 (38%), Gaps = 16/109 (14%)
Query: 287 QNIISLVVGILALVIATFSTGIDSKCFQYRKGDKPAEEDDVPYGY------GFFHFVFAT 340
Q IISL + +L VI T ++ + P D YG GF F
Sbjct: 158 QGIISLAITVLLAVIFTGFQALEYL-------EAPFTISDGIYGSTFYLATGFHGFHVII 210
Query: 341 GAMYFAMLLVGWNSHHSMKKWTID---VGWTSAWVRIVNEWLAVCVYLW 386
G ++ A+ L HH K + W V +V +L VC+Y W
Sbjct: 211 GTIFIAVCLFRLVLHHFSKSHHLGFEAAAWYWHMVDVVWLFLFVCIYYW 259
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.328 0.140 0.466
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,840,153
Number of Sequences: 164201
Number of extensions: 2002161
Number of successful extensions: 5464
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 5420
Number of HSP's gapped (non-prelim): 24
length of query: 402
length of database: 59,974,054
effective HSP length: 112
effective length of query: 290
effective length of database: 41,583,542
effective search space: 12059227180
effective search space used: 12059227180
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0153b.2


