

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0148.11
(156 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
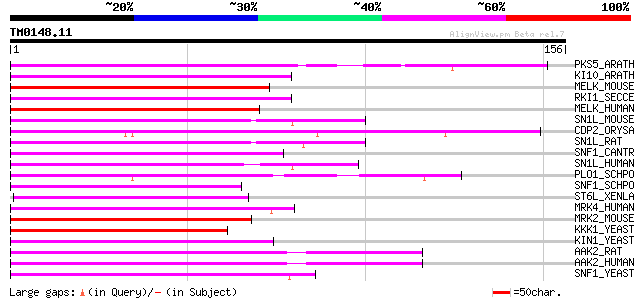
Score E
Sequences producing significant alignments: (bits) Value
PKS5_ARATH (O22932) SOS2-like protein kinase PKS5 (EC 2.7.1.37) 94 1e-19
KI10_ARATH (Q38997) SNF1-related protein kinase KIN10 (EC 2.7.1.... 66 4e-11
MELK_MOUSE (Q61846) Maternal embryonic leucine zipper kinase (EC... 65 5e-11
RKI1_SECCE (Q02723) Carbon catabolite derepressing protein kinas... 65 6e-11
MELK_HUMAN (Q14680) Maternal embryonic leucine zipper kinase (EC... 64 2e-10
SN1L_MOUSE (Q60670) Probable serine/threonine-protein kinase SNF... 59 3e-09
CDP2_ORYSA (P53683) Calcium-dependent protein kinase, isoform 2 ... 58 7e-09
SN1L_RAT (Q9R1U5) Probable serine/threonine-protein kinase SNF1L... 57 1e-08
SNF1_CANTR (O94168) Carbon catabolite derepressing protein kinas... 57 2e-08
SN1L_HUMAN (P57059) Probable serine/threonine-protein kinase SNF... 57 2e-08
PLO1_SCHPO (P50528) Serine/threonine-protein kinase plo1 (EC 2.7... 56 3e-08
SNF1_SCHPO (O74536) SNF1-like protein kinase ssp2 (EC 2.7.1.-) 56 4e-08
ST6L_XENLA (Q91819) Serine/threonine-protein kinase Eg2-like (EC... 55 6e-08
MRK4_HUMAN (Q96L34) MAP/microtubule affinity-regulating kinase 4... 55 6e-08
MRK2_MOUSE (Q05512) MAP/microtubule affinity-regulating kinase 2... 55 6e-08
KKK1_YEAST (P34244) Probable serine/threonine-protein kinase YKL... 55 6e-08
KIN1_YEAST (P13185) Protein kinase KIN1 (EC 2.7.1.-) 55 8e-08
AAK2_RAT (Q09137) 5'-AMP-activated protein kinase, catalytic alp... 55 8e-08
AAK2_HUMAN (P54646) 5'-AMP-activated protein kinase, catalytic a... 55 8e-08
SNF1_YEAST (P06782) Carbon catabolite derepressing protein kinas... 54 1e-07
>PKS5_ARATH (O22932) SOS2-like protein kinase PKS5 (EC 2.7.1.37)
Length = 435
Score = 94.0 bits (232), Expect = 1e-19
Identities = 57/154 (37%), Positives = 83/154 (53%), Gaps = 13/154 (8%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPF N+M MY K+ + E++FP W SP+ K+ +S++L +P RITI I+
Sbjct: 213 LFVLVAGYLPFNDPNVMNMYKKIYKGEYRFPRWMSPDLKRFVSRLLDINPETRITIDEIL 272
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
+ WF +G I D + + + SS E A K +NAF+ I S SSG DLS
Sbjct: 273 KDPWFVRGGFKQIKFHDDEIE--DQKVESSLE-------AVKSLNAFDLI-SYSSGLDLS 322
Query: 121 GFF---EEKRRGGSVFTSKCSVSEIASKIEGAAK 151
G F F S+ S +A ++EG A+
Sbjct: 323 GLFAGCSNSSGESERFLSEKSPEMLAEEVEGFAR 356
>KI10_ARATH (Q38997) SNF1-related protein kinase KIN10 (EC 2.7.1.-)
(AKIN10)
Length = 535
Score = 65.9 bits (159), Expect = 4e-11
Identities = 31/79 (39%), Positives = 44/79 (55%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LYALL G LPF EN+ ++ K+ + P SP ++ LI ++LV DP R+TI I
Sbjct: 230 LYALLCGTLPFDDENIPNLFKKIKGGIYTLPSHLSPGARDLIPRMLVVDPMKRVTIPEIR 289
Query: 61 RVSWFQKGFSASIPIPDPD 79
+ WFQ + +P PD
Sbjct: 290 QHPWFQAHLPRYLAVPPPD 308
>MELK_MOUSE (Q61846) Maternal embryonic leucine zipper kinase (EC
2.7.1.37) (Protein kinase PK38) (mPK38)
Length = 643
Score = 65.5 bits (158), Expect = 5e-11
Identities = 25/73 (34%), Positives = 45/73 (61%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY L+ G LPF +N+M +Y K++R +++ P W SP S L+ ++L DP RI++ +++
Sbjct: 199 LYVLMCGFLPFDDDNVMALYKKIMRGKYEVPKWLSPSSILLLQQMLQVDPKKRISMRNLL 258
Query: 61 RVSWFQKGFSASI 73
W + +S +
Sbjct: 259 NHPWVMQDYSCPV 271
>RKI1_SECCE (Q02723) Carbon catabolite derepressing protein kinase
(EC 2.7.1.-)
Length = 502
Score = 65.1 bits (157), Expect = 6e-11
Identities = 30/79 (37%), Positives = 45/79 (55%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LYALL G +PF +N+ ++ K+ + P + S + LIS++L+ DP RITI I
Sbjct: 205 LYALLCGAVPFDDDNIPNLFKKIKGGTYILPIYLSDLVRDLISRMLIVDPMKRITIGEIR 264
Query: 61 RVSWFQKGFSASIPIPDPD 79
+ SWFQ + +P PD
Sbjct: 265 KHSWFQNRLPRYLAVPPPD 283
>MELK_HUMAN (Q14680) Maternal embryonic leucine zipper kinase (EC
2.7.1.37) (hMELK) (Protein kinase PK38) (hPK38)
Length = 651
Score = 63.5 bits (153), Expect = 2e-10
Identities = 24/70 (34%), Positives = 43/70 (61%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY L+ G LPF +N+M +Y K++R ++ P W SP S L+ ++L DP RI++ +++
Sbjct: 199 LYVLMCGFLPFDDDNVMALYKKIMRGKYDVPKWLSPSSILLLQQMLQVDPKKRISMKNLL 258
Query: 61 RVSWFQKGFS 70
W + ++
Sbjct: 259 NHPWIMQDYN 268
>SN1L_MOUSE (Q60670) Probable serine/threonine-protein kinase SNF1LK
(EC 2.7.1.37) (HRT-20) (Myocardial SNF1-like kinase)
Length = 779
Score = 59.3 bits (142), Expect = 3e-09
Identities = 36/102 (35%), Positives = 53/102 (51%), Gaps = 3/102 (2%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY L+ G LPF NL T+ +VL F+ P + S + + LI ++LV DP RITI+ I
Sbjct: 214 LYVLVCGSLPFDGPNLPTLRQRVLEGRFRIPFFMSQDCETLIRRMLVVDPAKRITIAQIR 273
Query: 61 RVSWFQKGFSASIPIPDP--DESNFNSDLNSSSEQSTKVVAA 100
+ W Q + DP D + S+L +EQ ++ A
Sbjct: 274 QHRWMQAD-PTLLQQDDPAFDMQGYTSNLGDYNEQVLGIMQA 314
>CDP2_ORYSA (P53683) Calcium-dependent protein kinase, isoform 2 (EC
2.7.1.-) (CDPK 2)
Length = 533
Score = 58.2 bits (139), Expect = 7e-09
Identities = 43/159 (27%), Positives = 73/159 (45%), Gaps = 10/159 (6%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFP--PW--FSPESKKLISKILVADPNLRITI 56
LY LL+G PF E +++ +L+ E F PW S +K L+ K+L DP RIT
Sbjct: 275 LYILLSGVPPFWAETEKGIFDAILQGEIDFESQPWPSISESAKDLVRKMLTQDPKKRITS 334
Query: 57 SSIMRVSWFQKGFSASIPIPDPDESNFNS--DLNSSSEQSTKVVAAAKFINAFEFISSMS 114
+ +++ W + G ++ PI S +N + + KV+A+ + + M
Sbjct: 335 AQVLQHPWLRDGEASDKPIDSAVLSRMKQFRAMNKLKKMALKVIASNLNEEEIKGLKQMF 394
Query: 115 SGFDLSG----FFEEKRRGGSVFTSKCSVSEIASKIEGA 149
+ D +EE + G + SK S +E+ +E A
Sbjct: 395 TNMDTDNSGTITYEELKAGLAKLGSKLSEAEVKQLMEAA 433
>SN1L_RAT (Q9R1U5) Probable serine/threonine-protein kinase SNF1LK
(EC 2.7.1.37) (Salt-inducible protein kinase) (Protein
kinase KID2)
Length = 776
Score = 57.4 bits (137), Expect = 1e-08
Identities = 36/102 (35%), Positives = 53/102 (51%), Gaps = 3/102 (2%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY L+ G LPF NL T+ +VL F+ P + S + + LI ++LV DP RITI+ I
Sbjct: 214 LYVLVCGSLPFDGPNLPTLRQRVLEGRFRIPFFMSQDCETLIRRMLVVDPAKRITIAQIR 273
Query: 61 RVSWFQKGFSASIPIPDPDES--NFNSDLNSSSEQSTKVVAA 100
+ W Q + DP S + S+L +EQ ++ A
Sbjct: 274 QHRWMQAD-PTLLQQDDPAFSMQGYTSNLGDYNEQVLGIMQA 314
>SNF1_CANTR (O94168) Carbon catabolite derepressing protein kinase
(EC 2.7.1.-)
Length = 619
Score = 56.6 bits (135), Expect = 2e-08
Identities = 27/77 (35%), Positives = 43/77 (55%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY +L G LPF E + ++ K+ + P + SP +K L++++LV +P RITI IM
Sbjct: 239 LYVMLCGRLPFDDEFIPALFKKISNGVYTLPNYLSPGAKHLLTRMLVVNPLNRITIHEIM 298
Query: 61 RVSWFQKGFSASIPIPD 77
WF++ + PD
Sbjct: 299 EDEWFKQDMPDYLLPPD 315
>SN1L_HUMAN (P57059) Probable serine/threonine-protein kinase SNF1LK
(EC 2.7.1.37)
Length = 786
Score = 56.6 bits (135), Expect = 2e-08
Identities = 34/103 (33%), Positives = 53/103 (51%), Gaps = 9/103 (8%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY L+ G LPF NL T+ +VL F+ P + S + + LI ++LV DP RITI+ I
Sbjct: 217 LYVLVCGSLPFDGPNLPTLRQRVLEGRFRIPFFMSQDCESLIRRMLVVDPARRITIAQIR 276
Query: 61 RVSWFQKGFSASIPIPDP-----DESNFNSDLNSSSEQSTKVV 98
+ W + A +P P ++ S+L EQ+ ++
Sbjct: 277 QHRWMR----AEPCLPGPACPAFSAHSYTSNLGDYDEQALGIM 315
>PLO1_SCHPO (P50528) Serine/threonine-protein kinase plo1 (EC
2.7.1.37)
Length = 683
Score = 56.2 bits (134), Expect = 3e-08
Identities = 42/130 (32%), Positives = 66/130 (50%), Gaps = 12/130 (9%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPW--FSPESKKLISKILVADPNLRITISS 58
+YALL G PFQ + + T+Y K+ + FP S E+K LIS +L DP++R +I
Sbjct: 230 MYALLIGKPPFQDKEVKTIYRKIKANSYSFPSNVDISAEAKDLISSLLTHDPSIRPSIDD 289
Query: 59 IMRVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSS-GF 117
I+ +F G+ AS PDE + + SS+ + + F +F++S S GF
Sbjct: 290 IVDHEFFHTGYMASTL---PDEILHSMPIWPSSQ------SKSSFQRNLDFVASASGVGF 340
Query: 118 DLSGFFEEKR 127
S E+ +
Sbjct: 341 GNSAGVEKNK 350
>SNF1_SCHPO (O74536) SNF1-like protein kinase ssp2 (EC 2.7.1.-)
Length = 576
Score = 55.8 bits (133), Expect = 4e-08
Identities = 27/65 (41%), Positives = 38/65 (57%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY +L G LPF E + ++ KV + P + SP ++ LI +++VADP RITI I
Sbjct: 221 LYVMLVGRLPFDDEFIPNLFKKVNSCVYVMPDFLSPGAQSLIRRMIVADPMQRITIQEIR 280
Query: 61 RVSWF 65
R WF
Sbjct: 281 RDPWF 285
>ST6L_XENLA (Q91819) Serine/threonine-protein kinase Eg2-like (EC
2.7.1.37) (p46XlEg22)
Length = 408
Score = 55.1 bits (131), Expect = 6e-08
Identities = 23/66 (34%), Positives = 38/66 (56%)
Query: 2 YALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIMR 61
Y L G PF+ + Y ++ + EFQ+PP+ S E+K L+SK+L +PN R+ + ++
Sbjct: 327 YEFLVGKPPFETDTHQETYRRISKVEFQYPPYVSEEAKDLVSKLLKHNPNHRLPLKGVLE 386
Query: 62 VSWFQK 67
W K
Sbjct: 387 HPWIVK 392
>MRK4_HUMAN (Q96L34) MAP/microtubule affinity-regulating kinase 4
(EC 2.7.1.37) (MAP/microtubule affinity-regulating
kinase like 1)
Length = 752
Score = 55.1 bits (131), Expect = 6e-08
Identities = 26/82 (31%), Positives = 45/82 (54%), Gaps = 2/82 (2%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY L++G LPF NL + +VLR +++ P + S + + ++ + LV +P R T+ IM
Sbjct: 246 LYTLVSGSLPFDGHNLKELRERVLRGKYRVPFYMSTDCESILRRFLVLNPAKRCTLEQIM 305
Query: 61 RVSWFQKGFSAS--IPIPDPDE 80
+ W G+ P +P+E
Sbjct: 306 KDKWINIGYEGEELKPYTEPEE 327
>MRK2_MOUSE (Q05512) MAP/microtubule affinity-regulating kinase 2
(EC 2.7.1.37) (Serine/threonine-protein kinase Emk)
Length = 774
Score = 55.1 bits (131), Expect = 6e-08
Identities = 24/68 (35%), Positives = 41/68 (60%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY L++G LPF +NL + +VLR +++ P + S + + L+ K L+ +P+ R T+ IM
Sbjct: 240 LYTLVSGSLPFDGQNLKELRERVLRGKYRIPFYMSTDCENLLKKFLILNPSKRGTLEQIM 299
Query: 61 RVSWFQKG 68
+ W G
Sbjct: 300 KDRWMNVG 307
>KKK1_YEAST (P34244) Probable serine/threonine-protein kinase
YKL101W (EC 2.7.1.37)
Length = 1518
Score = 55.1 bits (131), Expect = 6e-08
Identities = 28/61 (45%), Positives = 38/61 (61%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ALL G LPF +N+ + KV ++Q P S E++ LISKILV DP RIT I+
Sbjct: 305 LFALLTGHLPFNDDNIKKLLLKVQSGKYQMPSNLSSEARDLISKILVIDPEKRITTQEIL 364
Query: 61 R 61
+
Sbjct: 365 K 365
>KIN1_YEAST (P13185) Protein kinase KIN1 (EC 2.7.1.-)
Length = 1064
Score = 54.7 bits (130), Expect = 8e-08
Identities = 24/74 (32%), Positives = 41/74 (54%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+ G +PF EN ++ K+ + + ++P S E L+SK+LV DP R T+ ++
Sbjct: 334 LFVLVCGKVPFDDENSSVLHEKIKQGKVEYPQHLSIEVISLLSKMLVVDPKRRATLKQVV 393
Query: 61 RVSWFQKGFSASIP 74
W +GF+ P
Sbjct: 394 EHHWMVRGFNGPPP 407
>AAK2_RAT (Q09137) 5'-AMP-activated protein kinase, catalytic
alpha-2 chain (EC 2.7.1.-) (AMPK alpha-2 chain)
Length = 552
Score = 54.7 bits (130), Expect = 8e-08
Identities = 34/116 (29%), Positives = 54/116 (46%), Gaps = 5/116 (4%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LYALL G LPF E++ T++ K+ F P + + L+ +L DP R TI I
Sbjct: 204 LYALLCGTLPFDDEHVPTLFKKIRGGVFYIPEYLNRSIATLLMHMLQVDPLKRATIKDIR 263
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSG 116
WF++ + + DP + D N +++ K V E ++S+ SG
Sbjct: 264 EHEWFKQDLPSYLFPEDP-----SYDANVIDDEAVKEVCEKFECTESEVMNSLYSG 314
>AAK2_HUMAN (P54646) 5'-AMP-activated protein kinase, catalytic
alpha-2 chain (EC 2.7.1.-) (AMPK alpha-2 chain)
Length = 552
Score = 54.7 bits (130), Expect = 8e-08
Identities = 34/116 (29%), Positives = 54/116 (46%), Gaps = 5/116 (4%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LYALL G LPF E++ T++ K+ F P + + L+ +L DP R TI I
Sbjct: 204 LYALLCGTLPFDDEHVPTLFKKIRGGVFYIPEYLNRSVATLLMHMLQVDPLKRATIKDIR 263
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSG 116
WF++ + + DP + D N +++ K V E ++S+ SG
Sbjct: 264 EHEWFKQDLPSYLFPEDP-----SYDANVIDDEAVKEVCEKFECTESEVMNSLYSG 314
>SNF1_YEAST (P06782) Carbon catabolite derepressing protein kinase
(EC 2.7.1.-)
Length = 633
Score = 54.3 bits (129), Expect = 1e-07
Identities = 28/90 (31%), Positives = 47/90 (52%), Gaps = 4/90 (4%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY +L LPF E++ ++ + + P + SP + LI ++L+ +P RI+I IM
Sbjct: 242 LYVMLCRRLPFDDESIPVLFKNISNGVYTLPKFLSPGAAGLIKRMLIVNPLNRISIHEIM 301
Query: 61 RVSWFQKGFSASIPIPD----PDESNFNSD 86
+ WF+ + PD P+E N N+D
Sbjct: 302 QDDWFKVDLPEYLLPPDLKPHPEEENENND 331
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.133 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,327,189
Number of Sequences: 164201
Number of extensions: 567242
Number of successful extensions: 2149
Number of sequences better than 10.0: 353
Number of HSP's better than 10.0 without gapping: 236
Number of HSP's successfully gapped in prelim test: 117
Number of HSP's that attempted gapping in prelim test: 1834
Number of HSP's gapped (non-prelim): 367
length of query: 156
length of database: 59,974,054
effective HSP length: 101
effective length of query: 55
effective length of database: 43,389,753
effective search space: 2386436415
effective search space used: 2386436415
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0148.11


