

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0136.7
(841 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
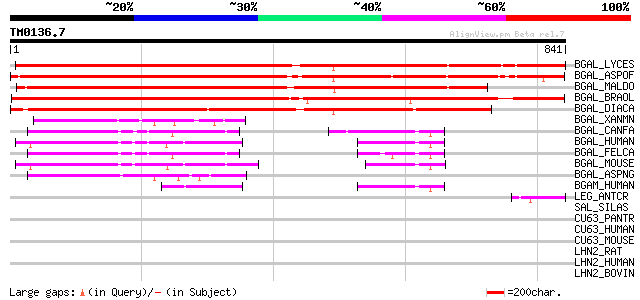
Score E
Sequences producing significant alignments: (bits) Value
BGAL_LYCES (P48980) Beta-galactosidase precursor (EC 3.2.1.23) (... 966 0.0
BGAL_ASPOF (P45582) Beta-galactosidase precursor (EC 3.2.1.23) (... 965 0.0
BGAL_MALDO (P48981) Beta-galactosidase precursor (EC 3.2.1.23) (... 863 0.0
BGAL_BRAOL (P49676) Beta-galactosidase precursor (EC 3.2.1.23) (... 842 0.0
BGAL_DIACA (Q00662) Putative beta-galactosidase precursor (EC 3.... 771 0.0
BGAL_XANMN (P48982) Beta-galactosidase precursor (EC 3.2.1.23) (... 162 3e-39
BGAL_CANFA (Q9TRY9) Beta-galactosidase precursor (EC 3.2.1.23) (... 156 2e-37
BGAL_HUMAN (P16278) Beta-galactosidase precursor (EC 3.2.1.23) (... 153 2e-36
BGAL_FELCA (O19015) Beta-galactosidase precursor (EC 3.2.1.23) (... 150 1e-35
BGAL_MOUSE (P23780) Beta-galactosidase precursor (EC 3.2.1.23) (... 148 5e-35
BGAL_ASPNG (P29853) Beta-galactosidase precursor (EC 3.2.1.23) (... 112 5e-24
BGAM_HUMAN (P16279) Beta-galactosidase-related protein precursor... 52 9e-06
LEG_ANTCR (P22031) D-galactoside-specific lectin (Sea urchin egg... 49 7e-05
SAL_SILAS (Q9PVW8) Rhamnose-binding lectin precursor (SAL) (RBL)... 40 0.026
CU63_PANTR (Q68US5) Protein C21orf63 homolog precursor 40 0.026
CU63_HUMAN (P58658) Protein C21orf63 precursor (Protein PRED34) ... 40 0.026
CU63_MOUSE (P58659) Protein C21orf63 homolog precursor 38 0.13
LHN2_RAT (O88923) Latrophilin 2 precursor (Calcium-independent a... 36 0.48
LHN2_HUMAN (O95490) Latrophilin 2 precursor (Calcium-independent... 36 0.48
LHN2_BOVIN (O97817) Latrophilin 2 precursor (Calcium-independent... 36 0.48
>BGAL_LYCES (P48980) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase) (Acid beta-galactosidase)
(Exo-(1-->4)-beta-D-galactanase)
Length = 835
Score = 966 bits (2496), Expect = 0.0
Identities = 476/846 (56%), Positives = 593/846 (69%), Gaps = 31/846 (3%)
Query: 10 LLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDG 69
+LL+ LLC++ +C A+V+YDH+A++++G+R++LISGSIHYPRSTPEMWPDLIQKAK+G
Sbjct: 7 MLLMLLLCLWV-SCGIASVSYDHKAIIVNGQRKILISGSIHYPRSTPEMWPDLIQKAKEG 65
Query: 70 GLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGF 129
G+DVI+TYVFWN HEP +G+Y FE R DLVKF+K V AGLYVHLRIGPYACAEWN+GGF
Sbjct: 66 GVDVIQTYVFWNGHEPEEGKYYFEERYDLVKFIKVVQEAGLYVHLRIGPYACAEWNFGGF 125
Query: 130 PLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVE 189
P+WL ++PGI FRT+NEPFKA MQ+FT KIVDMMK E LY +QGGPIILSQIENEYG +E
Sbjct: 126 PVWLKYVPGISFRTNNEPFKAAMQKFTTKIVDMMKAEKLYETQGGPIILSQIENEYGPME 185
Query: 190 GAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPNSKAKPK 249
G Y WAA MA L TGVPW+MC+Q++ PDPIINTCNGFYCD FTPN KPK
Sbjct: 186 WELGEPGKVYSEWAAKMAVDLGTGVPWIMCKQDDVPDPIINTCNGFYCDYFTPNKANKPK 245
Query: 250 FWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPFV 309
WTE + W FGG VPYRP ED+AFAVARF Q GG+ NYYMYHGGTNFGRTSGGPF+
Sbjct: 246 MWTEAWTAWFTEFGGPVPYRPAEDMAFAVARFIQTGGSFINYYMYHGGTNFGRTSGGPFI 305
Query: 310 ATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAVYKTE 369
ATSYDYDA +DE+G +RQPKWGHLKD+H+AIKLCE AL++ DPT+TSLG+ EA V+K+E
Sbjct: 306 ATSYDYDAPLDEFGSLRQPKWGHLKDLHRAIKLCEPALVSVDPTVTSLGNYQEARVFKSE 365
Query: 370 A-ECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFTT 428
+ C AFLAN + S A V F YNLP WS+SILPDCKN V NTA++ ++S T
Sbjct: 366 SGACAAFLANYNQHSFAKVAFGNMHYNLPPWSISILPDCKNTVYNTARVGAQSAQMKMTP 425
Query: 429 ESLKEVGSLEGPSPGWSW--ISEPIGISKADSFSRFGLLEQINTTADRSDYLWYSLRIDL 486
S G+SW +E + D+F+ GLLEQIN T D SDYLWY I++
Sbjct: 426 -----------VSRGFSWESFNEDAASHEDDTFTVVGLLEQINITRDVSDYLWYMTDIEI 474
Query: 487 ED-----DAGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTI 541
+ ++G L + S GHALH F+NG+LAG+ G+ +N K+ I L AG N I
Sbjct: 475 DPTEGFLNSGNWPWLTVFSAGHALHVFVNGQLAGTVYGSLENPKLTFSNGINLRAGVNKI 534
Query: 542 DLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDLGLPS 601
LLS+ VGL + GP F+TW AG+ GPV L GL G T D + ++W Y++GLKGE L L S
Sbjct: 535 SLLSIAVGLPNVGPHFETWNAGVLGPVSLNGLNEG-TRDLTWQKWFYKVGLKGEALSLHS 593
Query: 602 ---GTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYW 658
S +W S + + QPL+WYK F+AP GN+P+A+D MGKG+ W+NGQS+GR+W
Sbjct: 594 LSGSPSVEWVEGSLVAQKQPLSWYKTTFNAPDGNEPLALDMNTMGKGQVWINGQSLGRHW 653
Query: 659 PTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGD 718
P Y S G S CNY G +D KCL NCG+ SQ YHVPRSWL P N LV+FEE GGD
Sbjct: 654 PAYKS--SGSCSVCNYTGWFDEKKCLTNCGEGSQRWYHVPRSWLYPTGNLLVVFEEWGGD 711
Query: 719 PTQISIAIKLIGSSCSHVSESHPPPVDLWNS--DTESDRSGGPVLSLECPYPNEVITTIK 776
P I++ + IGS C+ + E P ++ W + DR P L+C P + I++IK
Sbjct: 712 PYGITLVKREIGSVCADIYEWQPQLLN-WQRLVSGKFDRPLRPKAHLKCA-PGQKISSIK 769
Query: 777 FASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTF-GDPCGGVTKSLA 835
FASFGTP G CGNF G C + ++ +K C+G SCS+ V+ F GDPC V K L+
Sbjct: 770 FASFGTPEGVCGNFQQGSCHAPRSYDAFKKNCVGKESCSVQVTPENFGGDPCRNVLKKLS 829
Query: 836 VEASCT 841
VEA C+
Sbjct: 830 VEAICS 835
>BGAL_ASPOF (P45582) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase)
Length = 832
Score = 965 bits (2494), Expect = 0.0
Identities = 474/851 (55%), Positives = 587/851 (68%), Gaps = 31/851 (3%)
Query: 1 MAMRRTQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWP 60
MA++ L++ L L V++P A+VTYDH++++I+G+RR+LISGSIHYPRSTPEMWP
Sbjct: 1 MALKLVLMLMVAL-LAAVWSPPAVTASVTYDHKSVIINGQRRILISGSIHYPRSTPEMWP 59
Query: 61 DLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYA 120
DLIQKAKDGGLDVI+TYVFWN HEP GQY F GR DLV+F+K V AGLY HLRIGPY
Sbjct: 60 DLIQKAKDGGLDVIQTYVFWNGHEPSPGQYYFGGRYDLVRFLKLVKQAGLYAHLRIGPYV 119
Query: 121 CAEWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQ 180
CAEWN+GGFP+WL ++PGI FRTDN PFKA M +FT KIV MMK E LY +QGGPIILSQ
Sbjct: 120 CAEWNFGGFPVWLKYVPGIHFRTDNGPFKAAMGKFTEKIVSMMKAEGLYETQGGPIILSQ 179
Query: 181 IENEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQF 240
IENEYG VE G A Y NWAA MA L+TGVPWVMC+Q++APDP+INTCNGFYCD F
Sbjct: 180 IENEYGPVEYYDGAAGKSYTNWAAKMAVGLNTGVPWVMCKQDDAPDPVINTCNGFYCDYF 239
Query: 241 TPNSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNF 300
+PN KPK WTE + GW FGGAVP RP ED+AFAVARF Q GG+ NYYMYHGGTNF
Sbjct: 240 SPNKDNKPKMWTEAWTGWFTGFGGAVPQRPAEDMAFAVARFIQKGGSFINYYMYHGGTNF 299
Query: 301 GRTSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSN 360
GRT+GGPF++TSYDYDA IDEYG +RQPKWGHL+D+HKAIKLCE AL++ +PTITSLG N
Sbjct: 300 GRTAGGPFISTSYDYDAPIDEYGLLRQPKWGHLRDLHKAIKLCEPALVSGEPTITSLGQN 359
Query: 361 LEAAVYKTEAECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSE 420
E+ VY++++ C AFLAN ++ ATV FNG YNLP WSVSILPDCK V NTA++ ++
Sbjct: 360 QESYVYRSKSSCAAFLANFNSRYYATVTFNGMHYNLPPWSVSILPDCKTTVFNTARVGAQ 419
Query: 421 SMISSFTTESLKEVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSDYLWY 480
+ TT ++ +G W +E ++F++ GL+EQ++TT DRSDYLWY
Sbjct: 420 T-----TTMKMQYLGGF-----SWKAYTEDTDALNDNTFTKDGLVEQLSTTWDRSDYLWY 469
Query: 481 SLRIDLEDD-----AGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLV 535
+ +D+ + G L + S GHA+H FING+L+G+ G+ DN K+ L
Sbjct: 470 TTYVDIAKNEEFLKTGKYPYLTVMSAGHAVHVFINGQLSGTAYGSLDNPKLTYSGSAKLW 529
Query: 536 AGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGE 595
AG N I +LS++VGL + G F+TW G+ GPV L GL G D S ++WTYQIGL GE
Sbjct: 530 AGSNKISILSVSVGLPNVGNHFETWNTGVLGPVTLTGLNEGKR-DLSLQKWTYQIGLHGE 588
Query: 596 DLGLPSGTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIG 655
L L S T + QPLTWYK F+AP GN+P+A+D MGKG+ W+NGQSIG
Sbjct: 589 TLSLHSLTGSSNVEWGEASQKQPLTWYKTFFNAPPGNEPLALDMNTMGKGQIWINGQSIG 648
Query: 656 RYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEES 715
RYWP Y G SC+YRG Y+ KCL NCG+ SQ YHVPRSWL P N LV+ EE
Sbjct: 649 RYWPAY--KASGSCGSCDYRGTYNEKKCLSNCGEASQRWYHVPRSWLIPTGNFLVVLEEW 706
Query: 716 GGDPTQISIAIKLIGSSCSHVSESHPPPVDLWNSDTESDRSGGPVLSLECPYPNEVITTI 775
GGDPT IS+ + + S C+ V E P +D W + G P + L C P + ++ I
Sbjct: 707 GGDPTGISMVKRSVASVCAEVEELQ-PTMDNWRTKA----YGRPKVHLSCD-PGQKMSKI 760
Query: 776 KFASFGTPHGNCGNFSHGDCSSKQALSIVQKA-----CIGSSSCSIGVSINTF-GDPCGG 829
KFASFGTP G CG+FS G C + ++ ++ C+G CS+ V+ F GDPC G
Sbjct: 761 KFASFGTPQGTCGSFSEGSCHAHKSYDAFEQEGLMQNCVGQEFCSVNVAPEVFGGDPCPG 820
Query: 830 VTKSLAVEASC 840
K LAVEA C
Sbjct: 821 TMKKLAVEAIC 831
>BGAL_MALDO (P48981) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase) (Acid beta-galactosidase)
(Exo-(1-->4)-beta-D-galactanase)
Length = 731
Score = 863 bits (2231), Expect = 0.0
Identities = 413/722 (57%), Positives = 518/722 (71%), Gaps = 21/722 (2%)
Query: 11 LLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGG 70
+LL C+++ A A+V+YDH+A++I+G++R+LISGSIHYPRSTPEMWPDLIQKAKDGG
Sbjct: 11 ILLLFSCIFSAAS--ASVSYDHKAIIINGQKRILISGSIHYPRSTPEMWPDLIQKAKDGG 68
Query: 71 LDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFP 130
LDVI+TYVFWN HEP G Y FE R DLVKF+K V GL+V+LRIGPY CAEWN+GGFP
Sbjct: 69 LDVIQTYVFWNGHEPSPGNYYFEERYDLVKFIKLVQQEGLFVNLRIGPYVCAEWNFGGFP 128
Query: 131 LWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVEG 190
+WL ++PGI FRTDNEPFKA MQ+FT KIV MMK E L+ +QGGPIILSQIENE+G VE
Sbjct: 129 VWLKYVPGIAFRTDNEPFKAAMQKFTEKIVSMMKAEKLFQTQGGPIILSQIENEFGPVEW 188
Query: 191 AYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPNSKAKPKF 250
G Y WAA MA LDTGVPW+MC+QE+APDP+I+TCNGFYC+ F PN KPK
Sbjct: 189 EIGAPGKAYTKWAAQMAVGLDTGVPWIMCKQEDAPDPVIDTCNGFYCENFKPNKDYKPKM 248
Query: 251 WTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPFVA 310
WTE + GW FGGAVP RP ED+AF+VARF Q GG+ NYYMYHGGTNFGRT+GGPF+A
Sbjct: 249 WTEVWTGWYTEFGGAVPTRPAEDVAFSVARFIQSGGSFLNYYMYHGGTNFGRTAGGPFMA 308
Query: 311 TSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAVYKTEA 370
TSYDYDA +DEYG R+PKWGHL+D+HKAIK CE AL++ DP++T LGSN EA V+K+E+
Sbjct: 309 TSYDYDAPLDEYGLPREPKWGHLRDLHKAIKSCESALVSVDPSVTKLGSNQEAHVFKSES 368
Query: 371 ECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFTTES 430
+C AFLAN D V F G Y+LP WS+SILPDCK V NTAK+ S+S S
Sbjct: 369 DCAAFLANYDAKYSVKVSFGGGQYDLPPWSISILPDCKTEVYNTAKVGSQS--------S 420
Query: 431 LKEVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSDYLWYSLRIDLEDDA 490
++ + P S+I E + D+ + GL EQIN T D +DYLWY I + D
Sbjct: 421 QVQMTPVHSGFPWQSFIEETTSSDETDTTTLDGLYEQINITRDTTDYLWYMTDITIGSDE 480
Query: 491 -----GAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLS 545
G +L I S GHAL+ FING+L+G+ G+ +N K++ + L +G N + LLS
Sbjct: 481 AFLKNGKSPLLTIFSAGHALNVFINGQLSGTVYGSLENPKLSFSQNVNLRSGINKLALLS 540
Query: 546 LTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDLGLPSGT-- 603
++VGL + G F+TW AG+ GP+ LKGL +G T D S +WTY+ GLKGE LGL + T
Sbjct: 541 ISVGLPNVGTHFETWNAGVLGPITLKGLNSG-TWDMSGWKWTYKTGLKGEALGLHTVTGS 599
Query: 604 -SGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYI 662
S +W ++ + QPLTWYK F+AP G+ P+A+D MGKG+ W+NGQS+GR+WP YI
Sbjct: 600 SSVEWVEGPSMAEKQPLTWYKATFNAPPGDAPLALDMGSMGKGQIWINGQSVGRHWPGYI 659
Query: 663 SPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPTQI 722
+ G C+Y G YD KC +CG+PSQ YH+PRSWL P N LV+FEE GGDP++I
Sbjct: 660 A--RGSCGDCSYAGTYDDKKCRTHCGEPSQRWYHIPRSWLTPTGNLLVVFEEWGGDPSRI 717
Query: 723 SI 724
S+
Sbjct: 718 SL 719
>BGAL_BRAOL (P49676) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase)
Length = 828
Score = 842 bits (2175), Expect = 0.0
Identities = 423/858 (49%), Positives = 554/858 (64%), Gaps = 50/858 (5%)
Query: 3 MRRTQFLLLLLWLLCVYA-PACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPD 61
M+ QF LL L+L+ + + + V++D RA+ IDG+RR+L+SGSIHYPRST +MWPD
Sbjct: 1 MKMKQFNLLSLFLILITSFGSANSTIVSHDERAITIDGQRRILLSGSIHYPRSTSDMWPD 60
Query: 62 LIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYAC 121
LI KAKDGGLD IETYVFWN HEP + QY+F G DLV+F+K + +AGLY LRIGPY C
Sbjct: 61 LISKAKDGGLDTIETYVFWNAHEPSRRQYDFSGNLDLVRFIKTIQSAGLYSVLRIGPYVC 120
Query: 122 AEWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQI 181
AEWNYGGFP+WLH +P ++FRT N F EMQ FT KIV+MMK+E+L+ASQGGPIIL+QI
Sbjct: 121 AEWNYGGFPVWLHNMPDMKFRTINPGFMNEMQNFTTKIVNMMKEESLFASQGGPIILAQI 180
Query: 182 ENEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFT 241
ENEYGNV +YG YI+W A+MA SLD GVPW+MCQQ +AP P+I TCNGFYCDQ+
Sbjct: 181 ENEYGNVISSYGAEGKAYIDWCANMANSLDIGVPWIMCQQPHAPQPMIETCNGFYCDQYK 240
Query: 242 PNSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFG 301
P++ + PK WTE + GW +GG PYR EDLAF+VARFFQ GGT QNYYMYHGGTNFG
Sbjct: 241 PSNPSSPKMWTENWTGWFKNWGGKHPYRTAEDLAFSVARFFQTGGTFQNYYMYHGGTNFG 300
Query: 302 RTSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNL 361
R +GGP++ TSYDYDA +DEYG + QPKWGHLK +H +K E+ L + + LG+++
Sbjct: 301 RVAGGPYITTSYDYDAPLDEYGNLNQPKWGHLKQLHTLLKSMEKPLTYGNISTIDLGNSV 360
Query: 362 EAAVYKTEAECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSES 421
A VY T + F+ N++ T+DA V F G YN+PAWSVS+LPDC NTA++N+++
Sbjct: 361 TATVYSTNEKSSCFIGNVNATADALVNFKGKDYNVPAWSVSVLPDCDKEAYNTARVNTQT 420
Query: 422 MISSFTTESLKEVGSLEGPSPGWSWISE-----PIGISKADSFSRFGLLEQINTTADRSD 476
I T +S E L+ W+W E I D ++ GL++Q + T D SD
Sbjct: 421 SI--ITEDSCDEPEKLK-----WTWRPEFTTQKTILKGSGDLIAK-GLVDQKDVTNDASD 472
Query: 477 YLWYSLRI--DLEDDAGAQTV-LHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPIT 533
YLWY R+ D +D ++ + L + S H LHA++NGK G+ + E +
Sbjct: 473 YLWYMTRVHLDKKDPIWSRNMSLRVHSNAHVLHAYVNGKYVGNQIVRDNKFDYRFEKKVN 532
Query: 534 LVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTL--DFSSKQWTYQIG 591
LV G N + LLS++VGLQ++GPFF++ GI GPV L G K T+ D S QW Y+IG
Sbjct: 533 LVHGTNHLALLSVSVGLQNYGPFFESGPTGINGPVKLVGYKGDETIEKDLSKHQWDYKIG 592
Query: 592 LKGEDLGLPSGTSG-----QWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGE 646
L G + L S S +W+++ LP ++ L+WYK NF AP G DP+ +D G+GKGE
Sbjct: 593 LNGFNHKLFSMKSAGHHHRKWSTEK-LPADRMLSWYKANFKAPLGKDPVIVDLNGLGKGE 651
Query: 647 AWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDS 706
W+NGQSIGRYWP++ S +GCT C+YRG Y S KC CGKP+Q YHVPRS+L
Sbjct: 652 VWINGQSIGRYWPSFNSSDEGCTEECDYRGEYGSDKCAFMCGKPTQRWYHVPRSFLNDKG 711
Query: 707 -NTLVLFEESGGDPTQISIAIKLIGSSCSHVSESHPPPVDLWNSDTESDRSGGPVLSLEC 765
NT+ LFEE GGDP+ + + G C+ E + +E
Sbjct: 712 HNTITLFEEMGGDPSMVKFKTVVTGRVCAKAHEHN---------------------KVEL 750
Query: 766 PYPNEVITTIKFASFGTPHGNCGNFSHGDC-SSKQALSIVQKACIGSSSCSIGVSINTFG 824
N I+ +KFASFG P G CG+F+ G C +K A+ +V K C+G +C++ VS + FG
Sbjct: 751 SCNNRPISAVKFASFGNPSGQCGSFAAGSCEGAKDAVKVVAKECVGKLNCTMNVSSHKFG 810
Query: 825 D--PCGGVTKSLAVEASC 840
CG K L VE C
Sbjct: 811 SNLDCGDSPKRLFVEVEC 828
>BGAL_DIACA (Q00662) Putative beta-galactosidase precursor (EC
3.2.1.23) (Lactase) (SR12 protein)
Length = 731
Score = 771 bits (1990), Expect = 0.0
Identities = 390/743 (52%), Positives = 494/743 (65%), Gaps = 36/743 (4%)
Query: 1 MAMRRTQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWP 60
M M +L+ + CVY NV YD+RA+ I+ +RR+L+SGSIHYPRSTPEMWP
Sbjct: 10 MKMMLVYVFVLITLISCVYG------NVWYDYRAIKINDQRRILLSGSIHYPRSTPEMWP 63
Query: 61 DLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYA 120
D+I+KAKD LDVI+TYVFWN HEP +G+Y FEGR DLVKF+K + AGL+VHLRIGP+A
Sbjct: 64 DIIEKAKDSQLDVIQTYVFWNGHEPSEGKYYFEGRYDLVKFIKLIHQAGLFVHLRIGPFA 123
Query: 121 CAEWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQ 180
CAEWN+GGFP+WL ++PGI+FRTDN PFK +MQ FT KIVDMMK E L+ QGGPIIL+Q
Sbjct: 124 CAEWNFGGFPVWLKYVPGIEFRTDNGPFKEKMQVFTTKIVDMMKAEKLFHWQGGPIILNQ 183
Query: 181 IENEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQE-NAPDPIINTCNGFYCDQ 239
IENEYG VE G Y +WAA MA SL+ GVPW+MC+Q+ + PD +I+TCNGFYC+
Sbjct: 184 IENEYGPVEWEIGAPGKAYTHWAAQMAQSLNAGVPWIMCKQDSDVPDNVIDTCNGFYCEG 243
Query: 240 FTPNSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTN 299
F P K+KPK WTE + GW +G VPYRP ED+AF+VARF Q GG+ NYYM+HGGTN
Sbjct: 244 FVPKDKSKPKMWTENWTGWYTEYGKPVPYRPAEDVAFSVARFIQNGGSFMNYYMFHGGTN 303
Query: 300 FGRTSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGS 359
F T+ G FV+TSYDYDA +DEYG R+PK+ HLK++HKAIK+CE AL+++D +T+LGS
Sbjct: 304 F-ETTAGRFVSTSYDYDAPLDEYGLPREPKYTHLKNLHKAIKMCEPALVSSDAKVTNLGS 362
Query: 360 NLEAAVYKTEA-ECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKIN 418
N EA VY + + C AFLAN D V F+G + LPAWS+SILPDCK V NTA++N
Sbjct: 363 NQEAHVYSSNSGSCAAFLANYDPKWSVKVTFSGMEFELPAWSISILPDCKKEVYNTARVN 422
Query: 419 SES-MISSFTTESLKEVGSLEGPSPGWSWISEPIGISKADS---FSRFGLLEQINTTADR 474
S + S T + + +W S + ADS F L EQIN T D+
Sbjct: 423 EPSPKLHSKMTPVISNL----------NWQSYSDEVPTADSPGTFREKKLYEQINMTWDK 472
Query: 475 SDYLWYSLRIDLEDD-----AGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVE 529
SDYLWY + L+ + G + L + S GH LH F+NG+L G G+ ++
Sbjct: 473 SDYLWYMTDVVLDGNEGFLKKGDEPWLTVNSAGHVLHVFVNGQLQGHAYGSLAKPQLTFS 532
Query: 530 IPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQ 589
+ + AG N I LLS VGL + G F+ + G+ GPV L GL G T D + + W+Y+
Sbjct: 533 QKVKMTAGVNRISLLSAVVGLANVGWHFERYNQGVLGPVTLSGLNEG-TRDLTWQYWSYK 591
Query: 590 IGLKGEDLGL-PSGTSG--QWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGE 646
IG KGE+ + SG S QW + QPL WYK F AP GNDP+A+D MGKG+
Sbjct: 592 IGTKGEEQQVYNSGGSSHVQWGPPAW---KQPLVWYKTTFDAPGGNDPLALDLGSMGKGQ 648
Query: 647 AWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDS 706
AW+NGQSIGR+W I+ C +CNY G Y +KCL +CGK SQ YHVPRSWLQP
Sbjct: 649 AWINGQSIGRHWSNNIAK-GSCNDNCNYAGTYTETKCLSDCGKSSQKWYHVPRSWLQPRG 707
Query: 707 NTLVLFEESGGDPTQISIAIKLI 729
N LV+FEE GGD +S+ + I
Sbjct: 708 NLLVVFEEWGGDTKWVSLVKRTI 730
>BGAL_XANMN (P48982) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase)
Length = 598
Score = 162 bits (410), Expect = 3e-39
Identities = 114/346 (32%), Positives = 161/346 (45%), Gaps = 35/346 (10%)
Query: 36 VIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGR 95
V DGK L+SG+IH+ R W D +QKA+ GL+ +ETYVFWNL EP QGQ++F G
Sbjct: 38 VRDGKPYQLLSGAIHFQRIPRAYWKDRLQKARALGLNTVETYVFWNLVEPQQGQFDFSGN 97
Query: 96 GDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRF 155
D+ FVK A GL V LR GPYACAEW GG+P WL I+ R+ + F A Q +
Sbjct: 98 NDVAAFVKEAAAQGLNVILRPGPYACAEWEAGGYPAWLFGKGNIRVRSRDPRFLAASQAY 157
Query: 156 TAKIVDMMKQENLYASQGGPIILSQIENEYGNVEGAYGPAAVPYINWAASMATSLDTGVP 215
+ + + L GGPII Q+ENEYG+ + A N A + D +
Sbjct: 158 LDALAKQV--QPLLNHNGGPIIAVQVENEYGSYADDHAYMA---DNRAMYVKAGFDKALL 212
Query: 216 WV-----MCQQENAPD--PIINTCNGFYCDQFTPNSKAKP-------KFWTEGYNGWLLY 261
+ M PD ++N G F K +P ++W ++ W
Sbjct: 213 FTSDGADMLANGTLPDTLAVVNFAPGEAKSAFDKLIKFRPDQPRMVGEYWAGWFDHWGKP 272
Query: 262 FGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPF----------VAT 311
+ E+ + + + G N YM+ GGT+FG +G F T
Sbjct: 273 HAATDARQQAEEFEWILRQ-----GHSANLYMFIGGTSFGFMNGANFQNNPSDHYAPQTT 327
Query: 312 SYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSL 357
SYDYDA +DE G PK+ ++D + + + T T+L
Sbjct: 328 SYDYDAILDEAGH-PTPKFALMRDAIARVTGVQPPALPAPITTTTL 372
Score = 42.7 bits (99), Expect = 0.004
Identities = 54/253 (21%), Positives = 91/253 (35%), Gaps = 40/253 (15%)
Query: 408 KNVVLNTAKINSESMISSFTTESLKEVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQ 467
++ + + ++ + TT +L E S W + PI I +FG
Sbjct: 349 RDAIARVTGVQPPALPAPITTTTLPATPLRESASL-WDNLPTPIAIDTPQPMEQFG---- 403
Query: 468 INTTADRSDYLWYSLRIDLEDDAGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVN 527
DY + R + L++ + +++ + GS + +
Sbjct: 404 -------QDYGYILYRTTITGPRKGP--LYLGDVRDVARVYVDQRPVGSVERRLQQVSLE 454
Query: 528 VEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWT 587
VEIP AG++T+D+L G ++G AG+ PV+L + F T
Sbjct: 455 VEIP----AGQHTLDVLVENSGRINYGTRMADGRAGLVDPVLLDSQQLTGWQAFPLPMRT 510
Query: 588 YQI--GLKGEDLGLPSGTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKG 645
G G+ + P+ G TL P Y +D GKG
Sbjct: 511 PDSIRGWTGKAVQGPAFHRG------TLRIGTPTDTY--------------LDMRAFGKG 550
Query: 646 EAWVNGQSIGRYW 658
AW NG ++GR+W
Sbjct: 551 FAWANGVNLGRHW 563
>BGAL_CANFA (Q9TRY9) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase) (Acid beta-galactosidase) (Fragment)
Length = 662
Score = 156 bits (395), Expect = 2e-37
Identities = 114/341 (33%), Positives = 162/341 (47%), Gaps = 29/341 (8%)
Query: 28 VTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLHEPVQ 87
+ Y H + DG+ ISGSIHY R W D + K K GL+ I+TYV WN HEP
Sbjct: 29 IDYSHNRFLKDGQPFRYISGSIHYSRVPRFYWKDRLLKMKMAGLNAIQTYVPWNFHEPQP 88
Query: 88 GQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFIPGIQFRTDNEP 147
GQY F G D+ F+K GL V LR GPY CAEW+ GG P WL I R+ +
Sbjct: 89 GQYQFSGEQDVEYFIKLAHELGLLVILRPGPYICAEWDMGGLPAWLLLKESIILRSSDPD 148
Query: 148 FKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVEGAYGPAAVPYINWAASMA 207
+ A + ++ ++ MK L GGPII Q+ENEYG+ Y Y+ + +
Sbjct: 149 YLAAVDKWLGVLLPKMKP--LLYQNGGPIITMQVENEYGS----YFTCDYDYLRFLQKLF 202
Query: 208 TSLDTGVPWVMCQQENAPDPIIN--TCNGFYCD-QFTPNS----------KAKPK---FW 251
G ++ + A + + G Y F P + K++PK
Sbjct: 203 HH-HLGNDVLLFTTDGANEKFLQCGALQGLYATVDFGPGANITAAFQIQRKSEPKGPLVN 261
Query: 252 TEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGG--PFV 309
+E Y GWL ++G E +A ++ G + N YM+ GGTNF +G P+
Sbjct: 262 SEFYTGWLDHWGQPHSTVRTEVVASSLHDILAHGANV-NLYMFIGGTNFAYWNGANMPYQ 320
Query: 310 A--TSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALI 348
A TSYDYDA + E G + + K+ L++V + + E I
Sbjct: 321 AQPTSYDYDAPLSEAGDLTE-KYFALREVIRKFEKVPEGFI 360
Score = 51.2 bits (121), Expect = 1e-05
Identities = 51/184 (27%), Positives = 80/184 (42%), Gaps = 16/184 (8%)
Query: 483 RIDLEDDAGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTID 542
R L D T L G A+++ + G +G + S V + + IT AG T+D
Sbjct: 414 RTTLPQDCSDPTPLSSPLSGVHDRAYVS--VDGVPQGVMERSNV-ITLNITGKAGA-TLD 469
Query: 543 LLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNG---STLDFSSKQWTYQIGLKGEDLGL 599
LL +G ++G + + + I+ + + L+ ++ G G + G
Sbjct: 470 LLVENMGRVNYGRYINDFKGLISNLTLGSSILTNWMIFPLNTEDAVRSHLGGWHGPNNGR 529
Query: 600 PSGTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIA----IDFTGMGKGEAWVNGQSIG 655
T +S TLP +Y NFS PSG + I F G KG+ W+NG ++G
Sbjct: 530 HDKTFAHRSSNYTLP-----AFYMGNFSIPSGIPDLPQDTFIQFPGWTKGQVWINGFNLG 584
Query: 656 RYWP 659
RYWP
Sbjct: 585 RYWP 588
>BGAL_HUMAN (P16278) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase) (Acid beta-galactosidase)
Length = 677
Score = 153 bits (387), Expect = 2e-36
Identities = 120/372 (32%), Positives = 168/372 (44%), Gaps = 37/372 (9%)
Query: 9 LLLLLWLLCVYAPACFCANVT-------YDHRALVIDGKRRVLISGSIHYPRSTPEMWPD 61
+LLLL +L + P N T Y + + DG+ ISGSIHY R W D
Sbjct: 8 ILLLLLVLLLLGPTRGLRNATQRMFEIDYSRDSFLKDGQPFRYISGSIHYSRVPRFYWKD 67
Query: 62 LIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYAC 121
+ K K GL+ I+TYV WN HEP GQY F D+ F++ GL V LR GPY C
Sbjct: 68 RLLKMKMAGLNAIQTYVPWNFHEPWPGQYQFSEDHDVEYFLRLAHELGLLVILRPGPYIC 127
Query: 122 AEWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQI 181
AEW GG P WL I R+ + + A + ++ ++ MK L GGP+I Q+
Sbjct: 128 AEWEMGGLPAWLLEKESILLRSSDPDYLAAVDKWLGVLLPKMKP--LLYQNGGPVITVQV 185
Query: 182 ENEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPII--NTCNGFY--- 236
ENEY G+Y Y+ + G V+ + A + G Y
Sbjct: 186 ENEY----GSYFACDFDYLRFLQKRFRH-HLGDDVVLFTTDGAHKTFLKCGALQGLYTTV 240
Query: 237 --------CDQFTPNSKAKPK---FWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLG 285
D F K +PK +E Y GWL ++G P+ ++ A A + + L
Sbjct: 241 DFGTGSNITDAFLSQRKCEPKGPLINSEFYTGWLDHWG--QPHSTIKTEAVASSLYDILA 298
Query: 286 -GTLQNYYMYHGGTNFGRTSG--GPFVA--TSYDYDAAIDEYGFIRQPKWGHLKDVHKAI 340
G N YM+ GGTNF +G P+ A TSYDYDA + E G + + + + K
Sbjct: 299 RGASVNLYMFIGGTNFAYWNGANSPYAAQPTSYDYDAPLSEAGDLTEKYFALRNIIQKFE 358
Query: 341 KLCEEALIATDP 352
K+ E + + P
Sbjct: 359 KVPEGPIPPSTP 370
Score = 49.7 bits (117), Expect = 3e-05
Identities = 40/139 (28%), Positives = 62/139 (43%), Gaps = 13/139 (9%)
Query: 528 VEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGST---LDFSSK 584
+ + IT AG T+DLL +G ++G + + + ++ + + T LD
Sbjct: 461 ITLNITGKAGA-TLDLLVENMGRVNYGAYINDFKGLVSNLTLSSNILTDWTIFPLDTEDA 519
Query: 585 QWTYQIGLKGEDLGLPSGTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIA----IDFT 640
++ G D G +S TLP +Y NFS PSG + I F
Sbjct: 520 VRSHLGGWGHRDSGHHDEAWAHNSSNYTLP-----AFYMGNFSIPSGIPDLPQDTFIQFP 574
Query: 641 GMGKGEAWVNGQSIGRYWP 659
G KG+ W+NG ++GRYWP
Sbjct: 575 GWTKGQVWINGFNLGRYWP 593
>BGAL_FELCA (O19015) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase) (Acid beta-galactosidase)
Length = 669
Score = 150 bits (380), Expect = 1e-35
Identities = 113/341 (33%), Positives = 160/341 (46%), Gaps = 29/341 (8%)
Query: 28 VTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLHEPVQ 87
+ Y H + DG+ ISGSIHY R W D + K K GL+ I+TYV WN HEP
Sbjct: 35 IDYGHNRFLKDGQPFRYISGSIHYFRVPRFYWKDRLLKMKMAGLNAIQTYVPWNFHEPQP 94
Query: 88 GQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFIPGIQFRTDNEP 147
GQY F G D+ F+K GL V LR GPY CAEW+ GG P WL I R+ +
Sbjct: 95 GQYQFSGEHDVEYFLKLAHELGLLVILRPGPYICAEWDMGGLPAWLLLKESIILRSSDPD 154
Query: 148 FKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVEGAYGPAAVPYINWAASMA 207
+ A + ++ ++ MK L GGPII Q+ENEY G+Y Y+ +
Sbjct: 155 YLAAVDKWLGVLLPKMKP--LLYQNGGPIITVQVENEY----GSYFTCDYDYLRFLQRRF 208
Query: 208 TSLDTGVPWVMCQQENAPDPII--NTCNGFYCD-QFTPNS----------KAKPK---FW 251
G ++ + A + + G Y F P++ K++P+
Sbjct: 209 RD-HLGGDVLLFTTDGAHEKFLQCGALQGIYATVDFGPDANITAAFQIQRKSEPRGPLVN 267
Query: 252 TEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGG--PF- 308
+E Y GWL ++G E +A ++ G + N YM+ GGTNF +G P+
Sbjct: 268 SEFYTGWLDHWGQPHSRVRTEVVASSLHDVLAHGANV-NLYMFIGGTNFAYWNGANIPYQ 326
Query: 309 -VATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALI 348
TSYDYDA + E G + K+ L+DV + + E I
Sbjct: 327 PQPTSYDYDAPLSEAGDLTD-KYFALRDVIRKFEKVPEGFI 366
Score = 47.8 bits (112), Expect = 1e-04
Identities = 42/146 (28%), Positives = 66/146 (44%), Gaps = 26/146 (17%)
Query: 528 VEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGST--------- 578
+ + IT AG T+DLL +G ++G + + + ++ L GS+
Sbjct: 462 ITLNITGQAGA-TLDLLVENMGRVNYGRYINDFKG------LISNLTLGSSVLTDWMIFP 514
Query: 579 LDFSSKQWTYQIGLKGEDLGLPSGTS-GQWNSQSTLPKNQPLTWYKINFSAPSGNDPIA- 636
LD ++ G G + G + +S TLP +Y NFS PSG +
Sbjct: 515 LDTEDAVRSHLGGWHGRNHGRQDNKAFAHHSSNYTLP-----AFYAGNFSIPSGIPDLPQ 569
Query: 637 ---IDFTGMGKGEAWVNGQSIGRYWP 659
I F+G KG+ W+NG ++GRYWP
Sbjct: 570 DTFIQFSGWTKGQVWINGFNLGRYWP 595
>BGAL_MOUSE (P23780) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase) (Acid beta-galactosidase)
Length = 647
Score = 148 bits (374), Expect = 5e-35
Identities = 127/395 (32%), Positives = 177/395 (44%), Gaps = 39/395 (9%)
Query: 10 LLLLWLLCVYAPACFCANVT-------YDHRALVIDGKRRVLISGSIHYPRSTPEMWPDL 62
L LL LL + A NVT Y + DG+ ISGSIHY R W D
Sbjct: 10 LPLLALLQLLGAAHGIYNVTQRTFKLDYSRDRFLKDGQPFRYISGSIHYFRIPRFYWEDR 69
Query: 63 IQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACA 122
+ K K GL+ I+ YV WN HEP GQY F G D+ F++ GL V LR GPY CA
Sbjct: 70 LLKMKMAGLNAIQMYVPWNFHEPQPGQYEFSGDRDVEHFIQLAHELGLLVILRPGPYICA 129
Query: 123 EWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIE 182
EW+ GG P WL I R+ + + + ++ A ++ MK L GGPII Q+E
Sbjct: 130 EWDMGGLPAWLLEKQSIVLRSSDPDYLVAVDKWLAVLLPKMKP--LLYQNGGPIITVQVE 187
Query: 183 NEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPII--NTCNGFYC--- 237
NEY G+Y Y+ + G ++ + A + ++ T Y
Sbjct: 188 NEY----GSYFACDYDYLRFLVH-RFRYHLGNDVILFTTDGASEKMLKCGTLQDLYATVD 242
Query: 238 --------DQFTPNSKAKPK---FWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLG- 285
F K +PK +E Y GWL ++G P+ V+ A + + L
Sbjct: 243 FGTGNNITQAFLVQRKFEPKGPLINSEFYTGWLDHWG--KPHSTVKTKTLATSLYNLLAR 300
Query: 286 GTLQNYYMYHGGTNFGRTSGG--PF--VATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIK 341
G N YM+ GGTNF +G P+ TSYDYDA + E G + + K+ L++V + K
Sbjct: 301 GANVNLYMFIGGTNFAYWNGANTPYEPQPTSYDYDAPLSEAGDLTK-KYFALREVIQMFK 359
Query: 342 LCEEALIATDPTITSLGSNLEAAVYKTEAECVAFL 376
E I + G + +KT AE + L
Sbjct: 360 EVPEGPIPPSTPKFAYG-KVALRKFKTVAEALGIL 393
Score = 55.5 bits (132), Expect = 6e-07
Identities = 40/128 (31%), Positives = 60/128 (46%), Gaps = 12/128 (9%)
Query: 540 TIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGST---LDFSSKQWTYQIGLKGED 596
T+D+L +G ++G F + + I+ I + T L+ + + G + D
Sbjct: 474 TLDILVENMGRVNYGRFINDFKGLISNMTINSTVLTNWTVFPLNTEAMVRNHLWGREASD 533
Query: 597 LGLPSGTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIA----IDFTGMGKGEAWVNGQ 652
G G S +S LP T+Y NFS PSG + I F G KG+ W+NG
Sbjct: 534 EGHLDGRSTSNSSDLILP-----TFYVGNFSIPSGIPDLPQDTFIQFPGWSKGQVWINGF 588
Query: 653 SIGRYWPT 660
++GRYWPT
Sbjct: 589 NLGRYWPT 596
>BGAL_ASPNG (P29853) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase)
Length = 1006
Score = 112 bits (279), Expect = 5e-24
Identities = 103/370 (27%), Positives = 154/370 (40%), Gaps = 47/370 (12%)
Query: 28 VTYDHRALVIDGKRRVLISGSIH-YPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLHEPV 86
VT+D ++L I+G+R ++ SG H + E+ D+ QK K G + + YV W L E
Sbjct: 46 VTWDDKSLFINGERIMIFSGEFHPFRLPVKELQLDIFQKVKALGFNCVSFYVDWALVEGK 105
Query: 87 QGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFIPGIQFRTDNE 146
G+Y +G DL F + AG+Y+ R GPY AE + GGFP WL + G R+ ++
Sbjct: 106 PGEYRADGIFDLEPFFDAASEAGIYLLARPGPYINAESSGGGFPGWLQRVNG-TLRSSDK 164
Query: 147 PFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEY-GNVEGAYGPAAVPYINWAAS 205
+ + + + + + + + GGPIIL Q ENEY G P V Y+ +
Sbjct: 165 AYLDATDNYVSHVAATIAKYQI--TNGGPIILYQPENEYTSGCSGVEFPDPV-YMQYVED 221
Query: 206 MATSLDTGVPWV----MCQQENAPDPIINTCNGFYCDQFTPN-SKAKPKFWTEG-----Y 255
A + +P + NAP + + D + A P W G +
Sbjct: 222 QARNAGVVIPLINNDASASGNNAPGTGKGAVDIYGHDSYPLGFDCANPTVWPSGDLPTNF 281
Query: 256 NGWLLYFGGAVPYRPVEDLAFAVARFFQLGG---------------------------TL 288
L PY VE F + GG +
Sbjct: 282 RTLHLEQSPTTPYAIVE---FQGGSYDPWGGPGFAACSELLNNEFERVFYKNDFSFQIAI 338
Query: 289 QNYYMYHGGTNFGRTSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALI 348
N YM GGTN+G G P TSYDY +A+ E I + K+ LK + K+ L
Sbjct: 339 MNLYMIFGGTNWGNL-GYPNGYTSYDYGSAVTESRNITREKYSELKLLGNFAKVSPGYLT 397
Query: 349 ATDPTITSLG 358
A+ +T+ G
Sbjct: 398 ASPGNLTTSG 407
>BGAM_HUMAN (P16279) Beta-galactosidase-related protein precursor
(Beta-galactosidase-like protein) (S-Gal)
(Elastin-binding protein) (EBP)
Length = 546
Score = 51.6 bits (122), Expect = 9e-06
Identities = 41/131 (31%), Positives = 62/131 (47%), Gaps = 10/131 (7%)
Query: 230 NTCNGFYCDQFTPNSKAKPK---FWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLG- 285
N+ D F K +PK +E Y GWL ++G P+ ++ A A + + L
Sbjct: 111 NSAGSNITDAFLSQRKCEPKGPLINSEFYTGWLDHWGQ--PHSTIKTEAVASSLYDILAR 168
Query: 286 GTLQNYYMYHGGTNFGRTSGG--PFVA--TSYDYDAAIDEYGFIRQPKWGHLKDVHKAIK 341
G N YM+ GGTNF +G P+ A TSYDYDA + E G + + + + K K
Sbjct: 169 GASVNLYMFIGGTNFAYWNGANSPYAAQPTSYDYDAPLSEAGDLTEKYFALRNIIQKFEK 228
Query: 342 LCEEALIATDP 352
+ E + + P
Sbjct: 229 VPEGPIPPSTP 239
Score = 49.7 bits (117), Expect = 3e-05
Identities = 40/139 (28%), Positives = 62/139 (43%), Gaps = 13/139 (9%)
Query: 528 VEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGST---LDFSSK 584
+ + IT AG T+DLL +G ++G + + + ++ + + T LD
Sbjct: 330 ITLNITGKAGA-TLDLLVENMGRVNYGAYINDFKGLVSNLTLSSNILTDWTIFPLDTEDA 388
Query: 585 QWTYQIGLKGEDLGLPSGTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIA----IDFT 640
++ G D G +S TLP +Y NFS PSG + I F
Sbjct: 389 VRSHLGGWGHRDSGHHDEAWAHNSSNYTLP-----AFYMGNFSIPSGIPDLPQDTFIQFP 443
Query: 641 GMGKGEAWVNGQSIGRYWP 659
G KG+ W+NG ++GRYWP
Sbjct: 444 GWTKGQVWINGFNLGRYWP 462
Score = 40.8 bits (94), Expect = 0.015
Identities = 27/75 (36%), Positives = 36/75 (48%), Gaps = 7/75 (9%)
Query: 9 LLLLLWLLCVYAPACFCANVT-------YDHRALVIDGKRRVLISGSIHYPRSTPEMWPD 61
+LLLL +L + P N T Y + + DG+ ISGSIHY R W D
Sbjct: 8 ILLLLLVLLLLGPTRGLRNATQRMFEIDYSRDSFLKDGQPFRYISGSIHYSRVPRFYWKD 67
Query: 62 LIQKAKDGGLDVIET 76
+ K K GL+ I+T
Sbjct: 68 RLLKMKMAGLNAIQT 82
>LEG_ANTCR (P22031) D-galactoside-specific lectin (Sea urchin egg
lectin) (SUEL)
Length = 105
Score = 48.5 bits (114), Expect = 7e-05
Identities = 29/87 (33%), Positives = 43/87 (49%), Gaps = 8/87 (9%)
Query: 761 LSLECPYPNEVITTIKFASFGTPHGNC-----GNFSHG-DCSSKQALSIVQKACIGSSSC 814
L++ CP ++ I A +G G G F+ C S + +V+ +C G SSC
Sbjct: 19 LTISCPEGEGIV--IYDAIYGRKRGEVCPGLFGAFTKNRKCRSSNSQQVVENSCEGKSSC 76
Query: 815 SIGVSINTFGDPCGGVTKSLAVEASCT 841
++ S + FGDPC G K LAV C+
Sbjct: 77 TVLASNSVFGDPCPGTAKYLAVTYICS 103
>SAL_SILAS (Q9PVW8) Rhamnose-binding lectin precursor (SAL) (RBL)
(Roe lectin)
Length = 308
Score = 40.0 bits (92), Expect = 0.026
Identities = 22/66 (33%), Positives = 29/66 (43%), Gaps = 3/66 (4%)
Query: 776 KFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTFGDPCGGVTKSLA 835
+ S G P+G N + C + L+ V C G SC++ F DPC G K L
Sbjct: 151 QICSNGLPNGLTQNTN---CYAANTLTTVAGLCNGKKSCTVEALNTIFSDPCSGTVKYLT 207
Query: 836 VEASCT 841
V CT
Sbjct: 208 VTYICT 213
Score = 35.0 bits (79), Expect = 0.83
Identities = 16/47 (34%), Positives = 24/47 (51%)
Query: 794 DCSSKQALSIVQKACIGSSSCSIGVSINTFGDPCGGVTKSLAVEASC 840
+C + L+ V C S+C+I + N FGDPC K L + +C
Sbjct: 261 NCYTSDTLNKVAAGCDHLSTCTIPANNNFFGDPCPNTYKYLRIVYAC 307
>CU63_PANTR (Q68US5) Protein C21orf63 homolog precursor
Length = 441
Score = 40.0 bits (92), Expect = 0.026
Identities = 20/48 (41%), Positives = 26/48 (53%), Gaps = 1/48 (2%)
Query: 794 DCSSKQALSIVQKACIGSSSCSIGVSINTFGDPC-GGVTKSLAVEASC 840
DC S AL ++ + C G C I V+ + FG PC GV K L V +C
Sbjct: 212 DCLSYSALQVLSRRCYGKQRCKIIVNNHHFGSPCLPGVKKYLTVTYAC 259
>CU63_HUMAN (P58658) Protein C21orf63 precursor (Protein PRED34)
(SUE21) (UNQ2504/PRO5993)
Length = 441
Score = 40.0 bits (92), Expect = 0.026
Identities = 20/48 (41%), Positives = 26/48 (53%), Gaps = 1/48 (2%)
Query: 794 DCSSKQALSIVQKACIGSSSCSIGVSINTFGDPC-GGVTKSLAVEASC 840
DC S AL ++ + C G C I V+ + FG PC GV K L V +C
Sbjct: 212 DCLSYSALQVLSRRCYGKQRCKIIVNNHHFGSPCLPGVKKYLTVTYAC 259
>CU63_MOUSE (P58659) Protein C21orf63 homolog precursor
Length = 440
Score = 37.7 bits (86), Expect = 0.13
Identities = 19/48 (39%), Positives = 24/48 (49%), Gaps = 1/48 (2%)
Query: 794 DCSSKQALSIVQKACIGSSSCSIGVSINTFGDPC-GGVTKSLAVEASC 840
DC S AL ++ + C G C + V FG PC GV K L V +C
Sbjct: 212 DCLSYTALQVLSRRCYGKQRCKVLVDNYHFGSPCLPGVKKYLTVAYAC 259
>LHN2_RAT (O88923) Latrophilin 2 precursor (Calcium-independent
alpha-latrotoxin receptor 2) (CIRL-2)
Length = 1487
Score = 35.8 bits (81), Expect = 0.48
Identities = 24/88 (27%), Positives = 38/88 (42%), Gaps = 7/88 (7%)
Query: 758 GPVLSLECPYPNEVITTIKFASFG-TPHGNCG----NFSHGDCSSKQALSIVQKACIGSS 812
G + L CP + ++ I+ A++G T C + DC A I+ + C +
Sbjct: 44 GYSIDLRCPGSDVIM--IESANYGRTDDKICDADPFQMENTDCYLPDAFKIMTQRCNNRT 101
Query: 813 SCSIGVSINTFGDPCGGVTKSLAVEASC 840
C + + F DPC G K L V+ C
Sbjct: 102 QCVVVTGSDVFPDPCPGTYKYLEVQYEC 129
>LHN2_HUMAN (O95490) Latrophilin 2 precursor (Calcium-independent
alpha-latrotoxin receptor 2) (Latrophilin homolog 1)
(Lectomedin 1)
Length = 1459
Score = 35.8 bits (81), Expect = 0.48
Identities = 24/88 (27%), Positives = 38/88 (42%), Gaps = 7/88 (7%)
Query: 758 GPVLSLECPYPNEVITTIKFASFG-TPHGNCG----NFSHGDCSSKQALSIVQKACIGSS 812
G + L CP + ++ I+ A++G T C + DC A I+ + C +
Sbjct: 44 GYSIDLRCPGSDVIM--IESANYGRTDDKICDADPFQMENTDCYLPDAFKIMTQRCNNRT 101
Query: 813 SCSIGVSINTFGDPCGGVTKSLAVEASC 840
C + + F DPC G K L V+ C
Sbjct: 102 QCIVVTGSDVFPDPCPGTYKYLEVQYEC 129
>LHN2_BOVIN (O97817) Latrophilin 2 precursor (Calcium-independent
alpha-latrotoxin receptor 2)
Length = 1478
Score = 35.8 bits (81), Expect = 0.48
Identities = 24/88 (27%), Positives = 38/88 (42%), Gaps = 7/88 (7%)
Query: 758 GPVLSLECPYPNEVITTIKFASFG-TPHGNCG----NFSHGDCSSKQALSIVQKACIGSS 812
G + L CP + ++ I+ A++G T C + DC A I+ + C +
Sbjct: 44 GYSIDLRCPGSDVIM--IESANYGRTDDKICDADPFQMENTDCYLPDAFKIMTQRCNNRT 101
Query: 813 SCSIGVSINTFGDPCGGVTKSLAVEASC 840
C + + F DPC G K L V+ C
Sbjct: 102 QCIVVTGSDVFPDPCPGTYKYLEVQYEC 129
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.136 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 110,903,983
Number of Sequences: 164201
Number of extensions: 5216861
Number of successful extensions: 10603
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 10496
Number of HSP's gapped (non-prelim): 58
length of query: 841
length of database: 59,974,054
effective HSP length: 119
effective length of query: 722
effective length of database: 40,434,135
effective search space: 29193445470
effective search space used: 29193445470
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0136.7


