

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
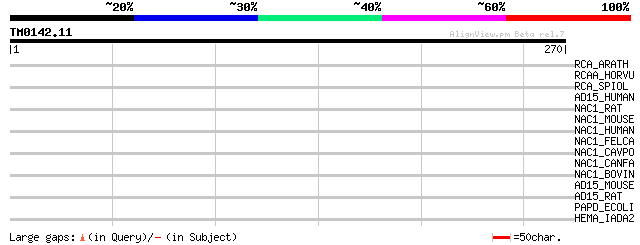
Query= TM0142.11
(270 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
RCA_ARATH (P10896) Ribulose bisphosphate carboxylase/oxygenase a... 40 0.005
RCAA_HORVU (Q40073) Ribulose bisphosphate carboxylase/oxygenase ... 40 0.006
RCA_SPIOL (P10871) Ribulose bisphosphate carboxylase/oxygenase a... 38 0.023
AD15_HUMAN (Q13444) ADAM 15 precursor (EC 3.4.24.-) (A disintegr... 31 2.8
NAC1_RAT (Q01728) Sodium/calcium exchanger 1 precursor (Na(+)/Ca... 31 3.7
NAC1_MOUSE (P70414) Sodium/calcium exchanger 1 precursor (Na(+)/... 31 3.7
NAC1_HUMAN (P32418) Sodium/calcium exchanger 1 precursor (Na(+)/... 31 3.7
NAC1_FELCA (P48767) Sodium/calcium exchanger 1 precursor (Na(+)/... 31 3.7
NAC1_CAVPO (P48766) Sodium/calcium exchanger 1 precursor (Na(+)/... 31 3.7
NAC1_CANFA (P23685) Sodium/calcium exchanger 1 precursor (Na(+)/... 31 3.7
NAC1_BOVIN (P48765) Sodium/calcium exchanger 1 precursor (Na(+)/... 31 3.7
AD15_MOUSE (O88839) ADAM 15 precursor (EC 3.4.24.-) (A disintegr... 31 3.7
AD15_RAT (Q9QYV0) ADAM 15 precursor (EC 3.4.24.-) (A disintegrin... 30 6.2
PAPD_ECOLI (P15319) Chaperone protein papD precursor 30 8.1
HEMA_IADA2 (P03446) Hemagglutinin precursor [Contains: Hemagglut... 30 8.1
>RCA_ARATH (P10896) Ribulose bisphosphate carboxylase/oxygenase
activase, chloroplast precursor (RuBisCO activase) (RA)
Length = 474
Score = 40.4 bits (93), Expect = 0.005
Identities = 27/91 (29%), Positives = 42/91 (45%), Gaps = 9/91 (9%)
Query: 176 FQVPKVLQNKPIEFFNEGLAEELG------KEGACEQAIFPDSNKVITKCAMLGNLTVEG 229
F+ P++ K +E+ N + E+ E QA D+N G +G
Sbjct: 383 FEQPEMTYEKLMEYGNMLVMEQENVKRVQLAETYLSQAALGDAN---ADAIGRGTFYGKG 439
Query: 230 GDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ +L + EGCTDP + ++P A DDGTC
Sbjct: 440 AQQVNLPVPEGCTDPVAENFDPTARSDDGTC 470
>RCAA_HORVU (Q40073) Ribulose bisphosphate carboxylase/oxygenase
activase A, chloroplast precursor (RuBisCO activase A)
(RA A)
Length = 464
Score = 40.0 bits (92), Expect = 0.006
Identities = 29/98 (29%), Positives = 48/98 (48%), Gaps = 11/98 (11%)
Query: 169 DGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNKVITKCAML 222
DGP + F+ PK+ K +E+ + + E+ + QA D+N+ K
Sbjct: 368 DGP--VTFEQPKMTVEKLLEYGHMLVQEQDNVKRVQLADTYMSQAALGDANQDAMKT--- 422
Query: 223 GNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
G+ +G + L + EGCTD ++ Y+P A DDG+C
Sbjct: 423 GSFYGKGAQQGTLPVPEGCTDQNAKNYDPTARSDDGSC 460
>RCA_SPIOL (P10871) Ribulose bisphosphate carboxylase/oxygenase
activase, chloroplast precursor (RuBisCO activase) (RA)
Length = 472
Score = 38.1 bits (87), Expect = 0.023
Identities = 29/103 (28%), Positives = 47/103 (45%), Gaps = 13/103 (12%)
Query: 169 DGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNK-VITKCAM 221
DGP F+ P++ K +E+ N + E+ + A D+NK I +
Sbjct: 376 DGPP--VFEQPEMTLQKLMEYGNMLVQEQENVKRVQLADQYMSSAALGDANKDAIDRGTF 433
Query: 222 LGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTCPLDL 264
G + + L + +GCTDP + Y+P A DDG+C +L
Sbjct: 434 FG----KAAQQVSLPVAQGCTDPEAKNYDPTARSDDGSCTYNL 472
>AD15_HUMAN (Q13444) ADAM 15 precursor (EC 3.4.24.-) (A disintegrin
and metalloproteinase domain 15)
(Metalloproteinase-like, disintegrin-like, and
cysteine-rich protein 15) (MDC-15) (Metalloprotease RGD
disintegrin protein) (Metargidin)
Length = 814
Score = 31.2 bits (69), Expect = 2.8
Identities = 11/25 (44%), Positives = 16/25 (64%)
Query: 220 AMLGNLTVEGGDRCDLNLVEGCTDP 244
A GN+ VE G++CD ++ C DP
Sbjct: 422 AFCGNMFVEPGEQCDCGFLDDCVDP 446
>NAC1_RAT (Q01728) Sodium/calcium exchanger 1 precursor
(Na(+)/Ca(2+)-exchange protein 1)
Length = 971
Score = 30.8 bits (68), Expect = 3.7
Identities = 26/92 (28%), Positives = 39/92 (42%), Gaps = 19/92 (20%)
Query: 17 PITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQE--IE------------- 61
PI HT L V + Y K K+ + N A+ V T +E IE
Sbjct: 702 PILGEHTKLEVIIEESYEFKSTVDKLIKKTNLALVVGTNSWREQFIEAITVSAGEDDDDD 761
Query: 62 ---EYKIPSWANFELGKASVYWKTMNG-LPPT 89
E K+PS ++ + +V+WK + +PPT
Sbjct: 762 ECGEEKLPSCFDYVMHFLTVFWKVLFAFVPPT 793
>NAC1_MOUSE (P70414) Sodium/calcium exchanger 1 precursor
(Na(+)/Ca(2+)-exchange protein 1)
Length = 970
Score = 30.8 bits (68), Expect = 3.7
Identities = 26/92 (28%), Positives = 39/92 (42%), Gaps = 19/92 (20%)
Query: 17 PITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQE--IE------------- 61
PI HT L V + Y K K+ + N A+ V T +E IE
Sbjct: 701 PILGEHTKLEVIIQESYEFKSTVDKLIKKTNLALVVGTNSWREQFIEAITVSAGEDDDDD 760
Query: 62 ---EYKIPSWANFELGKASVYWKTMNG-LPPT 89
E K+PS ++ + +V+WK + +PPT
Sbjct: 761 ECGEEKLPSCFDYVMHFLTVFWKVLFAFVPPT 792
>NAC1_HUMAN (P32418) Sodium/calcium exchanger 1 precursor
(Na(+)/Ca(2+)-exchange protein 1)
Length = 973
Score = 30.8 bits (68), Expect = 3.7
Identities = 26/92 (28%), Positives = 39/92 (42%), Gaps = 19/92 (20%)
Query: 17 PITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQE--IE------------- 61
PI HT L V + Y K K+ + N A+ V T +E IE
Sbjct: 704 PILGEHTKLEVIIEESYEFKSTVDKLIKKTNLALVVGTNSWREQFIEAITVSAGEDDDDD 763
Query: 62 ---EYKIPSWANFELGKASVYWKTMNG-LPPT 89
E K+PS ++ + +V+WK + +PPT
Sbjct: 764 ECGEEKLPSCFDYVMHFLTVFWKVLFAFVPPT 795
>NAC1_FELCA (P48767) Sodium/calcium exchanger 1 precursor
(Na(+)/Ca(2+)-exchange protein 1)
Length = 970
Score = 30.8 bits (68), Expect = 3.7
Identities = 26/92 (28%), Positives = 39/92 (42%), Gaps = 19/92 (20%)
Query: 17 PITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQE--IE------------- 61
PI HT L V + Y K K+ + N A+ V T +E IE
Sbjct: 701 PILGEHTKLEVIIEESYEFKSTVDKLIKKTNLALVVGTNSWREQFIEAITVSAGEDDDDD 760
Query: 62 ---EYKIPSWANFELGKASVYWKTMNG-LPPT 89
E K+PS ++ + +V+WK + +PPT
Sbjct: 761 ECGEEKLPSCFDYVMHFLTVFWKVLFAFVPPT 792
>NAC1_CAVPO (P48766) Sodium/calcium exchanger 1 precursor
(Na(+)/Ca(2+)-exchange protein 1)
Length = 970
Score = 30.8 bits (68), Expect = 3.7
Identities = 26/92 (28%), Positives = 39/92 (42%), Gaps = 19/92 (20%)
Query: 17 PITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQE--IE------------- 61
PI HT L V + Y K K+ + N A+ V T +E IE
Sbjct: 701 PILGEHTKLEVIIEESYEFKSTVDKLIKKTNLALVVGTNSWREQFIEAITVSAGEDDDDD 760
Query: 62 ---EYKIPSWANFELGKASVYWKTMNG-LPPT 89
E K+PS ++ + +V+WK + +PPT
Sbjct: 761 ECGEEKLPSCFDYVMHFLTVFWKVLFAFVPPT 792
>NAC1_CANFA (P23685) Sodium/calcium exchanger 1 precursor
(Na(+)/Ca(2+)-exchange protein 1)
Length = 970
Score = 30.8 bits (68), Expect = 3.7
Identities = 26/92 (28%), Positives = 39/92 (42%), Gaps = 19/92 (20%)
Query: 17 PITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQE--IE------------- 61
PI HT L V + Y K K+ + N A+ V T +E IE
Sbjct: 701 PILGEHTKLEVIIEESYEFKSTVDKLIKKTNLALVVGTNSWREQFIEAITVSAGEDDDDD 760
Query: 62 ---EYKIPSWANFELGKASVYWKTMNG-LPPT 89
E K+PS ++ + +V+WK + +PPT
Sbjct: 761 ECGEEKLPSCFDYVMHFLTVFWKVLFAFVPPT 792
>NAC1_BOVIN (P48765) Sodium/calcium exchanger 1 precursor
(Na(+)/Ca(2+)-exchange protein 1)
Length = 970
Score = 30.8 bits (68), Expect = 3.7
Identities = 26/92 (28%), Positives = 39/92 (42%), Gaps = 19/92 (20%)
Query: 17 PITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQE--IE------------- 61
PI HT L V + Y K K+ + N A+ V T +E IE
Sbjct: 701 PILGEHTRLEVIIEESYEFKSTVDKLIKKTNLALVVGTNSWREQFIEAITVSAGEDDDDD 760
Query: 62 ---EYKIPSWANFELGKASVYWKTMNG-LPPT 89
E K+PS ++ + +V+WK + +PPT
Sbjct: 761 ECGEEKLPSCFDYVMHFLTVFWKVLFAFVPPT 792
>AD15_MOUSE (O88839) ADAM 15 precursor (EC 3.4.24.-) (A disintegrin
and metalloproteinase domain 15)
(Metalloproteinase-like, disintegrin-like, and
cysteine-rich protein 15) (MDC-15) (Metalloprotease RGD
disintegrin protein) (Metargidin) (AD56)
Length = 864
Score = 30.8 bits (68), Expect = 3.7
Identities = 13/43 (30%), Positives = 23/43 (53%), Gaps = 3/43 (6%)
Query: 202 GACEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDP 244
G+C +F + ++ GN+ V+ G++CD + CTDP
Sbjct: 408 GSC---LFERQPSLAPMSSLCGNMFVDPGEQCDCGFPDECTDP 447
>AD15_RAT (Q9QYV0) ADAM 15 precursor (EC 3.4.24.-) (A disintegrin
and metalloproteinase domain 15)
(Metalloproteinase-like, disintegrin-like, and
cysteine-rich protein 15) (MDC-15) (Metalloprotease RGD
disintegrin protein) (Metargidin) (CRII-7)
Length = 816
Score = 30.0 bits (66), Expect = 6.2
Identities = 10/25 (40%), Positives = 17/25 (68%)
Query: 220 AMLGNLTVEGGDRCDLNLVEGCTDP 244
++ GN+ V+ G++CD + CTDP
Sbjct: 424 SLCGNMFVDPGEQCDCGFPDECTDP 448
>PAPD_ECOLI (P15319) Chaperone protein papD precursor
Length = 239
Score = 29.6 bits (65), Expect = 8.1
Identities = 19/76 (25%), Positives = 32/76 (42%), Gaps = 18/76 (23%)
Query: 52 VATTPAQEIEE--------YKIPSWANFELGKASVYWKTMNGLPPTSGE----------K 93
+AT P Q +E P + + S+++ + +PP S + K
Sbjct: 72 IATPPVQRLEPGAKSMVRLSTTPDISKLPQDRESLFYFNLREIPPRSEKANVLQIALQTK 131
Query: 94 LKLFYNPTSTQLAPNE 109
+KLFY P + + PNE
Sbjct: 132 IKLFYRPAAIKTRPNE 147
>HEMA_IADA2 (P03446) Hemagglutinin precursor [Contains:
Hemagglutinin HA1 chain; Hemagglutinin HA2 chain]
Length = 564
Score = 29.6 bits (65), Expect = 8.1
Identities = 16/55 (29%), Positives = 25/55 (45%), Gaps = 8/55 (14%)
Query: 87 PPTSGEKLKLFYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAMLRKDRGKAD 141
PPTS E++KL+ NP + E +F PR ++R +G+ D
Sbjct: 197 PPTSDEQVKLYKNPDTLSSVTTVEINRSFKPNIG--------PRPLVRGQQGRMD 243
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,995,084
Number of Sequences: 164201
Number of extensions: 1533509
Number of successful extensions: 2555
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 2542
Number of HSP's gapped (non-prelim): 15
length of query: 270
length of database: 59,974,054
effective HSP length: 108
effective length of query: 162
effective length of database: 42,240,346
effective search space: 6842936052
effective search space used: 6842936052
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0142.11


