

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0143b.7
(485 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
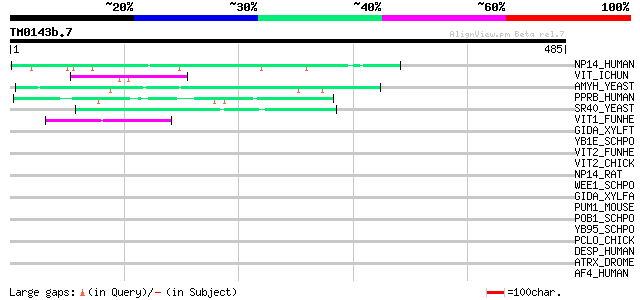
Score E
Sequences producing significant alignments: (bits) Value
NP14_HUMAN (Q14978) Nucleolar phosphoprotein p130 (Nucleolar 130... 52 3e-06
VIT_ICHUN (Q91062) Vitellogenin precursor (VTG) [Contains: Lipov... 47 1e-04
AMYH_YEAST (P08640) Glucoamylase S1/S2 precursor (EC 3.2.1.3) (G... 46 2e-04
PPRB_HUMAN (Q15648) Peroxisome proliferator-activated receptor b... 45 3e-04
SR40_YEAST (P32583) Suppressor protein SRP40 45 4e-04
VIT1_FUNHE (Q90508) Vitellogenin I precursor (VTG I) [Contains: ... 45 5e-04
GIDA_XYLFT (Q87DB3) Glucose inhibited division protein A 44 0.001
YB1E_SCHPO (P87179) Serine-rich protein C30B4.01c precursor 44 0.001
VIT2_FUNHE (Q98893) Vitellogenin II precursor (VTG II) [Contains... 42 0.005
VIT2_CHICK (P02845) Vitellogenin II precursor (Major vitellogeni... 42 0.005
NP14_RAT (P41777) Nucleolar phosphoprotein p130 (Nucleolar 130 k... 42 0.005
WEE1_SCHPO (P07527) Mitosis inhibitor protein kinase wee1 (EC 2.... 41 0.006
GIDA_XYLFA (Q9PBN4) Glucose inhibited division protein A 40 0.018
PUM1_MOUSE (Q80U78) Pumilio homolog 1 39 0.023
POB1_SCHPO (O74653) Pob1 protein (BOI protein homolog) 38 0.051
YB95_SCHPO (O42970) Hypothetical serine-rich protein C1E8.05 in ... 38 0.067
PCLO_CHICK (Q9PU36) Piccolo protein (Aczonin) (Fragment) 38 0.067
DESP_HUMAN (P15924) Desmoplakin (DP) (250/210 kDa paraneoplastic... 38 0.067
ATRX_DROME (Q9GQN5) Transcriptional regulator ATRX homolog (X-li... 38 0.067
AF4_HUMAN (P51825) AF-4 protein (Proto-oncogene AF4) (FEL protein) 38 0.067
>NP14_HUMAN (Q14978) Nucleolar phosphoprotein p130 (Nucleolar 130
kDa protein) (140 kDa nucleolar phosphoprotein)
(Nopp140) (Nucleolar and coiled-body phosphoprotein 1)
Length = 699
Score = 52.0 bits (123), Expect = 3e-06
Identities = 82/361 (22%), Positives = 135/361 (36%), Gaps = 27/361 (7%)
Query: 2 PKSRKRGATPTTFASQ-----KRAKGAGELTVGDPSPAKSASQRVEMQQGGDQ--GGTTP 54
PK++K TP T +Q K A+ A ++ G + + S+S ++ TP
Sbjct: 180 PKNQKPKITPVTVKAQTKAPPKPARAAPKIANGKAASSSSSSSSSSSSDDSEEEKAAATP 239
Query: 55 ---IPVEMALAKKTSRKRSS--RKSGSS-SSHHSKSSTSSSPLK-ESGPKRVGMSKMAVP 107
+P + +AK + ++ RKS SS S + P+K + GP A P
Sbjct: 240 KKTVPKKQVVAKAPVKAATTPTRKSSSSEDSSSDEEEEQKKPMKNKPGPYSYAPPPSAPP 299
Query: 108 RATWFSSKPPPVVLSSSSSSLESSNSSLDSNESDPVWDKL--TFYFNSKPIAWQHPGCEP 165
++PP + +ESS S D ++S +K T SK P +
Sbjct: 300 PKKSLGTQPPKKAV-EKQQPVESSEDSSDESDSSSEEEKKPPTKAVVSKATTKPPPAKKA 358
Query: 166 YYTRECRDNLSNFPPQDAQNMPSPAREENDPHVEVSGGEDQVSPIIQPDQVIG---VMLP 222
+ + + +A + P+ + + V+ V P P Q +G +L
Sbjct: 359 AESSSDSSDSDSSEDDEAPSKPAGTTKNSSNKPAVTTKSPAVKPAAAPKQPVGGGQKLLT 418
Query: 223 PKADVPASTFSEPSSRQLQTSPQQLQSSPQQLQQGA--RVALLSPHAKGGSASSKMAASS 280
KAD +S SS + +T + P+ + A A +P S+S ++SS
Sbjct: 419 RKADSSSSEEESSSSEEEKTKKMVATTKPKATAKAALSLPAKQAPQGSRDSSSDSDSSSS 478
Query: 281 HTSGERLSALLVEDPLSIFQSFFDGTLDLESPPRQAETTETAQSGGVVDDSRIEEAFGKL 340
E+ S V+ Q G S P A+ + S D EE KL
Sbjct: 479 EEEEEKTSKSAVKKKP---QKVAGGA--APSKPASAKKGKAESSNSSSSDDSSEEEEEKL 533
Query: 341 K 341
K
Sbjct: 534 K 534
Score = 43.5 bits (101), Expect = 0.001
Identities = 61/269 (22%), Positives = 97/269 (35%), Gaps = 23/269 (8%)
Query: 17 QKRAKGAGELTVGDPSPAKSA-----SQRVEMQQGGDQGGTTPIPVEMALAKKTSRKRSS 71
QK K + P AKS+ S + + TP+ V+ R++
Sbjct: 147 QKGVKPQAKAAKAPPKKAKSSDSDSDSSSEDEPPKNQKPKITPVTVKAQTKAPPKPARAA 206
Query: 72 RK--SGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSS--SSS 127
K +G ++S S SS+SSS K K VP+ + P + + SSS
Sbjct: 207 PKIANGKAASSSSSSSSSSSSDDSEEEKAAATPKKTVPKKQVVAKAPVKAATTPTRKSSS 266
Query: 128 LESSNSSLDSNESDPVWDKLTFYFNSKPIAWQHPGCEPYYTRECRDNLSNFPPQDAQNMP 187
E S+S + + P+ +K Y + P P P + +L PP+ A
Sbjct: 267 SEDSSSDEEEEQKKPMKNKPGPYSYAPP-----PSAPP-----PKKSLGTQPPKKAVEKQ 316
Query: 188 SPAREENDPHVEV-SGGEDQVSPIIQPDQVIGVMLPP---KADVPASTFSEPSSRQLQTS 243
P D E S E++ P + PP KA +S S+ S + +
Sbjct: 317 QPVESSEDSSDESDSSSEEEKKPPTKAVVSKATTKPPPAKKAAESSSDSSDSDSSEDDEA 376
Query: 244 PQQLQSSPQQLQQGARVALLSPHAKGGSA 272
P + + + V SP K +A
Sbjct: 377 PSKPAGTTKNSSNKPAVTTKSPAVKPAAA 405
Score = 31.6 bits (70), Expect = 4.8
Identities = 42/153 (27%), Positives = 60/153 (38%), Gaps = 23/153 (15%)
Query: 15 ASQKRAKGAGELTV---GDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSRKR-- 69
A+ K+ G G+ + D S ++ S E ++ TT A K+
Sbjct: 404 AAPKQPVGGGQKLLTRKADSSSSEEESSSSEEEKTKKMVATTKPKATAKAALSLPAKQAP 463
Query: 70 -----SSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSS 124
SS S SSSS + TS S +K+ K G A P SKP S+
Sbjct: 464 QGSRDSSSDSDSSSSEEEEEKTSKSAVKKKPQKVAGG---AAP------SKPA----SAK 510
Query: 125 SSSLESSNSSLDSNESDPVWDKLTFYFNSKPIA 157
ESSNSS + S+ +KL + +P A
Sbjct: 511 KGKAESSNSSSSDDSSEEEEEKLKGKGSPRPQA 543
>VIT_ICHUN (Q91062) Vitellogenin precursor (VTG) [Contains:
Lipovitellin LV-1N; Lipovitellin LV-1C; Lipovitellin
LV-2]
Length = 1823
Score = 47.0 bits (110), Expect = 1e-04
Identities = 41/111 (36%), Positives = 52/111 (45%), Gaps = 9/111 (8%)
Query: 54 PIPVEMALAKKTSRKRSSRKSGSSSSHHSKSSTSSSPLKES----GPKRVGMS-----KM 104
P P + + +S SS S SSSS S SS+SSS +S K V S K
Sbjct: 1248 PRPARSSSSSSSSDSSSSSSSSSSSSSSSSSSSSSSSESKSLEWLAVKDVNQSAFYNFKY 1307
Query: 105 AVPRATWFSSKPPPVVLSSSSSSLESSNSSLDSNESDPVWDKLTFYFNSKP 155
R S + P SSSSSS SS+SS S++SD +F +SKP
Sbjct: 1308 VPQRKPQTSRRHTPASSSSSSSSSSSSSSSSSSSDSDMTVSAESFEKHSKP 1358
Score = 43.1 bits (100), Expect = 0.002
Identities = 43/143 (30%), Positives = 59/143 (41%), Gaps = 31/143 (21%)
Query: 29 GDPSPAKSASQRVEMQQGGDQGGT-TP---IPVEMALAKKTSRKRSSRKSGSSSSHHSKS 84
G S + S+S D+ G TP V +A + + ++R S SSSS S S
Sbjct: 1130 GSSSSSSSSSSSSSSSSSSDKSGKKTPRQGSTVNLAAKRASKKQRGKDSSSSSSSSSSSS 1189
Query: 85 STSSSPLKESGPKR---------VGMSKMAVPRATWFSS------------------KPP 117
+S SP K G KR +G + ++ SS KP
Sbjct: 1190 DSSKSPHKHGGAKRQHAGHGAPHLGPQSHSSSSSSSSSSSSSSASKSFSTVKPPMTRKPR 1249
Query: 118 PVVLSSSSSSLESSNSSLDSNES 140
P SSSSSS +SS+SS S+ S
Sbjct: 1250 PARSSSSSSSSDSSSSSSSSSSS 1272
Score = 40.4 bits (93), Expect = 0.010
Identities = 41/152 (26%), Positives = 65/152 (41%), Gaps = 15/152 (9%)
Query: 4 SRKRGATPTTFASQKRAKGAGELTVGDPSPAKSASQRV-EMQQGGDQGGTTPIPVEMALA 62
S ++ ++ +S + +G+ T S A++R + Q+G D ++ + +
Sbjct: 1132 SSSSSSSSSSSSSSSSSDKSGKKTPRQGSTVNLAAKRASKKQRGKDSSSSSSSSSSSSDS 1191
Query: 63 KKTSRKRSSRK--------------SGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPR 108
K+ K K S SSSS S SS+SSS K + M++ P
Sbjct: 1192 SKSPHKHGGAKRQHAGHGAPHLGPQSHSSSSSSSSSSSSSSASKSFSTVKPPMTRKPRPA 1251
Query: 109 ATWFSSKPPPVVLSSSSSSLESSNSSLDSNES 140
+ SS SSSSSS SS+SS S+ S
Sbjct: 1252 RSSSSSSSSDSSSSSSSSSSSSSSSSSSSSSS 1283
Score = 35.4 bits (80), Expect = 0.33
Identities = 31/78 (39%), Positives = 40/78 (50%), Gaps = 6/78 (7%)
Query: 63 KKTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLS 122
K + S S SSSS S SS+SS + P++ +A RA SK S
Sbjct: 1123 KDAKKPPGSSSSSSSSSSSSSSSSSSDKSGKKTPRQGSTVNLAAKRA----SKKQRGKDS 1178
Query: 123 SSSSSLESSNSSLDSNES 140
SSSSS SS+SS DS++S
Sbjct: 1179 SSSSS--SSSSSSDSSKS 1194
>AMYH_YEAST (P08640) Glucoamylase S1/S2 precursor (EC 3.2.1.3)
(Glucan 1,4-alpha-glucosidase) (1,4-alpha-D-glucan
glucohydrolase)
Length = 1367
Score = 46.2 bits (108), Expect = 2e-04
Identities = 69/332 (20%), Positives = 116/332 (34%), Gaps = 16/332 (4%)
Query: 6 KRGATPTTFASQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKT 65
K+ TP S + + V PS + + S + + + P+P + ++
Sbjct: 308 KKTTTPVPTPSSSTTESSSA-PVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTES 366
Query: 66 SRKRSSRKSGSSSSHHSKSST---SSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLS 122
S + + SSS SST SS+P+ S V +T SS P V S
Sbjct: 367 SSAPVTSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAP-VTSS 425
Query: 123 SSSSSLESSNSSLDSNESDPVWDKLTFYFNSKPIAWQHPGCEPYYTRECRDNLSNFPPQD 182
++ SS SS + S PV T +S P+ + + +
Sbjct: 426 TTESSSAPVTSSTTESSSAPVTSSTT-ESSSAPVPTPSSSTTESSSAPVTSSTTESSSAP 484
Query: 183 AQNMPSPAREENDPHVEVSGGEDQVSPIIQPDQVIGVMLPPKADVPASTFSEPSSRQLQT 242
S E + V S E +P+ P A P+S+ +E SS + +
Sbjct: 485 VPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPAPTPSSSTTESSSAPVTS 544
Query: 243 SPQQLQSSP-----QQLQQGARVALLSPHAKGGSA-----SSKMAASSHTSGERLSALLV 292
S + S+P + + + S + SA SS SS S+
Sbjct: 545 STTESSSAPVPTPSSSTTESSSTPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTT 604
Query: 293 EDPLSIFQSFFDGTLDLESPPRQAETTETAQS 324
E + + T + S P + TTE++ +
Sbjct: 605 ESSSAPAPTPSSSTTESSSAPVTSSTTESSSA 636
Score = 40.4 bits (93), Expect = 0.010
Identities = 63/307 (20%), Positives = 114/307 (36%), Gaps = 22/307 (7%)
Query: 31 PSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSRKRSSRKSGSSSSHHSKSST--SS 88
P+P+ S ++ ++ PV + + +S +S + SSS+ + S+T SS
Sbjct: 396 PTPSSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAPVTSSTTESSS 455
Query: 89 SPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSNSSLDSNESD------P 142
+P+ S V +T SS P V + SSS+ ESS++ + S+ ++ P
Sbjct: 456 APVPTPSSSTTESSSAPVTSSTTESSSAP--VPTPSSSTTESSSAPVTSSTTESSSAPVP 513
Query: 143 VWDKLTFYFNSKPIAWQHPGCEPYYTRECRDNLSNFPPQDAQNMPSPAREENDPHVEVSG 202
T +S P + + + S E + V S
Sbjct: 514 TPSSSTTESSSAPAPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSTPVTSST 573
Query: 203 GEDQVSPIIQPDQVIGVMLPPKADVPASTFSEPSSRQLQT---SPQQLQSSP--QQLQQG 257
E +P+ P P+S+ +E SS T S + S+P +
Sbjct: 574 TESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPAPTPSSSTTESSSAPVTSSTTES 633
Query: 258 ARVALLSPHAKGGSASSKMAASSHTSGERLSALLVEDPLSIFQSFFDGTLDLESPPRQAE 317
+ + +P + +SS + +S S+ V P S T + S P +
Sbjct: 634 SSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPSS-------STTESSSAPVTSS 686
Query: 318 TTETAQS 324
TTE++ +
Sbjct: 687 TTESSSA 693
Score = 35.4 bits (80), Expect = 0.33
Identities = 55/260 (21%), Positives = 102/260 (39%), Gaps = 23/260 (8%)
Query: 31 PSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSRKRSSRKSGSSSSHHSKSSTSSSP 90
P+P+ S ++ TT E + A + S+ +S S+ S + +SS+P
Sbjct: 639 PTPSSSTTESSSAPVPTPSSSTT----ESSSAPVPTPSSSTTESSSAPVTSSTTESSSAP 694
Query: 91 LKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESS------NSSLDSNESDPVW 144
+ S + + + P ++ S PV SSS++ SS +SS + S PV
Sbjct: 695 VTSSTTES-SSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVT 753
Query: 145 DKLTFYFNSKPIAWQHPGCEPYYTRECRDNLSNFPPQDAQNMPSPAREENDPHVEVSGGE 204
T +S P+ + S+ + +P+P+ S E
Sbjct: 754 SSTT-ESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPSS---------STTE 803
Query: 205 DQVSPIIQPDQVIGVMLPPKADVPASTFSEPSSRQLQT-SPQQLQSSPQQLQQGARVALL 263
V+P+ P + + P S+ +E SS + T S +SS + + +
Sbjct: 804 SSVAPVPTPSSSSNITSSAPSSTPFSSSTESSSVPVPTPSSSTTESSSAPVSSSTTESSV 863
Query: 264 SPHAKGGSASSKMAASSHTS 283
+P S+SS + +S+ +S
Sbjct: 864 AP-VPTPSSSSNITSSAPSS 882
Score = 32.3 bits (72), Expect = 2.8
Identities = 53/278 (19%), Positives = 101/278 (36%), Gaps = 30/278 (10%)
Query: 50 GGTTPIPVEMALAKKTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRA 109
GGT + ++ ++ S+ +S +++S S+SST++S ES +
Sbjct: 206 GGTKSSTTTSSTSESSTTTSSTSESSTTTSSTSESSTTTSSTSESSTSSSTTAPATPTTT 265
Query: 110 TWFSSKPPPVVLSSSSSSLESSNSSLDSNESDPVWDKLTFYFNSKPIAWQHPGCEPYYTR 169
+ KP P +++S + +++ P K T + C T
Sbjct: 266 SCTKEKPTP----PTTTSCTKEKPTPPHHDTTPCTKKKTTTSKT---------CTKKTTT 312
Query: 170 ECRDNLSNFPPQDAQNMPSPA---REENDPHVEVSGGEDQVSPIIQPDQVIGVMLPPKAD 226
S+ + +P+P+ E + V S E +P+ P A
Sbjct: 313 PVPTPSSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSS--STTESSSAP 370
Query: 227 VPASTFSEPSSRQLQTSPQQLQSSPQQLQQGARVALLSPHAKGGSASSKMAASSHTSGER 286
V +ST +E SS + +S + S+P + S + S A + ++ E
Sbjct: 371 VTSST-TESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVTSSTTES 429
Query: 287 LSALLVEDPLSIFQSFFDGTLDLESPPRQAETTETAQS 324
SA + T + S P + TTE++ +
Sbjct: 430 SSAPVT-----------SSTTESSSAPVTSSTTESSSA 456
>PPRB_HUMAN (Q15648) Peroxisome proliferator-activated receptor
binding protein (PBP) (PPAR binding protein) (Thyroid
hormone receptor-associated protein complex 220 kDa
component) (Trap220) (Thyroid receptor interacting
protein 2) (TRIP2) (p53 regulatory
Length = 1581
Score = 45.4 bits (106), Expect = 3e-04
Identities = 75/291 (25%), Positives = 107/291 (35%), Gaps = 48/291 (16%)
Query: 4 SRKRGATPTTFASQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAK 63
S R TP T ++ G+ + P A ++ +Q IP +
Sbjct: 1026 SSNRPFTPPTSTGGSKSPGSAGRSQTPPGVATPPIPKITIQ----------IPKGTVMVG 1075
Query: 64 KTSRKRSSRKSGS-----SSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPP 118
K S SGS S SHHS SS+SSS SG + S+ + SSK
Sbjct: 1076 KPSSHSQYTSSGSVSSSGSKSHHSHSSSSSSSASTSGKMKSSKSEGS------SSSK--- 1126
Query: 119 VVLSSSSSSLESSNSSLDSNESDPVWDKLTFYFNSKPIAWQHPGCEPYYTRECRDNLSN- 177
LSSS S + S+ S S S K PG P S+
Sbjct: 1127 --LSSSMYSSQGSSGSSQSKNSSQSGGK--------------PGSSPITKHGLSSGSSST 1170
Query: 178 -FPPQDAQNM---PSPAREENDP-HVEVSGGEDQVSPIIQPDQVIGVMLPPKADVPASTF 232
PQ + PS ++ P H GG D+++ ++P V G KA P S+
Sbjct: 1171 KMKPQGKPSSLMNPSLSKPNISPSHSRPPGGSDKLASPMKP--VPGTPPSSKAKSPISSG 1228
Query: 233 SEPSSRQLQTSPQQLQSSPQQLQQGARVALLSPHAKGGSASSKMAASSHTS 283
S S +S ++SS G+ P + +ASS +SS +S
Sbjct: 1229 SGGSHMSGTSSSSGMKSSSGLGSSGSLSQKTPPSSNSCTASSSSFSSSGSS 1279
Score = 35.0 bits (79), Expect = 0.43
Identities = 25/76 (32%), Positives = 36/76 (46%), Gaps = 8/76 (10%)
Query: 65 TSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSS 124
+S+ +S SGS SH S +S+SS SG G P P ++S
Sbjct: 1218 SSKAKSPISSGSGGSHMSGTSSSSGMKSSSGLGSSGSLSQKTP--------PSSNSCTAS 1269
Query: 125 SSSLESSNSSLDSNES 140
SSS SS SS+ S+++
Sbjct: 1270 SSSFSSSGSSMSSSQN 1285
>SR40_YEAST (P32583) Suppressor protein SRP40
Length = 406
Score = 45.1 bits (105), Expect = 4e-04
Identities = 54/228 (23%), Positives = 86/228 (37%), Gaps = 7/228 (3%)
Query: 58 EMALAKKTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPP 117
++++ +K ++SS S SSSS S SS+SSS SG S + ++ S
Sbjct: 13 KLSVKEKEIEEKSSSSSSSSSSSSSSSSSSSSSSSSSGESSSSSSSSSSSSSSDSSDSSD 72
Query: 118 PVVLSSSSSSLESSNSSLDSNESDPVWDKLTFYFNSKPIAWQHPGCEPYYTRECRDNLSN 177
SSSSSS SS+SS DS S + +S + E E +
Sbjct: 73 SESSSSSSSSSSSSSSSSDSESSSESDSSSSGSSSSSSSSSDESSSESESEDETKKRARE 132
Query: 178 FPPQDAQNMPSPAREENDPHVEVSGGEDQVSPIIQPDQVIGVMLPPKADVPASTFSEPSS 237
+DA+ + + +P S + G ++D +S+ S SS
Sbjct: 133 SDNEDAK---ETKKAKTEPESSSSSESSSSGSSSSSESESG----SESDSDSSSSSSSSS 185
Query: 238 RQLQTSPQQLQSSPQQLQQGARVALLSPHAKGGSASSKMAASSHTSGE 285
S QSS + S + S S ++SS +S +
Sbjct: 186 DSESDSESDSQSSSSSSSSDSSSDSDSSSSDSSSDSDSSSSSSSSSSD 233
Score = 37.4 bits (85), Expect = 0.087
Identities = 36/141 (25%), Positives = 62/141 (43%), Gaps = 5/141 (3%)
Query: 4 SRKRGATPTTFASQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAK 63
S ++ ++ +S + +GE + S + S+S + D ++ + +
Sbjct: 30 SSSSSSSSSSSSSSSSSSSSGESSSSSSSSSSSSSS--DSSDSSDSESSSSSSSSSSSSS 87
Query: 64 KTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLS- 122
+S SS +S SSSS S SS+SSS S + +K + +K +
Sbjct: 88 SSSDSESSSESDSSSSGSSSSSSSSSDESSSESESEDETKKRARESDNEDAKETKKAKTE 147
Query: 123 --SSSSSLESSNSSLDSNESD 141
SSSSS SS+ S S+ES+
Sbjct: 148 PESSSSSESSSSGSSSSSESE 168
Score = 36.6 bits (83), Expect = 0.15
Identities = 63/311 (20%), Positives = 112/311 (35%), Gaps = 31/311 (9%)
Query: 25 ELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSRKRSSRKSGSSSSHHSKS 84
E + + S + S+S ++ E + + +S SS S SS S S
Sbjct: 18 EKEIEEKSSSSSSSSSSSSSSSSSSSSSSSSSGESSSSSSSSSSSSSSDSSDSSDSESSS 77
Query: 85 STSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSNSSLDSNESDPVW 144
S+SSS S S + ++ SS SSSSSS ESS+ S +E
Sbjct: 78 SSSSSSSSSSSSSDSESSSESDSSSSGSSS-------SSSSSSDESSSESESEDE----- 125
Query: 145 DKLTFYFNSKPIAWQHPGCEPYYTRECRDNLSNFPPQDAQNMPSPAREENDPHVEVSGGE 204
+K A + + T++ + + ++ + S + E++ E S +
Sbjct: 126 --------TKKRARESDNEDAKETKKAKTEPESSSSSESSSSGSSSSSESESGSE-SDSD 176
Query: 205 DQVSPIIQPDQVIGVMLPPKADVPASTFSEPSSRQLQTSPQQLQSSPQQLQQGARVALLS 264
S D ++D + + S SS +S SS + S
Sbjct: 177 SSSSSSSSSDS--------ESDSESDSQSSSSSSSSDSSSDSDSSSSDSSSDSDSSSSSS 228
Query: 265 PHAKGGSASSKMAASSHTSGERLSALLVEDPLSIFQSFFDGTLDLESPPRQAETTETAQS 324
+ + S ++ S +SG S+ + S +S + D +S ++E
Sbjct: 229 SSSSDSDSDSDSSSDSDSSGSSDSSSSSDS--SSDESTSSDSSDSDSDSDSGSSSELETK 286
Query: 325 GGVVDDSRIEE 335
D+S+ EE
Sbjct: 287 EATADESKAEE 297
Score = 32.3 bits (72), Expect = 2.8
Identities = 31/123 (25%), Positives = 49/123 (39%), Gaps = 11/123 (8%)
Query: 16 SQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSRKRSSRKSG 75
++KRA+ + + AK+ + + G ++ +S S +S
Sbjct: 126 TKKRARESDNEDAKETKKAKTEPESSSSSESSSSGSSS-----------SSESESGSESD 174
Query: 76 SSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSNSSL 135
S SS S SS+ S ES + S + + SS S SSSS SS+S
Sbjct: 175 SDSSSSSSSSSDSESDSESDSQSSSSSSSSDSSSDSDSSSSDSSSDSDSSSSSSSSSSDS 234
Query: 136 DSN 138
DS+
Sbjct: 235 DSD 237
>VIT1_FUNHE (Q90508) Vitellogenin I precursor (VTG I) [Contains:
Lipovitellin 1 (LV1); Phosvitin (PV); Lipovitellin 2
(LV2)]
Length = 1704
Score = 44.7 bits (104), Expect = 5e-04
Identities = 38/110 (34%), Positives = 52/110 (46%), Gaps = 1/110 (0%)
Query: 32 SPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSRKRSSRKSGSSSSHHSKSSTSSSPL 91
S + S S+ + +++A +S SSR+S SSSS S SS+SSS
Sbjct: 1089 SSSSSESRSSRSSSSSSSSSRSSRKIDLAARTNSSSSSSSRRSRSSSSS-SSSSSSSSSS 1147
Query: 92 KESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSNSSLDSNESD 141
S +R S + ++ SS+ SSSSSS SS SSL S SD
Sbjct: 1148 SSSSSRRSSSSSSSSSSSSSRSSRRVNSTRSSSSSSRTSSASSLASFFSD 1197
Score = 40.8 bits (94), Expect = 0.008
Identities = 38/104 (36%), Positives = 48/104 (45%), Gaps = 9/104 (8%)
Query: 38 SQRVEMQQGGDQGGTTPIPVEMALAK-KTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGP 96
S+ E ++GG PV + L K +SR+ SS S SSSS S+S +S S S
Sbjct: 1056 SEDEETEEGG--------PVLVKLNKILSSRRNSSSSSSSSSSSSSESRSSRSSSSSSSS 1107
Query: 97 KRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSNSSLDSNES 140
R R SS SSSSSS SS+SS S+ S
Sbjct: 1108 SRSSRKIDLAARTNSSSSSSSRRSRSSSSSSSSSSSSSSSSSSS 1151
Score = 40.4 bits (93), Expect = 0.010
Identities = 36/140 (25%), Positives = 68/140 (47%), Gaps = 12/140 (8%)
Query: 4 SRKRGATPTTFASQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAK 63
SR+ ++ ++ +S ++ + S + +S+++++ + ++ + +
Sbjct: 1077 SRRNSSSSSSSSSSSSSESRSSRSSSSSSSSSRSSRKIDLAARTNSSSSSSSRRSRSSSS 1136
Query: 64 KTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSS 123
+S SS S SSSS S SS+SSS S R ++ R++ SS+ +S
Sbjct: 1137 SSSSSSSSSSSSSSSSRRSSSSSSSSSSSSSRSSR----RVNSTRSSSSSSR------TS 1186
Query: 124 SSSSLES--SNSSLDSNESD 141
S+SSL S S+SS S+ SD
Sbjct: 1187 SASSLASFFSDSSSSSSSSD 1206
>GIDA_XYLFT (Q87DB3) Glucose inhibited division protein A
Length = 629
Score = 43.9 bits (102), Expect = 0.001
Identities = 48/202 (23%), Positives = 81/202 (39%), Gaps = 15/202 (7%)
Query: 294 DPLSIFQSFFDGTLDLESPPRQAETTETAQSGGVVDDSRIEEAFGKLKRLVFDAGFIDGI 353
+P +F S + L L A T + G+VDD+R K + + + G + +
Sbjct: 425 EPYRMFTSRAEYRLQLREDNADARLTAIGRDLGLVDDARWAHFNAKQEAVARECGRLSAL 484
Query: 354 RATP--QLGQEVKELLAFLLSQRLDPIQADALVELQTLLVNIMATLDKALAVEQLIETKK 411
ATP LG+EVKE L LS+ + I EL + + +L + Q+ E +
Sbjct: 485 WATPGNALGREVKETLGVTLSRETNIIDLMKRPELDYAALMRVPSLGPGVDDAQVAE--Q 542
Query: 412 AQVASNTNGLLPAKDEILKLTARKTLIAAEL-----------SSVDARLDELQREEVAAE 460
+++ G L + E + R A L + V +L +Q + V
Sbjct: 543 VEISVKYAGYLNRQSEEITRQQRHEATAIPLEFDYAAVRGLSTEVLQKLQHIQPQTVGQA 602
Query: 461 QRVKQSLDSRIAAARVQLEAFK 482
QR+ + I+ V LE +
Sbjct: 603 QRIPGMTPAAISLLLVHLERMR 624
>YB1E_SCHPO (P87179) Serine-rich protein C30B4.01c precursor
Length = 374
Score = 43.5 bits (101), Expect = 0.001
Identities = 38/121 (31%), Positives = 49/121 (40%), Gaps = 7/121 (5%)
Query: 22 GAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSRKRSSRKSGSSSSHH 81
G + TV S + + S TT P +S SS S SSSS
Sbjct: 121 GVLQTTVSSSSVSSTTSSSSSSSPSSSSTTTTTSP-------SSSSSSSSSSSSSSSSSS 173
Query: 82 SKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSNSSLDSNESD 141
S SS+SSS S S + ++ SS P +SSS S SS+SS S+ S
Sbjct: 174 SSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSVPITSSTSSSHSSSSSSSSSSSSSSR 233
Query: 142 P 142
P
Sbjct: 234 P 234
Score = 37.7 bits (86), Expect = 0.067
Identities = 33/139 (23%), Positives = 55/139 (38%), Gaps = 2/139 (1%)
Query: 4 SRKRGATPTTFASQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAK 63
S ++ TT S + + + S + S+S ++ + +
Sbjct: 142 SSPSSSSTTTTTSPSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSS 201
Query: 64 KTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFS--SKPPPVVL 121
+S S +SSSH S SS+SSS S P + +T+ S + P
Sbjct: 202 SSSSSSVPITSSTSSSHSSSSSSSSSSSSSSRPSSSSSFITTMSSSTFISTVTVTPSSSS 261
Query: 122 SSSSSSLESSNSSLDSNES 140
SS+SS + SS ++L N S
Sbjct: 262 SSTSSEVPSSTAALALNAS 280
Score = 37.7 bits (86), Expect = 0.067
Identities = 37/133 (27%), Positives = 53/133 (39%), Gaps = 15/133 (11%)
Query: 8 GATPTTFASQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSR 67
G TT +S + + PS + + + ++ + + +S
Sbjct: 121 GVLQTTVSSSSVSSTTSSSSSSSPSSSSTTTTTSPSSSSSSSSSSSSSSSSSSSSSSSSS 180
Query: 68 KRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSS 127
SS S SSSS S SS+SSS S +VP + SS SSSSSS
Sbjct: 181 SSSSSSSSSSSSSSSSSSSSSS-----------SSSSSVPITSSTSSSHS----SSSSSS 225
Query: 128 LESSNSSLDSNES 140
SS+SS S+ S
Sbjct: 226 SSSSSSSRPSSSS 238
Score = 36.6 bits (83), Expect = 0.15
Identities = 28/133 (21%), Positives = 57/133 (42%), Gaps = 4/133 (3%)
Query: 2 PKSRKRGATPTTFASQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMAL 61
P S ++ ++ +S + + + S + S+S ++ +P+ +
Sbjct: 155 PSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSVPITSST 214
Query: 62 AKKTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVL 121
+ S SS S SSSS S SS+ + + S +S + V ++ SS V
Sbjct: 215 SSSHSSSSSSSSSSSSSSRPSSSSSFITTMSSS----TFISTVTVTPSSSSSSTSSEVPS 270
Query: 122 SSSSSSLESSNSS 134
S+++ +L +S +S
Sbjct: 271 STAALALNASKAS 283
Score = 34.7 bits (78), Expect = 0.57
Identities = 29/83 (34%), Positives = 38/83 (44%)
Query: 74 SGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSNS 133
S SS S + SS+SSSP S S + ++ SS SSSSSS SS+S
Sbjct: 128 SSSSVSSTTSSSSSSSPSSSSTTTTTSPSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSS 187
Query: 134 SLDSNESDPVWDKLTFYFNSKPI 156
S S+ S + +S PI
Sbjct: 188 SSSSSSSSSSSSSSSSSSSSVPI 210
>VIT2_FUNHE (Q98893) Vitellogenin II precursor (VTG II) [Contains:
Lipovitellin 1 (LV1); Phosvitin (PV); Lipovitellin 2
(LV2)]
Length = 1687
Score = 41.6 bits (96), Expect = 0.005
Identities = 34/103 (33%), Positives = 44/103 (42%), Gaps = 6/103 (5%)
Query: 64 KTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSK------PP 117
K S K SS SGSS S S+SS+SSS S R S + +SK P
Sbjct: 1082 KNSTKASSSSSGSSRSSRSRSSSSSSSSSSSSSSRSSSSSSRSSSSLRRNSKMLDLADPL 1141
Query: 118 PVVLSSSSSSLESSNSSLDSNESDPVWDKLTFYFNSKPIAWQH 160
+ SSSS SS+SS S+ S K + + + H
Sbjct: 1142 NITSKRSSSSSSSSSSSSSSSSSSSSSSKTKWQLHERNFTKDH 1184
Score = 40.4 bits (93), Expect = 0.010
Identities = 34/83 (40%), Positives = 43/83 (50%), Gaps = 3/83 (3%)
Query: 62 AKKTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVL 121
+ ++SR RSS S SSSS S S+SSS S +R SKM + A +
Sbjct: 1094 SSRSSRSRSSSSSSSSSSSSSSRSSSSSSRSSSSLRR--NSKM-LDLADPLNITSKRSSS 1150
Query: 122 SSSSSSLESSNSSLDSNESDPVW 144
SSSSSS SS+SS S+ S W
Sbjct: 1151 SSSSSSSSSSSSSSSSSSSKTKW 1173
Score = 35.4 bits (80), Expect = 0.33
Identities = 30/85 (35%), Positives = 41/85 (47%), Gaps = 4/85 (4%)
Query: 60 ALAKKTSRKRSSRKSGSSSSHHSKSS----TSSSPLKESGPKRVGMSKMAVPRATWFSSK 115
A + + RSSR SSSS S SS +SSS + S R + + +SK
Sbjct: 1087 ASSSSSGSSRSSRSRSSSSSSSSSSSSSSRSSSSSSRSSSSLRRNSKMLDLADPLNITSK 1146
Query: 116 PPPVVLSSSSSSLESSNSSLDSNES 140
SSSSSS SS+SS S+++
Sbjct: 1147 RSSSSSSSSSSSSSSSSSSSSSSKT 1171
>VIT2_CHICK (P02845) Vitellogenin II precursor (Major vitellogenin)
[Contains: Lipovitellin I (LVI); Phosvitin (PV);
Lipovitellin II (LVII); YGP40]
Length = 1850
Score = 41.6 bits (96), Expect = 0.005
Identities = 38/143 (26%), Positives = 59/143 (40%), Gaps = 10/143 (6%)
Query: 8 GATPTTFASQKRAKGAGELTVGDPSPAKSASQR----------VEMQQGGDQGGTTPIPV 57
G P S + + T S A S +++ V+ + D ++
Sbjct: 1115 GTEPDAKTSSSSSSASSTATSSSSSSASSPNRKKPMDEEENDQVKQARNKDASSSSRSSK 1174
Query: 58 EMALAKKTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPP 117
+K++S K S+ SSSS S SS+S S S S + +++ SS+
Sbjct: 1175 SSNSSKRSSSKSSNSSKRSSSSSSSSSSSSRSSSSSSSSSSNSKSSSSSSKSSSSSSRSR 1234
Query: 118 PVVLSSSSSSLESSNSSLDSNES 140
SSSSSS SS+SS S+ S
Sbjct: 1235 SSSKSSSSSSSSSSSSSSKSSSS 1257
Score = 39.7 bits (91), Expect = 0.018
Identities = 33/78 (42%), Positives = 39/78 (49%), Gaps = 3/78 (3%)
Query: 66 SRKRSSRKSGSSSSHH---SKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLS 122
S KRSS S SSSS S SS+SSS K S S + R++ SS S
Sbjct: 1189 SSKRSSSSSSSSSSSSRSSSSSSSSSSNSKSSSSSSKSSSSSSRSRSSSKSSSSSSSSSS 1248
Query: 123 SSSSSLESSNSSLDSNES 140
SSSS SS SS S++S
Sbjct: 1249 SSSSKSSSSRSSSSSSKS 1266
Score = 39.3 bits (90), Expect = 0.023
Identities = 28/79 (35%), Positives = 38/79 (47%)
Query: 62 AKKTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVL 121
+ +S++ SS S SSSS S SS+SSS S + R+ S
Sbjct: 1186 SSNSSKRSSSSSSSSSSSSRSSSSSSSSSSNSKSSSSSSKSSSSSSRSRSSSKSSSSSSS 1245
Query: 122 SSSSSSLESSNSSLDSNES 140
SSSSSS +SS+S S+ S
Sbjct: 1246 SSSSSSSKSSSSRSSSSSS 1264
Score = 38.1 bits (87), Expect = 0.051
Identities = 28/79 (35%), Positives = 39/79 (48%)
Query: 62 AKKTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVL 121
+K ++ + S S SSSS S+SS+SSS + SK + + SS
Sbjct: 1184 SKSSNSSKRSSSSSSSSSSSSRSSSSSSSSSSNSKSSSSSSKSSSSSSRSRSSSKSSSSS 1243
Query: 122 SSSSSSLESSNSSLDSNES 140
SSSSSS S +SS S+ S
Sbjct: 1244 SSSSSSSSSKSSSSRSSSS 1262
Score = 37.0 bits (84), Expect = 0.11
Identities = 31/77 (40%), Positives = 41/77 (52%), Gaps = 6/77 (7%)
Query: 62 AKKTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVL 121
+ +S RSS S SSSS+ SKSS+SSS S + SK + ++ SS
Sbjct: 1198 SSSSSSSRSSSSSSSSSSN-SKSSSSSSKSSSSSSRSRSSSKSSSSSSSSSSSSS----- 1251
Query: 122 SSSSSSLESSNSSLDSN 138
S SSSS SS+SS S+
Sbjct: 1252 SKSSSSRSSSSSSKSSS 1268
Score = 33.9 bits (76), Expect = 0.97
Identities = 31/117 (26%), Positives = 50/117 (42%), Gaps = 4/117 (3%)
Query: 18 KRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSRKRSSRKSGSS 77
K+A+ + S + ++S+R + ++ + + ++S SS S S
Sbjct: 1159 KQARNKDASSSSRSSKSSNSSKRSSSKSSNSSKRSSSSSSSSSSSSRSSSSSSSSSSNSK 1218
Query: 78 SSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSNSS 134
SS S S+SSS S K S + ++ SS SSSSSS SS+ S
Sbjct: 1219 SSSSSSKSSSSSSRSRSSSKSSSSSSSSSSSSSSKSSSSR----SSSSSSKSSSHHS 1271
Score = 32.7 bits (73), Expect = 2.2
Identities = 32/118 (27%), Positives = 45/118 (38%), Gaps = 4/118 (3%)
Query: 16 SQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSRKRSSRKSG 75
S KR+ + S + S+S ++ + +K +S SR S
Sbjct: 1178 SSKRSSSKSSNSSKRSSSSSSSSSSSSRSSSSSSSSSSNSKSSSSSSKSSSSSSRSRSSS 1237
Query: 76 SSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSNS 133
SSS S SS+SSS S SK ++ S L+ SSSS SS S
Sbjct: 1238 KSSSSSSSSSSSSSSKSSSSRSSSSSSK----SSSHHSHSHHSGHLNGSSSSSSSSRS 1291
Score = 32.0 bits (71), Expect = 3.7
Identities = 45/143 (31%), Positives = 58/143 (40%), Gaps = 28/143 (19%)
Query: 2 PKSR--KRGATPTT-FASQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVE 58
PK+R K+G T F ++ AK T S A S + P+ E
Sbjct: 1099 PKNRPSKKGNTVLAEFGTEPDAK-----TSSSSSSASSTATSSSSSSASSPNRKKPMDEE 1153
Query: 59 MALAKKTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPP 118
+ K++ K SSSS SKSS SS K S K SK + SS
Sbjct: 1154 ----ENDQVKQARNKDASSSSRSSKSSNSS---KRSSSKSSNSSKRS-------SSS--- 1196
Query: 119 VVLSSSSSSLESSNSSLDSNESD 141
SSSSSS S+SS S+ S+
Sbjct: 1197 ---SSSSSSSSRSSSSSSSSSSN 1216
Score = 30.8 bits (68), Expect = 8.2
Identities = 21/91 (23%), Positives = 39/91 (42%)
Query: 4 SRKRGATPTTFASQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAK 63
S KR ++ ++ +S++ + + + S + S+S + ++ +
Sbjct: 1178 SSKRSSSKSSNSSKRSSSSSSSSSSSSRSSSSSSSSSSNSKSSSSSSKSSSSSSRSRSSS 1237
Query: 64 KTSRKRSSRKSGSSSSHHSKSSTSSSPLKES 94
K+S SS S SSS S S+SSS S
Sbjct: 1238 KSSSSSSSSSSSSSSKSSSSRSSSSSSKSSS 1268
>NP14_RAT (P41777) Nucleolar phosphoprotein p130 (Nucleolar 130 kDa
protein) (140 kDa nucleolar phosphoprotein) (Nopp140)
(Nucleolar and coiled-body phosphoprotein 1)
Length = 704
Score = 41.6 bits (96), Expect = 0.005
Identities = 58/264 (21%), Positives = 107/264 (39%), Gaps = 57/264 (21%)
Query: 53 TPIPVEMALAKKTSRKRSSRKSGSSSSHHSKSSTSSS--------PLKESGPKRVGMSKM 104
T P + K K ++ K+GSSSS S SS+ S PLK++ PK+ ++K
Sbjct: 203 TKAPAKPGPPAKAQPKAANGKAGSSSSSSSSSSSDDSEEEKKAAAPLKKTAPKKQVVAKA 262
Query: 105 AV-----------------------------PRATWFSSKPPPVVLSSSSS--------- 126
V +A +SS PPP V S S
Sbjct: 263 PVKVTAAPTQKSSSSEDSSSEEEEEQKKPMKKKAGPYSSVPPPSVSLSKKSVGAQSPKKA 322
Query: 127 --SLESSNSSLDSNESDPVWDKLTFYFNSKPIAWQHPGCEPYYTRECRDNLSNFPPQDAQ 184
+ ++SS DS+E + +K + + P +P ++ ++ S+ D+
Sbjct: 323 AAQTQPADSSADSSEESDSSSEEEKKTPAKTVVSKTP-AKPAPVKKKAESSSDSSDSDSS 381
Query: 185 NMPSPAREENDPHVEVSGGEDQVSPIIQPDQVIGVMLPPKADVPASTFSEPSSRQLQTSP 244
+PA+ + +S + V+P +P V P + PA + +P SR+ +S
Sbjct: 382 EDEAPAKPVSATKSPLS--KPAVTP--KPPAAKAVATPKQ---PAGSGQKPQSRKADSSS 434
Query: 245 QQLQSSPQQLQQGARVALLSPHAK 268
+ +SS + ++ + ++ +P A+
Sbjct: 435 SEEESSSSE-EEATKKSVTTPKAR 457
Score = 33.9 bits (76), Expect = 0.97
Identities = 42/143 (29%), Positives = 59/143 (40%), Gaps = 26/143 (18%)
Query: 2 PKSRKRGATPTTFASQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMAL 61
P + K ATP K+ G+G+ P K+ S E + + T V
Sbjct: 406 PPAAKAVATP------KQPAGSGQ----KPQSRKADSSSSEEESSSSEEEATKKSVTTPK 455
Query: 62 AKKTSR-------KRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSS 114
A+ T++ K++ R G SSS S +SSS ++ P + K A A
Sbjct: 456 ARVTAKAAPSLPAKQAPRAGGDSSSD---SESSSSEEEKKTPPKPPAKKKAAGAAV---P 509
Query: 115 KPPPV---VLSSSSSSLESSNSS 134
KP PV SSSSS S +SS
Sbjct: 510 KPTPVKKAAAESSSSSSSSEDSS 532
Score = 33.9 bits (76), Expect = 0.97
Identities = 56/275 (20%), Positives = 99/275 (35%), Gaps = 38/275 (13%)
Query: 27 TVGDPSPAKSASQRVEMQQGGDQGGTTP--------IPVEMALAKKTSRKRSSRKSGSSS 78
+VG SP K+A+Q D + P + ++K ++ +K SS
Sbjct: 313 SVGAQSPKKAAAQTQPADSSADSSEESDSSSEEEKKTPAKTVVSKTPAKPAPVKKKAESS 372
Query: 79 SHHSKSSTS--SSPLKESGPKRVGMSKMAV---PRATWFSSKPPPVVLSSSSSSLESSNS 133
S S S +S +P K + +SK AV P A + P S ++S
Sbjct: 373 SDSSDSDSSEDEAPAKPVSATKSPLSKPAVTPKPPAAKAVATPKQPAGSGQKPQSRKADS 432
Query: 134 SLDSNESDPVWDKLTFYFNSKPIAWQHPGCEPYYTRECRDNLSNFPPQDAQNMPSPAREE 193
S ES ++ T K + T + R P A+ P +
Sbjct: 433 SSSEEESSSSEEEAT----KKSVT----------TPKARVTAKAAPSLPAKQAPRAGGDS 478
Query: 194 NDPHVEVSGGEDQVSPIIQP--DQVIGVMLPPKADV-PASTFSEPSSRQLQTSPQQLQSS 250
+ S E++ +P P + G +P V A+ S SS + S ++ +
Sbjct: 479 SSDSESSSSEEEKKTPPKPPAKKKAAGAAVPKPTPVKKAAAESSSSSSSSEDSSEEEKKK 538
Query: 251 PQQLQQGARVALLSPHAKGGSASSKMAASSHTSGE 285
P+ + +P + G A+ A+ + +G+
Sbjct: 539 PK--------SKATPKPQAGKANGVPASQNGKAGK 565
>WEE1_SCHPO (P07527) Mitosis inhibitor protein kinase wee1 (EC
2.7.1.-) (P107 protein kinase homolog)
Length = 877
Score = 41.2 bits (95), Expect = 0.006
Identities = 57/244 (23%), Positives = 94/244 (38%), Gaps = 27/244 (11%)
Query: 13 TFASQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSRKRSSR 72
+F S + GA PSP +SQ + +TP A+K +K +
Sbjct: 161 SFFSSSMSNGAMSPPSHSPSPFLQSSQHIPP--------STP-------AQKLRKKNNFD 205
Query: 73 KSGSSSSHHSK-SSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESS 131
S+SH S +S S SP S P + S A P + ++ P + SS SS ++
Sbjct: 206 SFRISNSHISPFASGSFSPFATSSPNFLSTSTPAPPNS---NNANPSTLFSSIPSSRHTT 262
Query: 132 NSSLDSNESDPVWDKLTFYFNSKPIAWQHPGCEPYYTREC-RDNLSNFPPQDAQNMPSPA 190
++ SN + F ++P+ + G T+ +N S +DA M
Sbjct: 263 SNHFPSNSAQ----SSLFSPTARPLTARKLGFASSQTKSAVSNNHSRNSSKDASFMMKSF 318
Query: 191 REENDPHVEVSGGEDQVSPIIQPDQVIGVMLPPKADVPASTFSEPSSRQLQTSPQQLQSS 250
N H + E S + + ++ + P +T P S +L T+P Q
Sbjct: 319 IPSNRSHPQTQQNE---SSLFSDNSMVNSSSNSFSLFPNATLPNPPSSELLTTPFQQIKP 375
Query: 251 PQQL 254
P Q+
Sbjct: 376 PSQV 379
>GIDA_XYLFA (Q9PBN4) Glucose inhibited division protein A
Length = 629
Score = 39.7 bits (91), Expect = 0.018
Identities = 46/202 (22%), Positives = 79/202 (38%), Gaps = 15/202 (7%)
Query: 294 DPLSIFQSFFDGTLDLESPPRQAETTETAQSGGVVDDSRIEEAFGKLKRLVFDAGFIDGI 353
+P +F S + L L A T + G++DD+R K + + + +
Sbjct: 425 EPYRMFTSRAEYRLQLREDNADARLTAIGRDLGLIDDARWAHFNAKQEAVARECERLSAF 484
Query: 354 RATP--QLGQEVKELLAFLLSQRLDPIQADALVELQTLLVNIMATLDKALAVEQLIETKK 411
ATP LG+EVKE L LS+ + I EL + + +L + Q+ E +
Sbjct: 485 WATPGNALGREVKETLGVTLSRETNIIDLMKRPELDYAALMRVPSLGPGVDDAQVAE--Q 542
Query: 412 AQVASNTNGLLPAKDEILKLTARKTLIAAEL-----------SSVDARLDELQREEVAAE 460
+++ G L + E + R A L + V +L +Q + V
Sbjct: 543 VEISVKYAGYLNRQSEEITRQQRHEATAIPLEFDYAAVRGLSTEVLQKLQHIQPQTVGQA 602
Query: 461 QRVKQSLDSRIAAARVQLEAFK 482
QR+ + I+ V LE +
Sbjct: 603 QRIPGMTPAAISLLLVHLERMR 624
>PUM1_MOUSE (Q80U78) Pumilio homolog 1
Length = 1189
Score = 39.3 bits (90), Expect = 0.023
Identities = 38/164 (23%), Positives = 63/164 (38%), Gaps = 8/164 (4%)
Query: 188 SPAREENDPHVEVSGGEDQVSPIIQPDQVIGVMLPPKADVPASTFSEPSSRQLQTSPQQL 247
S A DP+ + P + P Q GV P PAS F + ++ +
Sbjct: 433 SAAPPGTDPYTAGLAAAATLGPAVVPHQYYGVT--PWGVYPASLFQQQAAAAAAATNSAT 490
Query: 248 QSSPQQLQQGARVALLSPHAKGGSASSKMAASSHTSGERLSALLVEDPLSIFQSFFDG-T 306
Q S Q QQG + L +GG++ + + + G++ L+ ++ +F G
Sbjct: 491 QQSAPQAQQGQQQVL-----RGGASQRPLTPNQNQQGQQTDPLVAAAAVNSALAFGQGLA 545
Query: 307 LDLESPPRQAETTETAQSGGVVDDSRIEEAFGKLKRLVFDAGFI 350
+ P A Q+G +V ++ G RLV A I
Sbjct: 546 AGMPGYPVLAPAAYYDQTGALVVNAGARNGLGAPVRLVAPAPVI 589
>POB1_SCHPO (O74653) Pob1 protein (BOI protein homolog)
Length = 871
Score = 38.1 bits (87), Expect = 0.051
Identities = 53/214 (24%), Positives = 88/214 (40%), Gaps = 27/214 (12%)
Query: 84 SSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSNSSLDSNESDPV 143
SS +S + + P+ V P AT P++ SS + S NS L S ++P+
Sbjct: 406 SSRNSKSTQSAVPENVSRFDSNEPSAT------SPILKRSSPTDSISQNSGLPSRLTEPI 459
Query: 144 WDKLTFYFNSKPIAWQHPGCEPYYTRECRDNLSNFPPQDAQNMPSP----AREEND---- 195
T + PG PY++ + NL + PQ + N+P+ A E +
Sbjct: 460 SSPSTSSIDVDKEGTSFPGL-PYHSS--KGNL--YAPQPSSNVPTKFTGGASESSSVPPR 514
Query: 196 PHVEVSGGEDQVSPIIQPDQVIGVMLPPK-----ADVPASTFSE-PSSRQLQTSPQQLQS 249
P G+ S I + + + PPK + P+S S PSS Q S +
Sbjct: 515 PIPSAMKGKAPASAI--SIEALEELDPPKITTIDGESPSSISSRLPSSNLEQGSSSSVTK 572
Query: 250 SPQQLQQGARVALLSPHAKGGSASSKMAASSHTS 283
SP+ + + A +KG S + K A +++ +
Sbjct: 573 SPESMPDPSAKASSPVTSKGVSINEKSAVNNYAT 606
>YB95_SCHPO (O42970) Hypothetical serine-rich protein C1E8.05 in
chromosome II precursor
Length = 317
Score = 37.7 bits (86), Expect = 0.067
Identities = 29/79 (36%), Positives = 44/79 (54%), Gaps = 2/79 (2%)
Query: 62 AKKTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSK-MAVPRATWFSSK-PPPV 119
+ +S+ SS KS SSSS SKSS+SSS +S SK A P ++ SSK
Sbjct: 166 SSSSSKSSSSSKSSSSSSSSSKSSSSSSSSSKSSSSSSSSSKSSASPSSSKSSSKFSSSS 225
Query: 120 VLSSSSSSLESSNSSLDSN 138
++S++ + SS+ ++ SN
Sbjct: 226 FITSTTPASSSSSGAIVSN 244
Score = 36.6 bits (83), Expect = 0.15
Identities = 30/93 (32%), Positives = 45/93 (48%), Gaps = 2/93 (2%)
Query: 70 SSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLE 129
SS + SSSS S SS SSS K S + S + +++ SS SSSSSS
Sbjct: 149 SSSSTPSSSSSSSSSSPSSSSSKSSSSSKSSSSSSSSSKSSSSSSSSSKSSSSSSSSS-- 206
Query: 130 SSNSSLDSNESDPVWDKLTFYFNSKPIAWQHPG 162
S++S S++S + +F ++ P + G
Sbjct: 207 KSSASPSSSKSSSKFSSSSFITSTTPASSSSSG 239
Score = 36.2 bits (82), Expect = 0.19
Identities = 30/79 (37%), Positives = 39/79 (48%), Gaps = 4/79 (5%)
Query: 62 AKKTSRKRSSRKSGSSSSHHSK-SSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVV 120
+K +S SS KS SSSS SK SS+SSS K S S +++ +S P
Sbjct: 176 SKSSSSSSSSSKSSSSSSSSSKSSSSSSSSSKSSASPSSSKSSSKFSSSSFITSTTP--- 232
Query: 121 LSSSSSSLESSNSSLDSNE 139
SSSSS SN+ S +
Sbjct: 233 ASSSSSGAIVSNAKTASTD 251
Score = 34.3 bits (77), Expect = 0.74
Identities = 26/110 (23%), Positives = 45/110 (40%), Gaps = 1/110 (0%)
Query: 31 PSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSRKRSSRKSGSSSSHHSKSSTSSSP 90
PS + S+S ++ + + K+S SS S SSSS S S +S+SP
Sbjct: 154 PSSSSSSSSSSPSSSSSKSSSSSKSSSSSSSSSKSSSSSSS-SSKSSSSSSSSSKSSASP 212
Query: 91 LKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSNSSLDSNES 140
+ S SS +V ++ ++S + S+S+ + S
Sbjct: 213 SSSKSSSKFSSSSFITSTTPASSSSSGAIVSNAKTASTDDSSSASSATSS 262
Score = 31.6 bits (70), Expect = 4.8
Identities = 34/128 (26%), Positives = 50/128 (38%), Gaps = 7/128 (5%)
Query: 15 ASQKRAKGAGELTVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSRKRSSRKS 74
+S + + PS + S S + + +K +S SS KS
Sbjct: 149 SSSSTPSSSSSSSSSSPSSSSSKSSSSSKSSSSSSSSSKSSSSSSSSSKSSSSSSSSSKS 208
Query: 75 GSS--SSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSN 132
+S SS S +SSS + + P S V A S+ SSS+SS SS
Sbjct: 209 SASPSSSKSSSKFSSSSFITSTTPASSSSSGAIVSNAKTASTDD-----SSSASSATSSV 263
Query: 133 SSLDSNES 140
SS+ S+ S
Sbjct: 264 SSVVSSAS 271
>PCLO_CHICK (Q9PU36) Piccolo protein (Aczonin) (Fragment)
Length = 5120
Score = 37.7 bits (86), Expect = 0.067
Identities = 55/249 (22%), Positives = 95/249 (38%), Gaps = 43/249 (17%)
Query: 31 PSPAKSASQRVEMQQGGDQGGTT-PIPVEMALAKK-TSRKRSSRKSGS----SSSHHSKS 84
P+PA +A ++Q G + + P P + A +KK +K+ GS S H++
Sbjct: 447 PTPASTAKPSPQLQPGQKKDASPKPDPSQQADSKKPVPQKKQPSMPGSPPVKSKQTHAEP 506
Query: 85 STSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSNSSLDSNESDPVW 144
S + + +S PK + S P + + +S+ L +S
Sbjct: 507 SDTGQQI-DSTPKSDQVKPTQAEEKQNQPSIQKPTMDTVPTSAAPGVKQDLADPQSPSTQ 565
Query: 145 DKLT---FYFNSKPIAWQHP-GCEPYYTRECRDNLSNFPPQDAQNMPSPAREENDPHVEV 200
K+T +KP A HP G +P D++ +P +R+++DP +
Sbjct: 566 QKVTDSPMPETTKPPADTHPAGDKP----------------DSKPLPQVSRQKSDPKLAS 609
Query: 201 SGGE---------DQVSPIIQPDQVIGVMLPPKADVP-------ASTFSEPSSRQLQTSP 244
G + +P+ + + PK D A T P+S Q P
Sbjct: 610 QSGAKSDAKTQKPSEPAPVKDDPKKLQTKPAPKPDTKPAPKGPQAGTGPRPTSAQPAPQP 669
Query: 245 QQLQSSPQQ 253
QQ Q +P+Q
Sbjct: 670 QQPQKTPEQ 678
Score = 35.4 bits (80), Expect = 0.33
Identities = 82/387 (21%), Positives = 146/387 (37%), Gaps = 38/387 (9%)
Query: 71 SRKSGSSSSHHSKSSTSSSPL--------KESGP--KRVGMSKMAVPRATWFSSKP--PP 118
SR GSSS H S + +SP +S P K++G +A S P P
Sbjct: 1628 SRGEGSSSLHASSFTPGTSPTSVSSLDEDSDSSPSHKKLGGESKQQRKARHRSHGPLLPT 1687
Query: 119 VVLSSSSSSLESSNSSLDSNESDPVWDKLTFYFNSKPIAWQHPGCEPYYTRE-CRDNLSN 177
+ SS L L E ++ +SK RE + SN
Sbjct: 1688 IEDSSEEEELREEEELLKEQEKQRELEQQQRKSSSKKSKKDKDELRAQRRRERPKTPPSN 1747
Query: 178 FPPQDAQNMPSPAREENDPHVEVSGGEDQVSPIIQ---------PDQVIGVMLPPKADVP 228
P + + R+ + + SP I+ P+++I V K
Sbjct: 1748 LSPIEDASPTEELRQAAEMEELHRSSCSEYSPSIESDPEGFEISPEKIIEVQKVYKLPTA 1807
Query: 229 ASTFSEPSSRQLQTSPQQLQSSPQQLQQGARVALLSPHAKGGSASSKMAASSHTSGERLS 288
S +S P+ +L + ++ +S + L+ V H S S ++A+ E+ S
Sbjct: 1808 VSLYS-PTDEKLIGALKE-ESGQKTLKSAEEVYEEMIHKTHKSKSFQIASEKDEVFEKES 1865
Query: 289 ---ALLVEDPL--SIFQSFFDGTLDLESPPRQAETTETAQSGGVVDDSRIEEAF----GK 339
+L+ED + S+ + ++GT+D RQ E+ E Q G R E K
Sbjct: 1866 LYGGMLIEDYIYESLIEDTYNGTVDTNLAMRQDESNEYIQQRGKEKKIRASEQIYDEPQK 1925
Query: 340 LKRLVFDAGFIDGIRATPQLGQEVKELLAFLLSQRLDPIQADALVELQTLLVNIMATLDK 399
+ L D ++ + + + QE ++++ + + + D+ V T LD
Sbjct: 1926 ITDLQEDYYSVEPLCSI--VPQEDIVSSSYIIPESHEIVVLDSTV---TSTTEEKQLLDA 1980
Query: 400 ALAVEQLIETKKAQVASNTNGLLPAKD 426
A E+L++ ++ Q+ ++ P D
Sbjct: 1981 EAAYEELMKKQRMQLTPGSSPTQPTSD 2007
>DESP_HUMAN (P15924) Desmoplakin (DP) (250/210 kDa paraneoplastic
pemphigus antigen)
Length = 2871
Score = 37.7 bits (86), Expect = 0.067
Identities = 31/131 (23%), Positives = 61/131 (45%), Gaps = 4/131 (3%)
Query: 358 QLGQEVKELLAFLLSQRLDPIQADALVELQTLLVNIMATLDKALAVEQLIETKKAQVASN 417
Q Q V+E + L+ ++ L L+ L +TL+ V+Q +E +K Q+ ++
Sbjct: 1804 QCTQVVQERESLLVKIKVLEQDKARLQRLEDELNRAKSTLEAETRVKQRLECEKQQIQND 1863
Query: 418 TNGLLPA----KDEILKLTARKTLIAAELSSVDARLDELQREEVAAEQRVKQSLDSRIAA 473
N ++ I K+ + + E +S+ + ++ LQ E E+R ++ L+
Sbjct: 1864 LNQWKTQYSRKEEAIRKIESEREKSEREKNSLRSEIERLQAEIKRIEERCRRKLEDSTRE 1923
Query: 474 ARVQLEAFKSR 484
+ QLE +SR
Sbjct: 1924 TQSQLETERSR 1934
>ATRX_DROME (Q9GQN5) Transcriptional regulator ATRX homolog
(X-linked nuclear protein) (dXNP) (d-xnp)
Length = 1311
Score = 37.7 bits (86), Expect = 0.067
Identities = 53/242 (21%), Positives = 104/242 (42%), Gaps = 33/242 (13%)
Query: 53 TPIPVEMALAKKTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRVGMSKMAVPRATWF 112
TP+ + + + SR+ S+ +S S+S S SS P + R + A +AT
Sbjct: 15 TPLTTDDSNSSSVSRRESATESKSASE-----SESSPPRSNTKQSRTHKNVKASGKAT-- 67
Query: 113 SSKPPPVVLSSSSSSLESSNSSLDSNESDPV-------WDKLTFYFNSKPIAWQHPGCEP 165
V SSS S +NSS + E +PV +KL + P + P
Sbjct: 68 -------VSSSSDSDQAVANSSANDEEKEPVCKIRIVPLEKLL----ASPKTKERPSRGS 116
Query: 166 YYTRECRDNLSNFPPQDAQNMPSPARE---ENDPHVEVSGGEDQVSPIIQPDQVIGVMLP 222
++ S+ P + PAR+ +N +E+S E+ + + ++ + V +P
Sbjct: 117 QQKNVTINDSSDEEPLKGSKLVLPARKSRNKNASIIELSDSEE----VDEEEESLLVAIP 172
Query: 223 PKADVPASTFSEPSSRQLQTSPQQLQSSPQQLQQGARVALLSPH-AKGGSASSKMAASSH 281
+ + + SS+ + S ++ Q + ++ + A+ S + + GS SS+ ++ +
Sbjct: 173 LPKEAQQTKPEKNSSKASKESIEKRQKAQKEATTSSARAIRSVNGTRRGSLSSERSSRAS 232
Query: 282 TS 283
+S
Sbjct: 233 SS 234
Score = 31.2 bits (69), Expect = 6.3
Identities = 27/115 (23%), Positives = 53/115 (45%), Gaps = 14/115 (12%)
Query: 27 TVGDPSPAKSASQRVEMQQGGDQGGTTPIPVEMALAKKTSRKRSSRKSGSSSSHHSKSST 86
T + + +K++ + +E +Q + TT + + R + + GS SS S ++
Sbjct: 180 TKPEKNSSKASKESIEKRQKAQKEATTS-------SARAIRSVNGTRRGSLSSERSSRAS 232
Query: 87 SSSPLKESGPKR--VGMSKMAVPRATWFSSKPPPVVLSSSSSSLESSNSSLDSNE 139
SS PKR V + ++++P+ +KP SS S E++ +S S +
Sbjct: 233 SSRAESPPRPKRCVVRLKRVSLPK-----TKPAQKPKKMSSDSEEAATTSKKSRQ 282
Score = 30.8 bits (68), Expect = 8.2
Identities = 24/100 (24%), Positives = 39/100 (39%), Gaps = 7/100 (7%)
Query: 1 MPKSRKRGATPTTFASQKRAKGAGELTVGDPSPAKSASQRVEMQQGGD-QGGTTPIPVEM 59
M + AT + + Q+R+K E +PA + E + GD + E+
Sbjct: 266 MSSDSEEAATTSKKSRQRRSKSESEADSDYEAPAAEEEEEEERKSSGDEEEAANSSDSEV 325
Query: 60 ALAKKTSRKRSSRKSGSSSSHHSKSSTSSSPLKESGPKRV 99
+K RK+S GSS + K+ G KR+
Sbjct: 326 MPQRKRRRKKSESDKGSSDFEPEEKQ------KKKGRKRI 359
>AF4_HUMAN (P51825) AF-4 protein (Proto-oncogene AF4) (FEL protein)
Length = 1210
Score = 37.7 bits (86), Expect = 0.067
Identities = 61/242 (25%), Positives = 91/242 (37%), Gaps = 34/242 (14%)
Query: 65 TSRKRSSRKSGSSSSHHS--KSSTSSSPLKESGPKRVGMSKMAVPRATWFSSKPPPVVLS 122
+S+ S+ + G+SS + S S E P++ S A P A S P PV +
Sbjct: 408 SSKTHSNSQQGTSSMLEDDLQLSDSEDSDSEQTPEKPPSSS-APPSAP--QSLPEPVASA 464
Query: 123 SSSSSLESSNSSLDSNESDPVWDKLTFYFNSKPIAWQHPGCEPYYTRECRDNLSNF---- 178
SSS+ S S DS+ + ++P+ P EP T + + L N+
Sbjct: 465 HSSSAESESTSDSDSSSDSESESSSSDSEENEPLETPAPEPEPPTTNKWQ--LDNWLTKV 522
Query: 179 -----PPQDAQNMPSPAREENDPHVEVSGGEDQVSPIIQPDQVIGVMLPPKADVPA---- 229
PP+ ++ P R H E G D + + PPK+ A
Sbjct: 523 SQPAAPPEGPRSTEPPRR-----HPESKGSSDSAT---SQEHSESKDPPPKSSSKAPRAP 574
Query: 230 STFSEPSSRQLQTSPQQLQSSPQQLQQGARVALLSPHAKGGSASSKMAASSHTSGERLSA 289
P R Q SP Q Q PQ+ G + K AS++ + + GER
Sbjct: 575 PEAPHPGKRSCQKSPAQ-QEPPQRQTVGTK-----QPKKPVKASARAGSRTSLQGEREPG 628
Query: 290 LL 291
LL
Sbjct: 629 LL 630
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.309 0.125 0.340
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 53,661,254
Number of Sequences: 164201
Number of extensions: 2279746
Number of successful extensions: 11237
Number of sequences better than 10.0: 299
Number of HSP's better than 10.0 without gapping: 62
Number of HSP's successfully gapped in prelim test: 246
Number of HSP's that attempted gapping in prelim test: 9480
Number of HSP's gapped (non-prelim): 958
length of query: 485
length of database: 59,974,054
effective HSP length: 114
effective length of query: 371
effective length of database: 41,255,140
effective search space: 15305656940
effective search space used: 15305656940
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0143b.7


