

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0136.17
(83 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
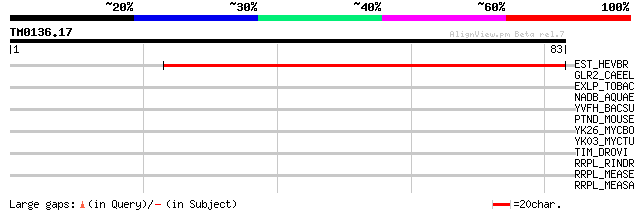
EST_HEVBR (Q7Y1X1) Esterase precursor (EC 3.1.1.-) (Early nodule... 89 3e-18
GLR2_CAEEL (Q10914) Glutamate receptor 2 precursor 29 2.5
EXLP_TOBAC (Q03211) Pistil-specific extensin-like protein precur... 29 2.5
NADB_AQUAE (O66973) L-aspartate oxidase (EC 1.4.3.16) (LASPO) (Q... 28 3.2
YVFH_BACSU (P71067) Putative L-lactate permease yvfH 27 7.1
PTND_MOUSE (Q64512) Protein tyrosine phosphatase, non-receptor t... 27 7.1
YK26_MYCBO (P64920) Hypothetical protein Mb2026c 27 9.3
YK03_MYCTU (P64919) Hypothetical protein Rv2003c/MT2059 27 9.3
TIM_DROVI (O17482) Timeless protein 27 9.3
RRPL_RINDR (P41357) RNA polymerase beta subunit (EC 2.7.7.48) (L... 27 9.3
RRPL_MEASE (P12576) RNA polymerase beta subunit (EC 2.7.7.48) (L... 27 9.3
RRPL_MEASA (P35975) RNA polymerase beta subunit (EC 2.7.7.48) (L... 27 9.3
>EST_HEVBR (Q7Y1X1) Esterase precursor (EC 3.1.1.-) (Early
nodule-specific protein homolog) (Latex allergen Hev b
13)
Length = 391
Score = 88.6 bits (218), Expect = 3e-18
Identities = 39/60 (65%), Positives = 47/60 (78%)
Query: 24 LNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSDGRIILDF 83
L+ +A + C+FPAIFN G SNSDTGG AAAF P PYG+T+FHR GRYSDGR+I+DF
Sbjct: 21 LSLAYASETCDFPAIFNFGDSNSDTGGKAAAFYPLNPPYGETFFHRSTGRYSDGRLIIDF 80
>GLR2_CAEEL (Q10914) Glutamate receptor 2 precursor
Length = 977
Score = 28.9 bits (63), Expect = 2.5
Identities = 15/47 (31%), Positives = 23/47 (48%)
Query: 9 HVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAF 55
H S +++ + N F K R F ++ G SN+D+ AAAF
Sbjct: 286 HFSLINITIFRIFDKNNKKFLKTRAEFHDVYRGGFSNTDSIPTAAAF 332
>EXLP_TOBAC (Q03211) Pistil-specific extensin-like protein precursor
(PELP)
Length = 426
Score = 28.9 bits (63), Expect = 2.5
Identities = 15/29 (51%), Positives = 15/29 (51%), Gaps = 4/29 (13%)
Query: 33 CNFPAIFNLGASNSDTGGLAAAFLPPKSP 61
CN P FN G S GGL LPPK P
Sbjct: 368 CNVPTNFNGGKS----GGLLKPLLPPKQP 392
>NADB_AQUAE (O66973) L-aspartate oxidase (EC 1.4.3.16) (LASPO)
(Quinolinate synthetase B)
Length = 510
Score = 28.5 bits (62), Expect = 3.2
Identities = 18/41 (43%), Positives = 20/41 (47%), Gaps = 2/41 (4%)
Query: 36 PAIFNLGASNS--DTGGLAAAFLPPKSPYGDTYFHRPAGRY 74
P I G N+ GG+AAA LP SPY AGRY
Sbjct: 42 PLILTRGIGNTYYSQGGIAAAVLPDDSPYLHYCDTLRAGRY 82
>YVFH_BACSU (P71067) Putative L-lactate permease yvfH
Length = 563
Score = 27.3 bits (59), Expect = 7.1
Identities = 17/54 (31%), Positives = 28/54 (51%)
Query: 8 RHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSP 61
R + F+S+F+ L+ + F K +PAI G S + L++ FL P+ P
Sbjct: 197 RQLPFLSVFIPLYLIIIMSGFRKALEIWPAILVSGVSFAVVQYLSSNFLGPELP 250
>PTND_MOUSE (Q64512) Protein tyrosine phosphatase, non-receptor type
13 (EC 3.1.3.48) (Protein tyrosine phosphatase PTP-BL)
(Protein-tyrosine phosphatase RIP) (protein tyrosine
phosphatase DPZPTP) (PTP36)
Length = 2453
Score = 27.3 bits (59), Expect = 7.1
Identities = 23/76 (30%), Positives = 34/76 (44%), Gaps = 4/76 (5%)
Query: 11 SFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAF--LPPKSPYGDTYF- 67
+F S + +T +KK+ ++ L A+N D G + PK+ G T
Sbjct: 272 TFSSPYQFKTSTPQMDALSKKKTWASSMDLLCAANRDISGETGRYQRCDPKTVTGRTSIT 331
Query: 68 -HRPAGRYSDGRIILD 82
+ GRYSDG I LD
Sbjct: 332 PRKKEGRYSDGSIALD 347
>YK26_MYCBO (P64920) Hypothetical protein Mb2026c
Length = 285
Score = 26.9 bits (58), Expect = 9.3
Identities = 12/26 (46%), Positives = 15/26 (57%)
Query: 46 SDTGGLAAAFLPPKSPYGDTYFHRPA 71
+D GGL FLP +P+ D Y R A
Sbjct: 180 ADGGGLVIGFLPRGTPWADLYALRAA 205
>YK03_MYCTU (P64919) Hypothetical protein Rv2003c/MT2059
Length = 285
Score = 26.9 bits (58), Expect = 9.3
Identities = 12/26 (46%), Positives = 15/26 (57%)
Query: 46 SDTGGLAAAFLPPKSPYGDTYFHRPA 71
+D GGL FLP +P+ D Y R A
Sbjct: 180 ADGGGLVIGFLPRGTPWADLYALRAA 205
>TIM_DROVI (O17482) Timeless protein
Length = 1343
Score = 26.9 bits (58), Expect = 9.3
Identities = 15/45 (33%), Positives = 24/45 (53%)
Query: 21 TTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDT 65
T+TL + C+F A N +S+S GG A++ P P G++
Sbjct: 1227 TSTLVCMNKLSNCSFAAPPNENSSSSGCGGTASSMSMPNMPDGNS 1271
>RRPL_RINDR (P41357) RNA polymerase beta subunit (EC 2.7.7.48) (Large
structural protein) (L protein)
Length = 2183
Score = 26.9 bits (58), Expect = 9.3
Identities = 8/20 (40%), Positives = 14/20 (70%)
Query: 64 DTYFHRPAGRYSDGRIILDF 83
D ++HRP+G+Y G ++ F
Sbjct: 1498 DIHYHRPSGKYQMGELLTSF 1517
>RRPL_MEASE (P12576) RNA polymerase beta subunit (EC 2.7.7.48) (Large
structural protein) (L protein)
Length = 2183
Score = 26.9 bits (58), Expect = 9.3
Identities = 8/20 (40%), Positives = 14/20 (70%)
Query: 64 DTYFHRPAGRYSDGRIILDF 83
D ++HRP+G+Y G ++ F
Sbjct: 1498 DVHYHRPSGKYQMGELLSSF 1517
>RRPL_MEASA (P35975) RNA polymerase beta subunit (EC 2.7.7.48) (Large
structural protein) (L protein)
Length = 2183
Score = 26.9 bits (58), Expect = 9.3
Identities = 8/20 (40%), Positives = 14/20 (70%)
Query: 64 DTYFHRPAGRYSDGRIILDF 83
D ++HRP+G+Y G ++ F
Sbjct: 1498 DVHYHRPSGKYQMGELLSSF 1517
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.328 0.143 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,296,441
Number of Sequences: 164201
Number of extensions: 381748
Number of successful extensions: 768
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 759
Number of HSP's gapped (non-prelim): 12
length of query: 83
length of database: 59,974,054
effective HSP length: 59
effective length of query: 24
effective length of database: 50,286,195
effective search space: 1206868680
effective search space used: 1206868680
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0136.17


