

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
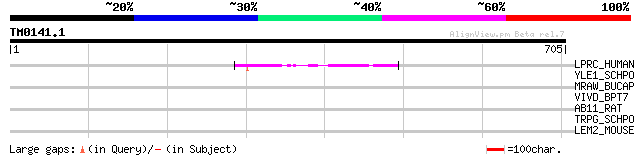
Query= TM0141.1
(705 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
LPRC_HUMAN (P42704) 130 kDa leucine-rich protein (LRP 130) (GP13... 45 5e-04
YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I 42 0.006
MRAW_BUCAP (O85295) S-adenosyl-methyltransferase mraW (EC 2.1.1.-) 35 0.89
VIVD_BPT7 (P03726) Internal virion protein D 33 2.6
AB11_RAT (O70127) Bile salt export pump (ATP-binding cassette, s... 32 5.7
TRPG_SCHPO (Q92370) Anthranilate synthase component II (EC 4.1.3... 32 7.5
LEM2_MOUSE (Q00690) E-selectin precursor (Endothelial leukocyte ... 31 9.8
>LPRC_HUMAN (P42704) 130 kDa leucine-rich protein (LRP 130) (GP130)
(Leucine-rich PPR-motif containing protein)
Length = 1273
Score = 45.4 bits (106), Expect = 5e-04
Identities = 51/215 (23%), Positives = 90/215 (41%), Gaps = 40/215 (18%)
Query: 286 ARKVFDGLQNRNVVV----WTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLG 341
A +++D LQ V + +L+ YLQN Y D L M++ + PN T++ L+
Sbjct: 25 AHRIWDTLQKLGAVYDVSHYNALLKVYLQNEYKFSPTDFLAKMEEANIQPNRVTYQRLIA 84
Query: 342 ACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTT 401
+ + GD+ A + LGF K+ D+ + VFS
Sbjct: 85 SYCNV-----GDIEGAS-KILGF--------------MKTKDLPVTEAVFS--------- 115
Query: 402 WNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEM 461
+++ G++ G + A + M+ AG P T++ +L+A A +D L
Sbjct: 116 --ALVTGHARAGDMENAENILTVMRDAGIEPGPDTYLALLNAYAEKGDIDHVKQTL---E 170
Query: 462 KNHKIEPGLEHYTCMVVL--YCRAGLLEKAENFMK 494
K K E L + ++ + +AG L ++ F K
Sbjct: 171 KVEKFELHLMDRDLLQIIFSFSKAGYLSMSQKFWK 205
>YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I
Length = 1261
Score = 42.0 bits (97), Expect = 0.006
Identities = 35/137 (25%), Positives = 66/137 (47%), Gaps = 11/137 (8%)
Query: 350 KHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDID---SSYNVFSDTMCRDVTT----W 402
K +L ++ LG+ LIN ++ GD D ++ N+F +T +V +
Sbjct: 870 KSSNLFCDNMKMLGYIPRASTFAHLINNSTRRGDTDDATTALNIFEETKRHNVKPSVFLY 929
Query: 403 NSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGL-DYLYNEM 461
N+++ E ++FQ+MK +G P +VT+ V++A + D+ L + L+ EM
Sbjct: 930 NAVLSKLGRARRTTECWKLFQEMKESGLLPTSVTYGTVINAACRIG--DESLAEKLFAEM 987
Query: 462 KNH-KIEPGLEHYTCMV 477
+N +P + Y M+
Sbjct: 988 ENQPNYQPRVAPYNTMI 1004
>MRAW_BUCAP (O85295) S-adenosyl-methyltransferase mraW (EC 2.1.1.-)
Length = 312
Score = 34.7 bits (78), Expect = 0.89
Identities = 24/85 (28%), Positives = 44/85 (51%), Gaps = 12/85 (14%)
Query: 594 ESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSP 653
E EI K ++ L ++KP G + IS H +ED + ++ +S+K + YGL P
Sbjct: 215 ELEEIKKGLENSLKILKPGGRLSIIS--FHSLEDRIVKNFMMKYSKKATVPYGL-----P 267
Query: 654 APIRIIKNLRMCDDCHTAVKLISKV 678
++ LR+C +K+I+++
Sbjct: 268 ITENKLETLRIC-----KLKIINRI 287
>VIVD_BPT7 (P03726) Internal virion protein D
Length = 1318
Score = 33.1 bits (74), Expect = 2.6
Identities = 21/61 (34%), Positives = 35/61 (56%), Gaps = 7/61 (11%)
Query: 351 HGDLL---HARLEKLGFKNYVI--VNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSM 405
HG+ L ++R GFKN ++N++ M ++G +D+ ++VF DT+ T WNS
Sbjct: 214 HGETLDEYNSRSTFFGFKNAAEAELSNSVAGMAFRAGRLDNGFDVFKDTI--TPTRWNSH 271
Query: 406 I 406
I
Sbjct: 272 I 272
>AB11_RAT (O70127) Bile salt export pump (ATP-binding cassette,
sub-family B, member 11) (Sister of P-glycoprotein)
Length = 1321
Score = 32.0 bits (71), Expect = 5.7
Identities = 20/56 (35%), Positives = 25/56 (43%), Gaps = 7/56 (12%)
Query: 317 TLDLLTCMDQEDTLPNECTFKVLLGACA-------GIALLKHGDLLHARLEKLGFK 365
T LL Q + + C F V+LG + G K G+LL RL K GFK
Sbjct: 782 TFSLLDKEQQRSEIHSMCLFFVILGCVSIFTQFLQGYTFAKSGELLTKRLRKFGFK 837
>TRPG_SCHPO (Q92370) Anthranilate synthase component II (EC
4.1.3.27) [Includes: Glutamine amidotransferase;
Indole-3-glycerol phosphate synthase (EC 4.1.1.48)
(IGPS); N-(5'-phosphoribosyl)anthranilate isomerase (EC
5.3.1.24) (PRAI)]
Length = 759
Score = 31.6 bits (70), Expect = 7.5
Identities = 22/81 (27%), Positives = 36/81 (44%), Gaps = 8/81 (9%)
Query: 333 ECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFS 392
+C F+ + G + HG + + LGF + N ++ +S +G I S +
Sbjct: 112 QCIFETMGGKVDSAGEIIHGKVSKINHDGLGFYQGIPQNISVTRYHSLAGKISSLPD--- 168
Query: 393 DTMCRDVTTW--NSMICGYSH 411
C DVT+W N +I G H
Sbjct: 169 ---CLDVTSWTENGVIMGARH 186
>LEM2_MOUSE (Q00690) E-selectin precursor (Endothelial leukocyte
adhesion molecule 1) (ELAM-1) (Leukocyte-endothelial
cell adhesion molecule 2) (LECAM2)
Length = 612
Score = 31.2 bits (69), Expect = 9.8
Identities = 24/83 (28%), Positives = 37/83 (43%), Gaps = 8/83 (9%)
Query: 566 NIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDV 625
N+K P W+ IR VN+V++ G P + E Q A +P N V +
Sbjct: 62 NLKHSPSYYWIGIRKVNNVWIWVGTGKPLTEE-----AQNWAPGEPNNKQRNEDCVEIYI 116
Query: 626 EDEQKEGYLN---CHSEKLAIAY 645
+ + G N C+ +KLA+ Y
Sbjct: 117 QRTKDSGMWNDERCNKKKLALCY 139
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.137 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 81,654,661
Number of Sequences: 164201
Number of extensions: 3382433
Number of successful extensions: 7037
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 7027
Number of HSP's gapped (non-prelim): 16
length of query: 705
length of database: 59,974,054
effective HSP length: 117
effective length of query: 588
effective length of database: 40,762,537
effective search space: 23968371756
effective search space used: 23968371756
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0141.1


