

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0148.13
(480 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
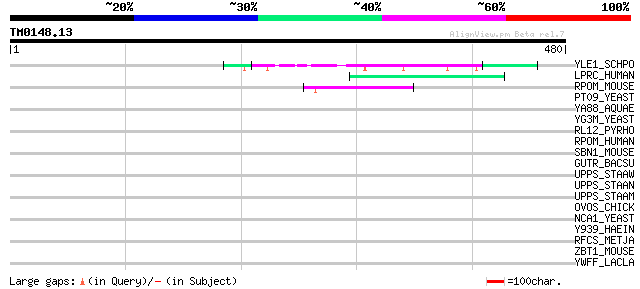
Score E
Sequences producing significant alignments: (bits) Value
YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I 57 1e-07
LPRC_HUMAN (P42704) 130 kDa leucine-rich protein (LRP 130) (GP13... 47 1e-04
RPOM_MOUSE (Q8BKF1) DNA-directed RNA polymerase, mitochondrial p... 44 7e-04
PT09_YEAST (P32522) PET309 protein, mitochondrial precursor 39 0.023
YA88_AQUAE (O67178) Hypothetical protein AQ_1088 38 0.051
YG3M_YEAST (P48237) Hypothetical 101.4 kDa protein in RPL24B-RSR... 37 0.086
RL12_PYRHO (O57705) 50S ribosomal protein L12P 35 0.56
RPOM_HUMAN (O00411) DNA-directed RNA polymerase, mitochondrial p... 34 0.95
SBN1_MOUSE (Q8R4Y4) Stabilin 1 precursor (FEEL-1 protein) 33 1.2
GUTR_BACSU (P39143) Transcription activator gutR 33 1.6
UPPS_STAAW (Q8NWZ5) Undecaprenyl pyrophosphate synthetase (EC 2.... 33 2.1
UPPS_STAAN (P60477) Undecaprenyl pyrophosphate synthetase (EC 2.... 33 2.1
UPPS_STAAM (P60476) Undecaprenyl pyrophosphate synthetase (EC 2.... 33 2.1
OVOS_CHICK (P20740) Ovostatin precursor (Ovomacroglobulin) 33 2.1
NCA1_YEAST (P32493) Nuclear control of ATPase messenger RNA expr... 33 2.1
Y939_HAEIN (P44080) Hypothetical protein HI0939 31 6.2
RFCS_METJA (Q58817) Replication factor C small subunit (RFC smal... 31 6.2
ZBT1_MOUSE (Q91VL9) Zinc finger and BTB domain containing protein 1 31 8.1
YWFF_LACLA (Q9CDP0) Hypothetical RNA methyltransferase ywfF (EC ... 31 8.1
>YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I
Length = 1261
Score = 57.0 bits (136), Expect = 1e-07
Identities = 51/206 (24%), Positives = 92/206 (43%), Gaps = 19/206 (9%)
Query: 210 LVETFIKSKTFN--YRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIV 267
L+ K+K F+ R F+HS + L+ + + W + ++LD IL+ +
Sbjct: 813 LISVLSKNKRFDAVQRVFEHSKH--LYRKISTKSLEKANW-FMALILDAM---ILSSSFA 866
Query: 268 MYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFL---GKEDKPLAAVNLLKQMRE 324
K D ++L G+ P T+ L++ G D A+N+ ++ +
Sbjct: 867 RQFKSSNLFCDNMKML-------GYIPRASTFAHLINNSTRRGDTDDATTALNIFEETKR 919
Query: 325 TGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEK 384
++P+V Y ++ L RA C F EM ++G +P V Y +I G+
Sbjct: 920 HNVKPSVFLYNAVLSKLGRARRTTECWKLFQEMKESGLLPTSVTYGTVINAACRIGDESL 979
Query: 385 AQEMFHEMISKEQV-PNVYTYNSMIR 409
A+++F EM ++ P V YN+MI+
Sbjct: 980 AEKLFAEMENQPNYQPRVAPYNTMIQ 1005
Score = 56.6 bits (135), Expect = 1e-07
Identities = 65/290 (22%), Positives = 117/290 (39%), Gaps = 26/290 (8%)
Query: 186 GFPATARTFNILIRTC---GEAGLAKDLVETFIKSKTFNYRPFKHSYNAILHSLLVLNQY 242
G+ A TF LI G+ A + F ++K N +P YNA+L L +
Sbjct: 883 GYIPRASTFAHLINNSTRRGDTDDATTALNIFEETKRHNVKPSVFLYNAVLSKLGRARRT 942
Query: 243 KLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEM-GRNGFSPDFHTYNL 301
++Q+M G +TY V+ A R+G + LF EM + + P YN
Sbjct: 943 TECWKLFQEMKESGLLPTSVTYGTVINAACRIGDESLAEKLFAEMENQPNYQPRVAPYNT 1002
Query: 302 LLHF----LGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLID--GLSRAGNLHACKHFFD 355
++ F + +K L N ++ T IEP+ Y L+D G + N+ + K +
Sbjct: 1003 MIQFEVQTMFNREKALFYYN---RLCATDIEPSSHTYKLLMDAYGTLKPVNVGSVKAVLE 1059
Query: 356 EMIKNGCMPDVVAYTVMITGYV-----VAGELEKAQEMFHEMISKEQVPNVY----TYNS 406
M + DV ++ Y+ V +++ A + ++K + + S
Sbjct: 1060 LMERT----DVPILSMHYAAYIHILGNVVSDVQAATSCYMNALAKHDAGEIQLDANLFQS 1115
Query: 407 MIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAR 456
I L + E ++ +M+ S N+ + +L+ G ++ AR
Sbjct: 1116 QIESLIANDRIVEGIQIVSDMKRYNVSLNAYIVNALIKGFTKVGMISKAR 1165
Score = 42.7 bits (99), Expect = 0.002
Identities = 28/119 (23%), Positives = 52/119 (43%), Gaps = 3/119 (2%)
Query: 353 FFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQ---EMFHEMISKEQVPNVYTYNSMIR 409
F D M G +P + +I G+ + A +F E P+V+ YN+++
Sbjct: 875 FCDNMKMLGYIPRASTFAHLINNSTRRGDTDDATTALNIFEETKRHNVKPSVFLYNAVLS 934
Query: 410 GLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKGKY 468
L A + E + +EM+ G P SV Y ++++ G + A ++ +M + Y
Sbjct: 935 KLGRARRTTECWKLFQEMKESGLLPTSVTYGTVINAACRIGDESLAEKLFAEMENQPNY 993
Score = 38.5 bits (88), Expect = 0.039
Identities = 26/112 (23%), Positives = 46/112 (40%), Gaps = 3/112 (2%)
Query: 322 MRETGIEPTVLHYTTLIDGLSRAGNLH---ACKHFFDEMIKNGCMPDVVAYTVMITGYVV 378
M+ G P + LI+ +R G+ + F+E ++ P V Y +++
Sbjct: 879 MKMLGYIPRASTFAHLINNSTRRGDTDDATTALNIFEETKRHNVKPSVFLYNAVLSKLGR 938
Query: 379 AGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETK 430
A + ++F EM +P TY ++I C G A + EME +
Sbjct: 939 ARRTTECWKLFQEMKESGLLPTSVTYGTVINAACRIGDESLAEKLFAEMENQ 990
Score = 36.2 bits (82), Expect = 0.19
Identities = 58/319 (18%), Positives = 108/319 (33%), Gaps = 49/319 (15%)
Query: 159 YHLIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNILIRTC---GEAGLAKDLVETFI 215
Y+ +++ W+L EM E G T+ T+ +I G+ LA+ L
Sbjct: 929 YNAVLSKLGRARRTTECWKLFQEMKESGLLPTSVTYGTVINAACRIGDESLAEKLFAEME 988
Query: 216 KSKTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLG 275
NY+P YN +I++ Q M
Sbjct: 989 NQP--NYQPRVAPYNT------------MIQFEVQTMF---------------------- 1012
Query: 276 KLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPL---AAVNLLKQMRETGIEPTVL 332
++ + + P HTY LL+ G KP+ + +L+ M T + +
Sbjct: 1013 NREKALFYYNRLCATDIEPSSHTYKLLMDAYGTL-KPVNVGSVKAVLELMERTDVPILSM 1071
Query: 333 HYTTLIDGLSRAGN--LHACKHFFDEMIKNGC---MPDVVAYTVMITGYVVAGELEKAQE 387
HY I L + A + + + K+ D + I + + + +
Sbjct: 1072 HYAAYIHILGNVVSDVQAATSCYMNALAKHDAGEIQLDANLFQSQIESLIANDRIVEGIQ 1131
Query: 388 MFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCS-PNSVVYVSLLSCL 446
+ +M N Y N++I+G G +A +E +G S Y +++
Sbjct: 1132 IVSDMKRYNVSLNAYIVNALIKGFTKVGMISKARYYFDLLECEGMSGKEPSTYENMVRAY 1191
Query: 447 QNAGKVANAREVMRQMMEK 465
+ A E++ Q+ K
Sbjct: 1192 LSVNDGRKAMEIVEQLKRK 1210
>LPRC_HUMAN (P42704) 130 kDa leucine-rich protein (LRP 130) (GP130)
(Leucine-rich PPR-motif containing protein)
Length = 1273
Score = 47.0 bits (110), Expect = 1e-04
Identities = 29/134 (21%), Positives = 53/134 (38%)
Query: 295 DFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFF 354
D YN LL + + + + L +M E I+P + Y LI G++
Sbjct: 40 DVSHYNALLKVYLQNEYKFSPTDFLAKMEEANIQPNRVTYQRLIASYCNVGDIEGASKIL 99
Query: 355 DEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMA 414
M ++ ++TG+ AG++E A+ + M P TY +++
Sbjct: 100 GFMKTKDLPVTEAVFSALVTGHARAGDMENAENILTVMRDAGIEPGPDTYLALLNAYAEK 159
Query: 415 GKFDEACSMLKEME 428
G D L+++E
Sbjct: 160 GDIDHVKQTLEKVE 173
Score = 43.5 bits (101), Expect = 0.001
Identities = 28/113 (24%), Positives = 48/113 (41%)
Query: 354 FDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCM 413
+D + K G + DV Y ++ Y+ + +M PN TY +I C
Sbjct: 29 WDTLQKLGAVYDVSHYNALLKVYLQNEYKFSPTDFLAKMEEANIQPNRVTYQRLIASYCN 88
Query: 414 AGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
G + A +L M+TK V+ +L++ AG + NA ++ M + G
Sbjct: 89 VGDIEGASKILGFMKTKDLPVTEAVFSALVTGHARAGDMENAENILTVMRDAG 141
Score = 35.4 bits (80), Expect = 0.33
Identities = 35/150 (23%), Positives = 67/150 (44%), Gaps = 8/150 (5%)
Query: 257 FSSDILT--YNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLL--LHFLGKEDKP 312
+ SD++T Y ++ R K++ L +E R S T N L + L K K
Sbjct: 584 YESDMVTGGYAALINLCCRHDKVEDALNLKEEFDRLDSSAVLDTGNYLGLVRVLAKHGKL 643
Query: 313 LAAVNLLKQMRETGI---EPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNG-CMPDVVA 368
A+ +LK+M+E + + T L + +++G + G + K + ++ G P
Sbjct: 644 QDAIKILKEMKEKDVLIKDTTALSFFHMLNGAALRGEIETVKQLHEAIVTLGLAEPSTNI 703
Query: 369 YTVMITGYVVAGELEKAQEMFHEMISKEQV 398
++T ++ G+L A E+ + K +V
Sbjct: 704 SFPLVTVHLEKGDLSTALEVAIDCYEKYKV 733
>RPOM_MOUSE (Q8BKF1) DNA-directed RNA polymerase, mitochondrial
precursor (EC 2.7.7.6) (MtRPOL)
Length = 1207
Score = 44.3 bits (103), Expect = 7e-04
Identities = 30/99 (30%), Positives = 46/99 (46%), Gaps = 4/99 (4%)
Query: 255 DGFSSDILT---YNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDK 311
+G +LT YN VM R G + +F + G SPD +Y L +G+ D+
Sbjct: 224 NGDRQQVLTLHMYNTVMLGWARKGSFRELVYVFLMLKDAGLSPDLCSYAAALQCMGRRDQ 283
Query: 312 PLAAV-NLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHA 349
+ + LKQM E G +P +L +++ RA L A
Sbjct: 284 DVRTIQRCLKQMMEEGFQPQLLFTDLVLEEEDRAALLRA 322
Score = 40.8 bits (94), Expect = 0.008
Identities = 23/84 (27%), Positives = 41/84 (48%), Gaps = 1/84 (1%)
Query: 390 HEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNA 449
H ++QV ++ YN+++ G G F E + ++ G SP+ Y + L C+
Sbjct: 222 HNNGDRQQVLTLHMYNTVMLGWARKGSFRELVYVFLMLKDAGLSPDLCSYAAALQCMGRR 281
Query: 450 GK-VANAREVMRQMMEKGKYAHLL 472
+ V + ++QMME+G LL
Sbjct: 282 DQDVRTIQRCLKQMMEEGFQPQLL 305
>PT09_YEAST (P32522) PET309 protein, mitochondrial precursor
Length = 965
Score = 39.3 bits (90), Expect = 0.023
Identities = 29/149 (19%), Positives = 60/149 (39%)
Query: 264 YNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMR 323
++ ++ A RL D Q F + G P Y +L++ + + + + L Q+
Sbjct: 314 FDYLLVAHSRLHNWDALQQQFNALFGIGKLPSIQHYGILMYTMARIGELDSVNKLYTQLL 373
Query: 324 ETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELE 383
G+ PT +L+ + G+ AC F+ K P +T+M+ Y +L+
Sbjct: 374 RRGMIPTYAVLQSLLYAHYKVGDFAACFSHFELFKKYDITPSTATHTIMLKVYRGLNDLD 433
Query: 384 KAQEMFHEMISKEQVPNVYTYNSMIRGLC 412
A + + V + +++ +C
Sbjct: 434 GAFRILKRLSEDPSVEITEGHFALLIQMC 462
Score = 38.1 bits (87), Expect = 0.051
Identities = 20/123 (16%), Positives = 53/123 (42%)
Query: 342 SRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNV 401
SR N A + F+ + G +P + Y +++ GEL+ +++ +++ + +P
Sbjct: 322 SRLHNWDALQQQFNALFGIGKLPSIQHYGILMYTMARIGELDSVNKLYTQLLRRGMIPTY 381
Query: 402 YTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQ 461
S++ G F S + + +P++ + +L + + A ++++
Sbjct: 382 AVLQSLLYAHYKVGDFAACFSHFELFKKYDITPSTATHTIMLKVYRGLNDLDGAFRILKR 441
Query: 462 MME 464
+ E
Sbjct: 442 LSE 444
Score = 37.4 bits (85), Expect = 0.086
Identities = 27/127 (21%), Positives = 54/127 (42%), Gaps = 3/127 (2%)
Query: 215 IKSKTFNYRPFKHSYNAILHSLLV---LNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAK 271
+ S + Y +H N + L+ L+ + ++ + + G I Y I+MY
Sbjct: 297 VSSTYWKYPDAQHDQNQFDYLLVAHSRLHNWDALQQQFNALFGIGKLPSIQHYGILMYTM 356
Query: 272 YRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTV 331
R+G+LD L+ ++ R G P + LL+ K A + + ++ I P+
Sbjct: 357 ARIGELDSVNKLYTQLLRRGMIPTYAVLQSLLYAHYKVGDFAACFSHFELFKKYDITPST 416
Query: 332 LHYTTLI 338
+T ++
Sbjct: 417 ATHTIML 423
Score = 34.7 bits (78), Expect = 0.56
Identities = 28/145 (19%), Positives = 57/145 (39%), Gaps = 5/145 (3%)
Query: 329 PTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEM 388
P++ HY L+ ++R G L + + ++++ G +P ++ + G+
Sbjct: 344 PSIQHYGILMYTMARIGELDSVNKLYTQLLRRGMIPTYAVLQSLLYAHYKVGDFAACFSH 403
Query: 389 FHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQN 448
F + P+ T+ M++ D A +LK + + + +LL +Q
Sbjct: 404 FELFKKYDITPSTATHTIMLKVYRGLNDLDGAFRILKRLSEDPSVEITEGHFALL--IQM 461
Query: 449 AGKVAN---AREVMRQMMEKGKYAH 470
K N A+E+ M E H
Sbjct: 462 CCKTTNHLIAQELFNLMTEHYNIQH 486
>YA88_AQUAE (O67178) Hypothetical protein AQ_1088
Length = 761
Score = 38.1 bits (87), Expect = 0.051
Identities = 42/167 (25%), Positives = 75/167 (44%), Gaps = 12/167 (7%)
Query: 301 LLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKN 360
LL FLG+ + A L K ++ + ++ + Y L G L +H+++ +
Sbjct: 77 LLYFFLGRVED--AERVLKKALKFSDVDDAL--YARLGALYYSQGKLEEAQHYWERALSL 132
Query: 361 GCMPDVVAYTVMITGYVVAGELEKAQEMFHEMIS-KEQVPNVYTYNSMIRGLCMAGKFDE 419
+ Y + + ++ GELEKA ++F + K ++I L + DE
Sbjct: 133 NPNKVEILYNLGVL-HLNKGELEKALDLFERALRLKPDFREAEEKKTLI--LLSLNRIDE 189
Query: 420 ACS-MLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEK 465
+E+E +PN VY+ L + L AG++A AR V ++ EK
Sbjct: 190 LVEEYYRELEK---NPNEEVYIKLGNTLYTAGRLAEARAVFQEGAEK 233
>YG3M_YEAST (P48237) Hypothetical 101.4 kDa protein in RPL24B-RSR1
intergenic region
Length = 864
Score = 37.4 bits (85), Expect = 0.086
Identities = 54/253 (21%), Positives = 96/253 (37%), Gaps = 44/253 (17%)
Query: 219 TFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLD 278
T Y PF HS+L L++ MLL+ + D++ +N MY
Sbjct: 191 TVKYLPF--------HSILHLDE----------MLLEQINGDVVKFNTHMY--------- 223
Query: 279 QFQILFKEMGR---NGFSPDFHTYNLLL---HFLGKEDKPLAAVNLLKQMRETGIEPTVL 332
+ +F +G F+ D ++L L + DK L +E G V
Sbjct: 224 --ECIFNNLGNLKPTNFNQDGTNDKVILKMKELLERYDKALKITEERINKKE-GFPSKVP 280
Query: 333 HYTTLIDG-----LSRAGNLHACKHFFDEMIKN-GCMPDVVAYTVMITGYVVAGELEKAQ 386
T I ++ + H +F + + G P+ T +I Y ++A
Sbjct: 281 KMTQAILNNCLKYSTKCSSFHDMDYFITKFRDDYGITPNKQNLTTVIQFYSRKEMTKQAW 340
Query: 387 EMFHEM--ISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLS 444
F M +S + P++ TYN+M+R F +A + +E++ P + Y+ +
Sbjct: 341 NTFDTMKFLSTKHFPDICTYNTMLRICEKERNFPKALDLFQEIQDHNIKPTTNTYIMMAR 400
Query: 445 CLQNAGKVANARE 457
L ++ A E
Sbjct: 401 VLASSSSNAVVSE 413
>RL12_PYRHO (O57705) 50S ribosomal protein L12P
Length = 108
Score = 34.7 bits (78), Expect = 0.56
Identities = 28/98 (28%), Positives = 40/98 (40%), Gaps = 11/98 (11%)
Query: 301 LLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKN 360
LLLH +GKE + NL ++ G+EP L+ L DE+I+
Sbjct: 8 LLLHSVGKE---INEENLKAVLQAAGVEPEEARIKALVAALEGVN--------IDEVIEK 56
Query: 361 GCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQV 398
MP VA GE +K +E E +E+V
Sbjct: 57 AAMPVAVAAAPAAAPAEAGGEEKKEEEKKEEEEKEEEV 94
>RPOM_HUMAN (O00411) DNA-directed RNA polymerase, mitochondrial
precursor (EC 2.7.7.6) (MtRPOL)
Length = 1230
Score = 33.9 bits (76), Expect = 0.95
Identities = 20/76 (26%), Positives = 36/76 (47%), Gaps = 4/76 (5%)
Query: 390 HEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCL--- 446
H K ++ + YN+++ G G F E +L ++ G +P+ + Y + L C+
Sbjct: 250 HGQRQKRKLLTLDMYNAVMLGWARQGAFKELVYVLFMVKDAGLTPDLLSYAAALQCMGRQ 309
Query: 447 -QNAGKVANAREVMRQ 461
Q+AG + E M Q
Sbjct: 310 DQDAGTIERCLEQMSQ 325
Score = 33.5 bits (75), Expect = 1.2
Identities = 23/87 (26%), Positives = 38/87 (43%), Gaps = 1/87 (1%)
Query: 264 YNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAV-NLLKQM 322
YN VM R G + + + G +PD +Y L +G++D+ + L+QM
Sbjct: 264 YNAVMLGWARQGAFKELVYVLFMVKDAGLTPDLLSYAAALQCMGRQDQDAGTIERCLEQM 323
Query: 323 RETGIEPTVLHYTTLIDGLSRAGNLHA 349
+ G++ L L+ RA L A
Sbjct: 324 SQEGLKLQALFTAVLLSEEDRATVLKA 350
>SBN1_MOUSE (Q8R4Y4) Stabilin 1 precursor (FEEL-1 protein)
Length = 2571
Score = 33.5 bits (75), Expect = 1.2
Identities = 19/58 (32%), Positives = 34/58 (57%), Gaps = 5/58 (8%)
Query: 6 RLGPKMVHRLSHCLIIS--RRLCDRP--FGGDGFECVEEPLKKLGSD-YDSSVDEWFQ 58
R+G H L+ C + +R+C P FGGDGF C + +++L ++ + S+ +WF+
Sbjct: 953 RVGNGGCHGLATCKAVGGGQRVCTCPPHFGGDGFSCYGDIIQELEANAHFSAFSQWFK 1010
>GUTR_BACSU (P39143) Transcription activator gutR
Length = 829
Score = 33.1 bits (74), Expect = 1.6
Identities = 32/123 (26%), Positives = 51/123 (41%), Gaps = 10/123 (8%)
Query: 308 KEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDG-LSRAGNLHACKHFFDEMIKN-----G 361
K+ + A +L ++ I PT+ H L+ G LS A H E G
Sbjct: 672 KDGQQEKAETILNEIENQPISPTIQHRVLLVRGDLSFARGYHVEAIQLYEAANEISSTYG 731
Query: 362 CMPDVVAYTVMITGYVVAGELEKAQEMFHEMI----SKEQVPNVYTYNSMIRGLCMAGKF 417
+ AY + YV + EKA+E F +M+ + QV +Y + M + L G+
Sbjct: 732 GEKTIEAYFNLGVAYVKCDQFEKAEEAFEQMLYDKHNANQVELIYYHYGMAQLLYRKGEK 791
Query: 418 DEA 420
+A
Sbjct: 792 TKA 794
>UPPS_STAAW (Q8NWZ5) Undecaprenyl pyrophosphate synthetase (EC
2.5.1.31) (UPP synthetase)
(Di-trans-poly-cis-decaprenylcistransferase)
(Undecaprenyl diphosphate synthase) (UDS)
Length = 256
Score = 32.7 bits (73), Expect = 2.1
Identities = 21/80 (26%), Positives = 37/80 (46%), Gaps = 1/80 (1%)
Query: 292 FSPDFHTYNLLLHFLGKEDK-PLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHAC 350
F P+ N+ + +G DK P + + + +E T L I+ RA +H+
Sbjct: 103 FLPELIEKNVKVETIGFTDKLPKSTIEAINNAKEKTANNTGLKLIFAINYGGRAELVHSI 162
Query: 351 KHFFDEMIKNGCMPDVVAYT 370
K+ FDE+ + G D++ T
Sbjct: 163 KNMFDELHQQGLNSDIIDET 182
>UPPS_STAAN (P60477) Undecaprenyl pyrophosphate synthetase (EC
2.5.1.31) (UPP synthetase)
(Di-trans-poly-cis-decaprenylcistransferase)
(Undecaprenyl diphosphate synthase) (UDS)
Length = 256
Score = 32.7 bits (73), Expect = 2.1
Identities = 21/80 (26%), Positives = 37/80 (46%), Gaps = 1/80 (1%)
Query: 292 FSPDFHTYNLLLHFLGKEDK-PLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHAC 350
F P+ N+ + +G DK P + + + +E T L I+ RA +H+
Sbjct: 103 FLPELIEKNVKVETIGFTDKLPKSTIEAINNAKEKTANNTGLKLIFAINYGGRAELVHSI 162
Query: 351 KHFFDEMIKNGCMPDVVAYT 370
K+ FDE+ + G D++ T
Sbjct: 163 KNMFDELHQQGLNSDIIDET 182
>UPPS_STAAM (P60476) Undecaprenyl pyrophosphate synthetase (EC
2.5.1.31) (UPP synthetase)
(Di-trans-poly-cis-decaprenylcistransferase)
(Undecaprenyl diphosphate synthase) (UDS)
Length = 256
Score = 32.7 bits (73), Expect = 2.1
Identities = 21/80 (26%), Positives = 37/80 (46%), Gaps = 1/80 (1%)
Query: 292 FSPDFHTYNLLLHFLGKEDK-PLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHAC 350
F P+ N+ + +G DK P + + + +E T L I+ RA +H+
Sbjct: 103 FLPELIEKNVKVETIGFTDKLPKSTIEAINNAKEKTANNTGLKLIFAINYGGRAELVHSI 162
Query: 351 KHFFDEMIKNGCMPDVVAYT 370
K+ FDE+ + G D++ T
Sbjct: 163 KNMFDELHQQGLNSDIIDET 182
>OVOS_CHICK (P20740) Ovostatin precursor (Ovomacroglobulin)
Length = 1473
Score = 32.7 bits (73), Expect = 2.1
Identities = 19/82 (23%), Positives = 37/82 (44%), Gaps = 10/82 (12%)
Query: 357 MIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMI----------SKEQVPNVYTYNS 406
++ N M V + ++ Y+ LE M H +I S++ + ++YT
Sbjct: 1091 ILVNNAMKGGVENELSLSAYITIALLEAGHSMSHTVIRNAFYCLETASEKNITDIYTQAL 1150
Query: 407 MIRGLCMAGKFDEACSMLKEME 428
+ C+AGK + S L+E++
Sbjct: 1151 VAYAFCLAGKAEICESFLRELQ 1172
>NCA1_YEAST (P32493) Nuclear control of ATPase messenger RNA
expression protein, mitochondrial precursor
Length = 518
Score = 32.7 bits (73), Expect = 2.1
Identities = 39/150 (26%), Positives = 67/150 (44%), Gaps = 31/150 (20%)
Query: 299 YNLLLHFLGKEDKP------------LAAVN--LLKQMRETGIEPTVLHYTTLIDGLSRA 344
YN+L E+ P LAA+N E + P V+ L+ +
Sbjct: 239 YNILCKVEANENPPYSPTLVTALENLLAAINNRFFPGRCENSLHPIVIEQ--LLSYFIKT 296
Query: 345 GNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGEL--EKAQEMFHEMISK-----EQ 397
GNL+ K+F +IK G +P+ +I Y+ A ++ +K+ ++F ++ SK +
Sbjct: 297 GNLNESKNFLGHLIKKGILPE----ATIINRYLEAIDVHFDKSTKIF-DIRSKFAFIADL 351
Query: 398 VPNVYTYNS--MIRGLC-MAGKFDEACSML 424
P + Y + + + L M FDE CS+L
Sbjct: 352 APIIENYGTIDLFKFLIPMCRHFDELCSLL 381
>Y939_HAEIN (P44080) Hypothetical protein HI0939
Length = 238
Score = 31.2 bits (69), Expect = 6.2
Identities = 28/114 (24%), Positives = 49/114 (42%), Gaps = 4/114 (3%)
Query: 303 LHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNL--HACKHFFDEMIKN 360
L +GK+ + L L ++ E+ + L +S+ N ++C FF ++ KN
Sbjct: 56 LQLIGKDLRRLGFRALNAKLTESNLSLFELDEQGTAIFISQEDNAPPNSCVLFFYDLNKN 115
Query: 361 GCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMA 414
GC+ T M G + +E+F +S + + TY S+I C A
Sbjct: 116 GCIGKGSPKTCMKKGKNTS--KSSTEELFGYKVSNKMIKTKLTYQSVIPTNCTA 167
>RFCS_METJA (Q58817) Replication factor C small subunit (RFC small
subunit) (Clamp loader small subunit) [Contains: Mja
RFC-1 intein; Mja RFC-2 intein; Mja RFC-3 intein]
Length = 1847
Score = 31.2 bits (69), Expect = 6.2
Identities = 40/184 (21%), Positives = 79/184 (42%), Gaps = 35/184 (19%)
Query: 56 WFQLNEESSGNQKPFSLRKGFLESVKLDAKRVLEILRQDGPGLDARVVLDELQVRPSGLL 115
++ + +S N + F GF + KL K++LEI++ + P LD + +D+ ++R
Sbjct: 327 YYHIVISNSSNLRTFLDNIGFSQERKL--KKLLEIIKDENPNLDV-ITIDKEKIR----Y 379
Query: 116 VREVL-VGILKSINHEN--RTRCAKLAYKFFVWCSQQEAYQHTANAYHLIMNIYAECEEY 172
+R+ L V + + I +N +C K+ + L+ IY EE
Sbjct: 380 IRDRLKVKLTRDIEKDNWSYNKCRKITQE-------------------LLKEIYYRLEEL 420
Query: 173 KALWRLVGEMIEKGFPATARTFNILIRTCGEAGLAKDLVETFIKSKTFNYRPFKHSYNAI 232
K + + + E I + A + + G+ D + +I+ K +P +Y I
Sbjct: 421 KEIEKALEENILIDWDEVAERRKEIAE---KTGIRSDRILEYIRGKR---KPSLKNYIKI 474
Query: 233 LHSL 236
++L
Sbjct: 475 ANTL 478
>ZBT1_MOUSE (Q91VL9) Zinc finger and BTB domain containing protein 1
Length = 713
Score = 30.8 bits (68), Expect = 8.1
Identities = 12/31 (38%), Positives = 20/31 (63%)
Query: 135 CAKLAYKFFVWCSQQEAYQHTANAYHLIMNI 165
C KL + + C+ + +YQ+T AYH I++I
Sbjct: 227 CEKLLDEHVLTCTNRHSYQNTTRAYHRIVDI 257
>YWFF_LACLA (Q9CDP0) Hypothetical RNA methyltransferase ywfF (EC
2.1.1.-)
Length = 513
Score = 30.8 bits (68), Expect = 8.1
Identities = 30/118 (25%), Positives = 51/118 (42%), Gaps = 2/118 (1%)
Query: 79 SVKLDAKRVLEILRQDGPGLDARVVLDELQVRPSGLLVREVLVGILKSINHENRTRCAKL 138
++K+ K LEI R G V+ L P L EVLV I ++ + +R + K+
Sbjct: 3 NLKIGQKLQLEIERMGINGEGIGVISGRLVFIPYALPGEEVLVEITENARNFSRAKLVKI 62
Query: 139 AYKFFVWCSQQEAYQHTANAYHLIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNI 196
K Q+ H + H++ Y E+K ++ + +EK PA R + +
Sbjct: 63 IEKSPNRVKPQDRAYHEMSQSHIMHLSYPMQLEFKR--DVMRQALEKYKPAGWRNYEL 118
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.139 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 56,131,897
Number of Sequences: 164201
Number of extensions: 2370582
Number of successful extensions: 4944
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 4902
Number of HSP's gapped (non-prelim): 45
length of query: 480
length of database: 59,974,054
effective HSP length: 114
effective length of query: 366
effective length of database: 41,255,140
effective search space: 15099381240
effective search space used: 15099381240
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0148.13


