

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0137.5
(214 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
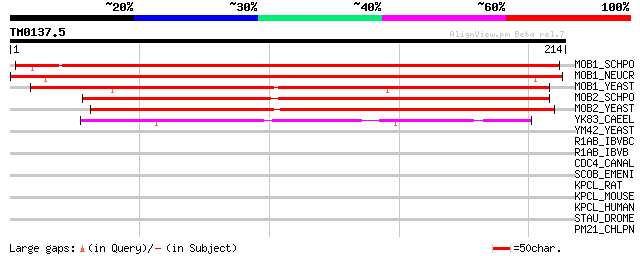
Sequences producing significant alignments: (bits) Value
MOB1_SCHPO (O94360) Maintenance of ploidy protein mob1 251 7e-67
MOB1_NEUCR (Q9P601) Probable maintenance of ploidy protein mob1 221 1e-57
MOB1_YEAST (P40484) Maintenance of ploidy protein MOB1 (MPS1 bin... 195 6e-50
MOB2_SCHPO (O74558) Maintenance of ploidy protein mob2 173 3e-43
MOB2_YEAST (P43563) Maintenance of ploidy protein MOB2 (MPS1 bin... 159 6e-39
YK83_CAEEL (P34349) Hypothetical protein C30A5.3 in chromosome III 51 2e-06
YM42_YEAST (Q03214) Hypothetical 162.7 kDa protein in SIP18-SPT2... 33 0.65
R1AB_IBVBC (Q91QT2) Replicase polyprotein 1ab (pp1ab) (ORF1ab po... 32 1.4
R1AB_IBVB (P27920) Replicase polyprotein 1ab (pp1ab) (ORF1ab pol... 32 1.4
CDC4_CANAL (P53699) Cell division control protein 4 30 5.5
SCOB_EMENI (Q00659) Sulfur metabolite repression control protein 29 7.1
KPCL_RAT (Q64617) Protein kinase C, eta type (EC 2.7.1.-) (nPKC-... 29 7.1
KPCL_MOUSE (P23298) Protein kinase C, eta type (EC 2.7.1.-) (nPK... 29 7.1
KPCL_HUMAN (P24723) Protein kinase C, eta type (EC 2.7.1.-) (nPK... 29 7.1
STAU_DROME (P25159) Maternal effect protein staufen 29 9.3
PM21_CHLPN (Q9Z6U5) Probable outer membrane protein pmp21 precur... 29 9.3
>MOB1_SCHPO (O94360) Maintenance of ploidy protein mob1
Length = 210
Score = 251 bits (642), Expect = 7e-67
Identities = 114/211 (54%), Positives = 160/211 (75%), Gaps = 2/211 (0%)
Query: 3 LFGLG-RNQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAVNT 61
+FG + KTFR +K+ +G+K QL+++ +ATLGSG+L EAVKLP GED+NEW+A+NT
Sbjct: 1 MFGFSNKTAKTFRVRKTE-AGTKHYQLRQYAEATLGSGSLMEAVKLPKGEDLNEWIAMNT 59
Query: 62 VDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDW 121
+DF+ Q+N+L+GT+TEFCT ++CP M AGP YEY W D KP +SAP Y+ L+DW
Sbjct: 60 MDFYTQINMLYGTITEFCTAASCPQMNAGPSYEYYWQDDKIYTKPTRMSAPDYINNLLDW 119
Query: 122 IESQLDDETIFPQKLGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLNT 181
+ +LDD+ +FP ++G FP NFR V++ IF+RLFR+YAHIY SHF +V+++ E++LNT
Sbjct: 120 TQEKLDDKKLFPTEIGVEFPKNFRKVIQQIFRRLFRIYAHIYCSHFHVMVAMELESYLNT 179
Query: 182 CFKHFVLFTWEFRLIDKAELAPLEDLVDSII 212
FKHFV F EF L+D E AP++DLVDS++
Sbjct: 180 SFKHFVFFCREFGLMDNKEYAPMQDLVDSMV 210
>MOB1_NEUCR (Q9P601) Probable maintenance of ploidy protein mob1
Length = 219
Score = 221 bits (562), Expect = 1e-57
Identities = 106/217 (48%), Positives = 145/217 (65%), Gaps = 4/217 (1%)
Query: 1 MSLFGLGRNQKT---FRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWL 57
MS F NQ+T FRP+ S G+ QL+++ +ATLG G+LR+ VKLP GED NEWL
Sbjct: 1 MSSFLTTVNQRTRNQFRPRASGKGGATSYQLRQYAEATLGGGSLRKVVKLPEGEDENEWL 60
Query: 58 AVNTVDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEY 117
AVN VDF+NQ+N+L+G +TEFC+P CP M A ++EY W D K+P ++ AP Y+E
Sbjct: 61 AVNMVDFYNQINLLYGAITEFCSPQTCPEMKATDEFEYLWQDTENYKRPTKMPAPAYIEQ 120
Query: 118 LMDWIESQLDDETIFPQKLGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEA 177
LM W++ +D+E + P ++G PFP +F +V+ IFKR++RVYAHIY H+ I L E
Sbjct: 121 LMSWVQGNIDNEAVLPSRIGVPFPKSFPALVRQIFKRMYRVYAHIYCHHYPVIRELGLEP 180
Query: 178 HLNTCFKHFVLFTWEFRLIDKAEL-APLEDLVDSIIQ 213
HLNT FK +VLF E L + PL DLVDS+++
Sbjct: 181 HLNTSFKQYVLFIDEHNLATGKDFWGPLGDLVDSMLR 217
>MOB1_YEAST (P40484) Maintenance of ploidy protein MOB1 (MPS1 binder
1)
Length = 236
Score = 195 bits (496), Expect = 6e-50
Identities = 99/202 (49%), Positives = 136/202 (67%), Gaps = 3/202 (1%)
Query: 9 NQKTFRPKKSAPSGSKGAQLQKHIDATLGS-GNLREAVKLPPGEDINEWLAVNTVDFFNQ 67
+QK F ++ + + +++ ++ TLGS G L +AVKLP GED NEWLAV+ VDF+NQ
Sbjct: 30 HQKPFLQPQAGTTVTTHQDIKQIVEMTLGSEGVLNQAVKLPRGEDENEWLAVHCVDFYNQ 89
Query: 68 VNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDWIESQLD 127
+N+L+G++TEFC+P CP M A +YEY WA + P+ VSAPKYVE LM W + Q D
Sbjct: 90 INMLYGSITEFCSPQTCPRMIATNEYEYLWAFQKG-QPPVSVSAPKYVECLMRWCQDQFD 148
Query: 128 DETIFPQKLGAPFPPNF-RDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLNTCFKHF 186
DE++FP K+ FP F + V++ I +RLFRVYAHIY HF +I+ L + LNT F+HF
Sbjct: 149 DESLFPSKVTGTFPEGFIQRVIQPILRRLFRVYAHIYCHHFNEILELNLQTVLNTSFRHF 208
Query: 187 VLFTWEFRLIDKAELAPLEDLV 208
LF EF L+ A+ PL +LV
Sbjct: 209 CLFAQEFELLRPADFGPLLELV 230
>MOB2_SCHPO (O74558) Maintenance of ploidy protein mob2
Length = 244
Score = 173 bits (438), Expect = 3e-43
Identities = 75/180 (41%), Positives = 116/180 (63%), Gaps = 2/180 (1%)
Query: 29 QKHIDATLGSGNLREAVKLPPGEDINEWLAVNTVDFFNQVNILFGTLTEFCTPSNCPSMT 88
Q + L GN V LP D++EW+A+N + F +N + FCT CP M+
Sbjct: 58 QPFVRTHLVKGNFSTIVSLPRFVDLDEWVALNVYELFTYLNHFYDVFATFCTVKTCPVMS 117
Query: 89 AGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDWIESQLDDETIFPQKLGAPFPPNFRDVV 148
A ++Y W D +KP+ + AP+Y+EY++ WIE++L D+ +FP K G PFP NF +V
Sbjct: 118 AAANFDYTWLDNN--RKPVHLPAPQYIEYVLAWIENRLHDQNVFPTKAGLPFPSNFLVIV 175
Query: 149 KTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLNTCFKHFVLFTWEFRLIDKAELAPLEDLV 208
K I+K++FR++AH+Y++H+ +I+ L EAH N+ F HF+ F EF+L+DK + APL+DL+
Sbjct: 176 KAIYKQMFRIFAHMYYAHYAEILHLSLEAHWNSFFAHFIAFGKEFQLLDKRDTAPLKDLI 235
>MOB2_YEAST (P43563) Maintenance of ploidy protein MOB2 (MPS1 binder
2)
Length = 259
Score = 159 bits (401), Expect = 6e-39
Identities = 71/179 (39%), Positives = 111/179 (61%), Gaps = 2/179 (1%)
Query: 32 IDATLGSGNLREAVKLPPGEDINEWLAVNTVDFFNQVNILFGTLTEFCTPSNCPSMTAGP 91
+ L G+ + V+LP D+ EW+A+N +FF +N +G + E+ TP P+M AGP
Sbjct: 73 VRTALVKGSFKTIVQLPKYVDLGEWIALNVFEFFTNLNQFYGVVAEYVTPDAYPTMNAGP 132
Query: 92 KYEYRWADGVTIKKPIEVSAPKYVEYLMDWIESQLDDETIFPQKLGAPFPPNFRDVVKTI 151
+Y W D + + + A +Y++ + WI ++++D+ +FP K G PFP F V+ I
Sbjct: 133 HTDYLWLDANN--RQVSLPASQYIDLALTWINNKVNDKNLFPTKNGLPFPQQFSRDVQRI 190
Query: 152 FKRLFRVYAHIYHSHFQKIVSLKEEAHLNTCFKHFVLFTWEFRLIDKAELAPLEDLVDS 210
++FR++AHIYH HF KIV L EAH N+ F HF+ F EF++ID+ E+APL L++S
Sbjct: 191 MVQMFRIFAHIYHHHFDKIVHLSLEAHWNSFFSHFISFAKEFKIIDRKEMAPLLPLIES 249
>YK83_CAEEL (P34349) Hypothetical protein C30A5.3 in chromosome III
Length = 223
Score = 50.8 bits (120), Expect = 2e-06
Identities = 41/180 (22%), Positives = 79/180 (43%), Gaps = 18/180 (10%)
Query: 28 LQKHIDATLGSGNLREAVKLPPGEDINE--WLAVNTVDFFNQVNILFGTLTEFCTPSNCP 85
+Q++I T+ + A L P D +E W + F ++N L L C P C
Sbjct: 39 IQQYIQQTIKANPADVATILTPPLDQDEGVWKYEHLRQFCIELNGLALLLQRECIPETCQ 98
Query: 86 SMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDWIESQLDDETIFPQKLGAPFPPNFR 145
MTA ++ + A K P E A Y + +D + L+ FP ++ N +
Sbjct: 99 QMTATEQWIFLCA---AHKNPNECPAIDYTRHTLDGAATLLNSNKYFPSRV------NIK 149
Query: 146 DV----VKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLNTCFKHFVLFTWEFRLIDKAEL 201
++ + ++ +R++R+++H + H + + E HL K F + ++ L+ + L
Sbjct: 150 EISISKLGSVARRVYRIFSHAFFHHRKLFDEFENETHL---CKRFTTYVSKYNLMQQEHL 206
>YM42_YEAST (Q03214) Hypothetical 162.7 kDa protein in SIP18-SPT21
intergenic region
Length = 1411
Score = 32.7 bits (73), Expect = 0.65
Identities = 29/100 (29%), Positives = 42/100 (42%), Gaps = 6/100 (6%)
Query: 72 FGTLTEFCTPSNCPSMTAGPKYEYR-WADGVTI--KKPIEV-SAPKYVEYLMDWIESQLD 127
F +LTEF + T P Y++ W +G+ I K E ++P Y L + E L
Sbjct: 349 FNSLTEFKLGYTQSTETTLPGYDFTFWENGMEIYDKSKYETKTSPVY--NLRQYYEKSLA 406
Query: 128 DETIFPQKLGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHF 167
T K G+ +P F K R+Y H+ HF
Sbjct: 407 VFTAIVAKFGSSYPDLFAKHTTLPQKEFERLYFHLLSEHF 446
>R1AB_IBVBC (Q91QT2) Replicase polyprotein 1ab (pp1ab) (ORF1ab
polyprotein) [Includes: Replicase polyprotein 1a (pp1a)
(ORF1a)] [Contains: p87; p195 (EC 3.4.24.-) (Papain-like
proteinase) (PL-PRO); Peptide HD2 (p41); 3C-like
proteinase (EC 3.4.24.-) (3CL-
Length = 6629
Score = 31.6 bits (70), Expect = 1.4
Identities = 20/45 (44%), Positives = 25/45 (55%), Gaps = 4/45 (8%)
Query: 104 KKPIEVSAPKYVEYLMDWI-ESQLDDETIFPQKLGAPFPPNFRDV 147
K+PIEV VE L+ I E DD +FP+ AP PP F +V
Sbjct: 715 KEPIEVDTDLTVEQLLSVIYEKMCDDLKLFPE---APEPPPFENV 756
>R1AB_IBVB (P27920) Replicase polyprotein 1ab (pp1ab) (ORF1ab
polyprotein) [Includes: Replicase polyprotein 1a (pp1a)
(ORF1a)] [Contains: p87; p195 (EC 3.4.24.-) (Papain-like
proteinase) (PL-PRO); Peptide HD2 (p41); 3C-like
proteinase (EC 3.4.24.-) (3CL-P
Length = 6629
Score = 31.6 bits (70), Expect = 1.4
Identities = 20/45 (44%), Positives = 25/45 (55%), Gaps = 4/45 (8%)
Query: 104 KKPIEVSAPKYVEYLMDWI-ESQLDDETIFPQKLGAPFPPNFRDV 147
K+PIEV VE L+ I E DD +FP+ AP PP F +V
Sbjct: 715 KEPIEVDTDLTVEQLLSVIYEKMCDDLKLFPE---APEPPPFENV 756
>CDC4_CANAL (P53699) Cell division control protein 4
Length = 684
Score = 29.6 bits (65), Expect = 5.5
Identities = 17/62 (27%), Positives = 26/62 (41%), Gaps = 10/62 (16%)
Query: 161 HIYHSHFQKIVSLKEEAHLNTCFKHFV---LFTWEFR-------LIDKAELAPLEDLVDS 210
H+ H ++ S + H TCF + + W F L+ A L L DLVD
Sbjct: 515 HVLQGHLDRVYSTAIDFHSKTCFSGSMDSNINVWNFETGELKKVLVGHASLVGLLDLVDD 574
Query: 211 II 212
++
Sbjct: 575 VL 576
>SCOB_EMENI (Q00659) Sulfur metabolite repression control protein
Length = 678
Score = 29.3 bits (64), Expect = 7.1
Identities = 19/68 (27%), Positives = 33/68 (47%), Gaps = 1/68 (1%)
Query: 18 SAPSGSKGAQLQKHIDAT-LGSGNLREAVKLPPGEDINEWLAVNTVDFFNQVNILFGTLT 76
S +G +G + H DA+ L SG++ + VK+ ED + + D+ N V + + T
Sbjct: 423 STYTGHRGGVIGLHFDASILASGSVDKTVKIWNFEDKSTFSLRGHTDWVNAVRVDTSSRT 482
Query: 77 EFCTPSNC 84
F +C
Sbjct: 483 VFSASDDC 490
>KPCL_RAT (Q64617) Protein kinase C, eta type (EC 2.7.1.-)
(nPKC-eta) (PKC-L)
Length = 683
Score = 29.3 bits (64), Expect = 7.1
Identities = 17/57 (29%), Positives = 28/57 (48%), Gaps = 1/57 (1%)
Query: 77 EFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDWIESQLDDETIFP 133
++ P M GP ++ WA GV + + + AP E D E+ L+DE ++P
Sbjct: 519 DYIAPEILQEMLYGPAVDW-WAMGVLLYEMLCGHAPFEAENEDDLFEAILNDEVVYP 574
>KPCL_MOUSE (P23298) Protein kinase C, eta type (EC 2.7.1.-)
(nPKC-eta) (PKC-L)
Length = 683
Score = 29.3 bits (64), Expect = 7.1
Identities = 17/57 (29%), Positives = 28/57 (48%), Gaps = 1/57 (1%)
Query: 77 EFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDWIESQLDDETIFP 133
++ P M GP ++ WA GV + + + AP E D E+ L+DE ++P
Sbjct: 519 DYIAPEILQEMLYGPAVDW-WAMGVLLYEMLCGHAPFEAENEDDLFEAILNDEVVYP 574
>KPCL_HUMAN (P24723) Protein kinase C, eta type (EC 2.7.1.-)
(nPKC-eta) (PKC-L)
Length = 682
Score = 29.3 bits (64), Expect = 7.1
Identities = 17/57 (29%), Positives = 28/57 (48%), Gaps = 1/57 (1%)
Query: 77 EFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDWIESQLDDETIFP 133
++ P M GP ++ WA GV + + + AP E D E+ L+DE ++P
Sbjct: 518 DYIAPEILQEMLYGPAVDW-WAMGVLLYEMLCGHAPFEAENEDDLFEAILNDEVVYP 573
>STAU_DROME (P25159) Maternal effect protein staufen
Length = 1026
Score = 28.9 bits (63), Expect = 9.3
Identities = 28/92 (30%), Positives = 42/92 (45%), Gaps = 5/92 (5%)
Query: 31 HIDATLGSGNLREAVKLPPGEDINEWLAVNTVDFFNQVNILFGTLTEFCTPSNCPSMTAG 90
+IDAT N + G+D VN + +N++ + LTE P++C + T
Sbjct: 288 NIDATGALSNEDTSSSGRGGKDKTPMCLVNELARYNKITHQY-RLTEERGPAHCKTFTVT 346
Query: 91 PKY---EYRWADGVTIKKPIEVSAPKYVEYLM 119
EY ADG IKK ++A K +E M
Sbjct: 347 LMLGDEEYS-ADGFKIKKAQHLAASKAIEETM 377
>PM21_CHLPN (Q9Z6U5) Probable outer membrane protein pmp21 precursor
(Polymorphic membrane protein 21)
Length = 1609
Score = 28.9 bits (63), Expect = 9.3
Identities = 27/105 (25%), Positives = 46/105 (43%), Gaps = 26/105 (24%)
Query: 25 GAQLQKHIDATLGSGNLREAVKLPPGEDIN-----------EWLAVNTVDFFNQVNILFG 73
GA L +HI T +GNLR + L GE+ + L+ N V+ + N++F
Sbjct: 695 GAILAQHIFITDNTGNLRFSGNLGGGEESSTVGDLAIVGGGALLSTNEVNVCSNQNVVF- 753
Query: 74 TLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYL 118
++ T + C S G + K +++SA VE++
Sbjct: 754 --SDNVTSNGCDS------------GGAILAKKVDISANHSVEFV 784
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.139 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,566,501
Number of Sequences: 164201
Number of extensions: 1118196
Number of successful extensions: 2253
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 2241
Number of HSP's gapped (non-prelim): 16
length of query: 214
length of database: 59,974,054
effective HSP length: 106
effective length of query: 108
effective length of database: 42,568,748
effective search space: 4597424784
effective search space used: 4597424784
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0137.5


