

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0136.12
(245 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
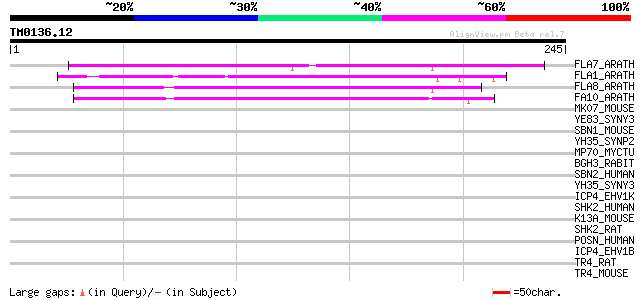
Score E
Sequences producing significant alignments: (bits) Value
FLA7_ARATH (Q9SJ81) Fasciclin-like arabinogalactan protein 7 pre... 117 3e-26
FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan protein 1 pre... 91 2e-18
FLA8_ARATH (O22126) Fasciclin-like arabinogalactan protein 8 pre... 87 5e-17
FA10_ARATH (Q9LZX4) Fasciclin-like arabinogalactan protein 10 pr... 80 4e-15
MK07_MOUSE (Q9WVS8) Mitogen-activated protein kinase 7 (EC 2.7.1... 41 0.002
YE83_SYNY3 (P74615) Hypothetical protein sll1483 precursor 40 0.007
SBN1_MOUSE (Q8R4Y4) Stabilin 1 precursor (FEEL-1 protein) 38 0.020
YH35_SYNP2 (O33752) Hypothetical protein sll1735 homolog 38 0.026
MP70_MYCTU (Q50769) Immunogenic protein MPT70 precursor 38 0.026
BGH3_RABIT (Q95215) Transforming growth factor-beta induced prot... 37 0.033
SBN2_HUMAN (Q8WWQ8) Stabilin 2 precursor (FEEL-2 protein) (Fasci... 35 0.22
YH35_SYNY3 (P73392) Hypothetical protein sll1735 34 0.28
ICP4_EHV1K (P17473) Trans-acting transcriptional protein ICP4 (1... 34 0.37
SHK2_HUMAN (Q9UPX8) SH3 and multiple ankyrin repeat domains prot... 33 0.63
K13A_MOUSE (Q9EQW7) Kinesin-like protein KIF13A 33 0.63
SHK2_RAT (Q9QX74) SH3 and multiple ankyrin repeat domains protei... 33 0.82
POSN_HUMAN (Q15063) Periostin precursor (PN) (Osteoblast specifi... 32 1.4
ICP4_EHV1B (P28925) Trans-acting transcriptional protein ICP4 (1... 32 1.4
TR4_RAT (P55094) Orphan nuclear receptor TR4 31 2.4
TR4_MOUSE (P49117) Orphan nuclear receptor TR4 (Orphan nuclear r... 31 2.4
>FLA7_ARATH (Q9SJ81) Fasciclin-like arabinogalactan protein 7
precursor
Length = 254
Score = 117 bits (293), Expect = 3e-26
Identities = 75/215 (34%), Positives = 112/215 (51%), Gaps = 8/215 (3%)
Query: 27 SPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAF 86
+PAPAP+ ++ +L AG + T + L +T V Q N+ G+T F P D+AF
Sbjct: 35 TPAPAPAPENVNLTELLSVAGPFHTFLDYLLSTGVIETFQNQANNTEEGITIFVPKDDAF 94
Query: 87 SNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTL--SNPVRTQAGENPDRLALNVTS 144
K L++L Q +L+ FH LP + S+S F L S PV T AG + +L T
Sbjct: 95 KAQKNPPLSNLTKDQLKQLVLFHALPHYYSLSEFKNLSQSGPVSTFAG---GQYSLKFTD 151
Query: 145 SGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFF---VAKPPAPAPAPEKAKA 201
TV + + V SV+S + +AVYQV++VLLP F V PAPAPAP +
Sbjct: 152 VSGTVRIDSLWTRTKVSSSVFSTDPVAVYQVNRVLLPEAIFGTDVPPMPAPAPAPIVSAP 211
Query: 202 SKKKSADDTEGPAGDDSAAVSVKQRLVWGPLAVAM 236
S S D+EG + S+ + Q+L+ P+++ +
Sbjct: 212 SDSPSVADSEGASSPKSSHKNSGQKLLLAPISMVI 246
>FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan protein 1
precursor
Length = 424
Score = 91.3 bits (225), Expect = 2e-18
Identities = 66/215 (30%), Positives = 101/215 (46%), Gaps = 25/215 (11%)
Query: 22 TLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAP 81
T AA +PAPA + + + A G L T A++ + + G+T F P
Sbjct: 174 TAAAPTPAPAEMN-----LTGIMSAHGCKVFAETLLTNPGASKTYQESLEG--GMTVFCP 226
Query: 82 NDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALN 141
D+A P + N L +K + F +PT+ SM+ + + P+ T A + ++ L
Sbjct: 227 GDDAMKGFLPKYKN-LTAPKKEAFLDFLAVPTYYSMAMLKSNNGPMNTLATDGANKFELT 285
Query: 142 VTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVA---KPPAPAPAPE- 197
V + G V + T I V + ++ + LA+Y DKVLLP++ F A + PAPAPAPE
Sbjct: 286 VQNDGEKVTLKTRINTVKIVDTLIDEQPLAIYATDKVLLPKELFKASAVEAPAPAPAPED 345
Query: 198 ------------KAKASKKKSADDTEG-PAGDDSA 219
KAK KKK+A + P GD +
Sbjct: 346 GDVADSPKAAKGKAKGKKKKAAPSPDNDPFGDSDS 380
>FLA8_ARATH (O22126) Fasciclin-like arabinogalactan protein 8
precursor (AtAGP8)
Length = 420
Score = 86.7 bits (213), Expect = 5e-17
Identities = 58/181 (32%), Positives = 89/181 (49%), Gaps = 5/181 (2%)
Query: 29 APAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSN 88
APAPS+S ++I +L+KAG T L+ + + T +A GLT FAP+D AF
Sbjct: 180 APAPSASLSNITGLLEKAGCKTFANLLVSSGVLKTYESAV----EKGLTVFAPSDEAFKA 235
Query: 89 LKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGNT 148
L L + L+++H L + + T N + T A + L ++SG+
Sbjct: 236 EGVPDLTKLTQAEVVSLLEYHALAEYKPKGSLKTNKNNISTLATNGAGKFDLTTSTSGDE 295
Query: 149 VNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFF-VAKPPAPAPAPEKAKASKKKSA 207
V + TG+ + +V + ++ VD VLLP + F +K P+PAPAPE A A
Sbjct: 296 VILHTGVAPSRLADTVLDATPVVIFTVDNVLLPAELFGKSKSPSPAPAPEPVTAPTPSPA 355
Query: 208 D 208
D
Sbjct: 356 D 356
>FA10_ARATH (Q9LZX4) Fasciclin-like arabinogalactan protein 10
precursor
Length = 422
Score = 80.1 bits (196), Expect = 4e-15
Identities = 56/189 (29%), Positives = 88/189 (45%), Gaps = 7/189 (3%)
Query: 29 APAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSN 88
APAPSS+ I L + G T LL ++ V + + GLT FAP+D AF
Sbjct: 180 APAPSSAGVSNITGLLEKAGCKTFANLLVSSGVIKTFESTV---EKGLTVFAPSDEAFKA 236
Query: 89 LKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGNT 148
L +L + L+++H L + + T + + T A + L ++SG+
Sbjct: 237 RGVPDLTNLTQAEVVSLLEYHALAEYKPKGSLKTNKDAISTLATNGAGKYDLTTSTSGDE 296
Query: 149 VNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPAPAPAPEKAKA---SKKK 205
V + TG+ + +V + + ++ VD VLLP + F K +PAPAPE A + K
Sbjct: 297 VILHTGVGPSRLADTVVDETPVVIFTVDNVLLPAELF-GKSSSPAPAPEPVSAPTPTPAK 355
Query: 206 SADDTEGPA 214
S E P+
Sbjct: 356 SPSPVEAPS 364
>MK07_MOUSE (Q9WVS8) Mitogen-activated protein kinase 7 (EC
2.7.1.37) (Extracellular signal-regulated kinase 5)
(ERK-5) (BMK1 kinase)
Length = 806
Score = 41.2 bits (95), Expect = 0.002
Identities = 27/66 (40%), Positives = 32/66 (47%), Gaps = 7/66 (10%)
Query: 160 VGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPAPAPAPEKAKASKKKSADDTEGPAGDDSA 219
+ G V SDN D+ LL R +A+PPAPAPAP A A SA T P G S
Sbjct: 556 LAGLVLSDN-------DRSLLERWTRMARPPAPAPAPAPAPAPAPSSAQPTSTPTGPVSQ 608
Query: 220 AVSVKQ 225
+ Q
Sbjct: 609 STGPLQ 614
>YE83_SYNY3 (P74615) Hypothetical protein sll1483 precursor
Length = 180
Score = 39.7 bits (91), Expect = 0.007
Identities = 39/154 (25%), Positives = 72/154 (46%), Gaps = 18/154 (11%)
Query: 33 SSSPTD-IIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSNLKP 91
S + TD + I++ A G T L+ + A + A +++ T FAP ++AF+ L
Sbjct: 38 SQAATDSAMTIVEVAAGNETFSTLVAAVKAADLVEA--LSAEGPFTVFAPTNDAFAALPA 95
Query: 92 GFLNSL----NDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGN 147
G + SL N + +++ +H++P ++ + S V + AGE L V
Sbjct: 96 GTVESLLLPENKDKLVKILTYHVVPGKITAAQVQ--SGEVASLAGE---ALTFKVKDGKV 150
Query: 148 TVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLP 181
VN T + V + N + ++ +D+V+LP
Sbjct: 151 KVNKAT-----VISADVDASNGV-IHVIDQVILP 178
>SBN1_MOUSE (Q8R4Y4) Stabilin 1 precursor (FEEL-1 protein)
Length = 2571
Score = 38.1 bits (87), Expect = 0.020
Identities = 34/160 (21%), Positives = 67/160 (41%), Gaps = 25/160 (15%)
Query: 17 FHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGL 76
FH+ T L P P P S + +IL +T +L+ + + +++
Sbjct: 488 FHTVTALRWQLPPPLPGDSKKTVGQILASTEVFTRFETILENCGLPS-----ILDGPGPF 542
Query: 77 TFFAPNDNAFSNLKPG---FLNSLNDQQKNELIQFHL-------LPTFVSMSNFDTLSNP 126
T FAP++ A +L+ G +L + + EL+++H+ + +S T++N
Sbjct: 543 TVFAPSNEAVDSLRDGRLIYLFTAGLSKLQELVRYHIYNHGQLTVEKLISKGRVLTMANQ 602
Query: 127 VRTQAGENPDRLAL----------NVTSSGNTVNMTTGIV 156
V T R+ L +V ++ ++M GI+
Sbjct: 603 VLTVNISEGGRILLGPGGIPVRRVDVPAANGVIHMLEGIL 642
>YH35_SYNP2 (O33752) Hypothetical protein sll1735 homolog
Length = 133
Score = 37.7 bits (86), Expect = 0.026
Identities = 33/130 (25%), Positives = 55/130 (41%), Gaps = 18/130 (13%)
Query: 39 IIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAG---LTFFAPNDNAFSNLKPGFLN 95
I+ I G++TL+ +Q A L+ + AG T FAPND+AF+ L PG +
Sbjct: 4 IVDIAVNNEGFSTLVAAVQA--------AGLVETLAGEGDFTVFAPNDDAFAKLPPGTIQ 55
Query: 96 SL--NDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENP-----DRLALNVTSSGNT 148
+L N Q ++ +H+ ++ + L + P D N T
Sbjct: 56 TLVQNPPQLARILTYHVAAGRLTKDDLIKLGEVTSIEGSPIPINGEDDFEVKNATVLAAD 115
Query: 149 VNMTTGIVNV 158
+ GI++V
Sbjct: 116 IEADNGIIHV 125
>MP70_MYCTU (Q50769) Immunogenic protein MPT70 precursor
Length = 193
Score = 37.7 bits (86), Expect = 0.026
Identities = 35/134 (26%), Positives = 56/134 (41%), Gaps = 15/134 (11%)
Query: 50 TTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSL--NDQQKNELIQ 107
TTL L + Q+ Q+N ++ T FAP + AFS L ++ L N ++
Sbjct: 71 TTLTAAL-SGQLNPQVNLVDTLNSGQYTVFAPTNAAFSKLPASTIDELKTNSSLLTSILT 129
Query: 108 FHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSD 167
+H++ S +N G +VT +G ++ G +V GG S
Sbjct: 130 YHVVAGQTSPANV----------VGTRQTLQGASVTVTGQGNSLKVGNADVVCGG--VST 177
Query: 168 NQLAVYQVDKVLLP 181
VY +D VL+P
Sbjct: 178 ANATVYMIDSVLMP 191
>BGH3_RABIT (Q95215) Transforming growth factor-beta induced protein
IG-H3 precursor (Beta IG-H3) (Kerato-epithelin)
(RGD-containing collagen associated protein) (RGD-CAP)
Length = 683
Score = 37.4 bits (85), Expect = 0.033
Identities = 34/145 (23%), Positives = 60/145 (40%), Gaps = 17/145 (11%)
Query: 39 IIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSL- 97
++ +LK ++ L+ +Q ++ + +N T FAP + AF L PG LN L
Sbjct: 505 VMDVLKGDNRFSMLVAAIQFRRLT-----ETLNREGAYTVFAPTNEAFQALPPGELNKLL 559
Query: 98 -NDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGNTVNMTTGIV 156
N ++ +++++H+ + TL Q + L V+S N V++
Sbjct: 560 GNAKELADILKYHVGEEILVSGGIGTLVRLKSLQGDK------LEVSSKNNAVSVN---- 609
Query: 157 NVTVGGSVYSDNQLAVYQVDKVLLP 181
V S VY + VL P
Sbjct: 610 KEPVAESDIMATNGVVYAITSVLQP 634
>SBN2_HUMAN (Q8WWQ8) Stabilin 2 precursor (FEEL-2 protein)
(Fasciclin EGF-like laminin-type EGF-like and link
domain-containing scavenger receptor-1) (FAS1 EGF-like
and X-link domain containing adhesion molecule-2)
(Hyaluronan receptor for endocytosis) [C
Length = 2551
Score = 34.7 bits (78), Expect = 0.22
Identities = 24/98 (24%), Positives = 48/98 (48%), Gaps = 12/98 (12%)
Query: 77 TFFAPNDNAFSNLKPGFLNSL----NDQQKNELIQFHLLPTFVSMSNFDTLSNP-VRTQA 131
T F PN+ A +N+K G L+ L ++ EL+++H++P F + +S P +R+ A
Sbjct: 554 TIFVPNNEALNNMKDGTLDYLLSPEGSRKLLELVRYHIVP-FTQLEVATLISTPHIRSMA 612
Query: 132 GENPDRLALNVTSSGNTVNMTTGIVNVTV---GGSVYS 166
+ + N T +G + + + + G +Y+
Sbjct: 613 NQ---LIQFNTTDNGQILANDVAMEEIEITAKNGRIYT 647
>YH35_SYNY3 (P73392) Hypothetical protein sll1735
Length = 133
Score = 34.3 bits (77), Expect = 0.28
Identities = 32/145 (22%), Positives = 60/145 (41%), Gaps = 18/145 (12%)
Query: 38 DIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSL 97
+I+ I ++TL+ T V ++ S T FAP D AF+ L PG + +L
Sbjct: 3 NIVEIAVSDERFSTLV-----TAVTAANLVDVLQSPGPFTVFAPTDTAFAKLPPGTITTL 57
Query: 98 --NDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGNTVNMTTGI 155
N Q ++ +H++ + ++ LS T +A++ T N T I
Sbjct: 58 VQNIPQLARILTYHVVAGKFTQADLCRLS----TVDSVEGSPIAIDCTEGFEVKNATVII 113
Query: 156 VNVTVGGSVYSDNQLAVYQVDKVLL 180
++ + ++ +D V+L
Sbjct: 114 PDIEADNGI-------IHVIDNVIL 131
>ICP4_EHV1K (P17473) Trans-acting transcriptional protein ICP4 (155
kDa immediate-early protein)
Length = 1487
Score = 33.9 bits (76), Expect = 0.37
Identities = 18/49 (36%), Positives = 24/49 (48%)
Query: 172 VYQVDKVLLPRDFFVAKPPAPAPAPEKAKASKKKSADDTEGPAGDDSAA 220
++ VD L V PP+PAP P KA + SA + GP +AA
Sbjct: 52 MFGVDDAPLSTPVVVIPPPSPAPEPRGGKAKRSPSAAGSGGPPTPAAAA 100
>SHK2_HUMAN (Q9UPX8) SH3 and multiple ankyrin repeat domains protein 2
(Shank2)
Length = 1253
Score = 33.1 bits (74), Expect = 0.63
Identities = 36/162 (22%), Positives = 61/162 (37%), Gaps = 9/162 (5%)
Query: 72 SNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQA 131
SN F P +F + + ++ + ++ H L T ++S ++S + ++
Sbjct: 873 SNCLPASFLPPPESFDAVADSGIEEVDSRSSSD----HHLETTSTISTVSSIST-LSSEG 927
Query: 132 GENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPA 191
GEN D + V+ + ++ N A+YQ V D FV PPA
Sbjct: 928 GENVDTCTVYADGQAFMVDKPPVPPKPKMKPIIHKSN--ALYQDALVEEDVDSFVIPPPA 985
Query: 192 PAPAPEKAKASKKKSADDTEGPAGDDSAAVSVKQRLVWGPLA 233
P P P A+ K D +K ++ GP A
Sbjct: 986 PPPPPGSAQPGMAKVLQPRTSKLWGD--VTEIKSPILSGPKA 1025
>K13A_MOUSE (Q9EQW7) Kinesin-like protein KIF13A
Length = 1749
Score = 33.1 bits (74), Expect = 0.63
Identities = 22/88 (25%), Positives = 45/88 (51%), Gaps = 6/88 (6%)
Query: 142 VTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDK----VLLPRDFFVAKPPAPAPAPE 197
V+ S T ++T+G + + + SD +AV D + ++ A+PP P+ AP+
Sbjct: 1538 VSRSPTTSSLTSGYFSHSASNATLSD--MAVPSSDSSDQLAVSTKEVECAEPPGPSLAPD 1595
Query: 198 KAKASKKKSADDTEGPAGDDSAAVSVKQ 225
+AS ++ + G D++ AV +++
Sbjct: 1596 VRRASNQELTEVGRGSGKDETIAVPLEE 1623
>SHK2_RAT (Q9QX74) SH3 and multiple ankyrin repeat domains protein 2
(Shank2) (Proline-rich synapse associated protein 1)
(ProSAP1) (Cortactin-binding protein 1) (CortBP1)
(GKAP/SAPAP interacting protein) (SPANK-3)
Length = 1474
Score = 32.7 bits (73), Expect = 0.82
Identities = 37/162 (22%), Positives = 62/162 (37%), Gaps = 9/162 (5%)
Query: 72 SNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQA 131
SN + F P +F + + ++ + ++ H L T ++S ++S + ++
Sbjct: 1094 SNCLPSSFLPPPESFDAVTDSGIEEVDSRSSSD----HHLETTSTISTVSSIST-LSSEG 1148
Query: 132 GENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPA 191
GE+ D + V+ + V+ N A+YQ D FV PPA
Sbjct: 1149 GESMDTCTVYADGQAFVVDKPPVPPKPKMKPIVHKSN--ALYQDTLPEEDTDGFVIPPPA 1206
Query: 192 PAPAPEKAKASKKKSADDTEGPAGDDSAAVSVKQRLVWGPLA 233
P P P A+A K D VK ++ GP A
Sbjct: 1207 PPPPPGSAQAGVAKVIQPRTSKLWGD--VTEVKSPILSGPKA 1246
>POSN_HUMAN (Q15063) Periostin precursor (PN) (Osteoblast specific
factor 2) (OSF-2)
Length = 836
Score = 32.0 bits (71), Expect = 1.4
Identities = 30/122 (24%), Positives = 53/122 (42%), Gaps = 23/122 (18%)
Query: 68 QLINSNAGLTFFAPNDNAFSNLKPGFLNS-LNDQQKNE-LIQFHLLPTFVSMSNFDTLSN 125
+ + + T FAP + AF L G L + D+ +E L+++H+L T
Sbjct: 261 EALGRDGHFTLFAPTNEAFEKLPRGVLERFMGDKVASEALMKYHILNTL----------- 309
Query: 126 PVRTQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSD------NQLAVYQVDKVL 179
Q E+ A+ T GNT+ + ++TV G + N ++ +D+VL
Sbjct: 310 ----QCSESIMGGAVFETLEGNTIEIGCDGDSITVNGIKMVNKKDIVTNNGVIHLIDQVL 365
Query: 180 LP 181
+P
Sbjct: 366 IP 367
>ICP4_EHV1B (P28925) Trans-acting transcriptional protein ICP4 (155
kDa immediate-early protein)
Length = 1487
Score = 32.0 bits (71), Expect = 1.4
Identities = 17/49 (34%), Positives = 23/49 (46%)
Query: 172 VYQVDKVLLPRDFFVAKPPAPAPAPEKAKASKKKSADDTEGPAGDDSAA 220
++ VD L V PP+P P P KA + SA + GP +AA
Sbjct: 52 MFGVDDAPLSTPAVVIPPPSPTPEPRGGKAKRSPSAAGSGGPPTPAAAA 100
>TR4_RAT (P55094) Orphan nuclear receptor TR4
Length = 596
Score = 31.2 bits (69), Expect = 2.4
Identities = 36/144 (25%), Positives = 58/144 (40%), Gaps = 35/144 (24%)
Query: 57 QTTQVATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVS 116
+T+Q A A ++ S A L+ N +A S ++P DQ +E+ +
Sbjct: 268 ETSQGALGTLANVVTSLANLSESLNNGDA-SEMQP------EDQSASEITRA-------- 312
Query: 117 MSNFDTLSNPVRTQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVY---SDNQLAVY 173
FDTL+ + T +P LA + +SG GGS++ D +
Sbjct: 313 ---FDTLAKALNTTDSASPPSLADGIDASG--------------GGSIHVISRDQSTPII 355
Query: 174 QVDKVLLPRDFFVAKPPAPAPAPE 197
+V+ LL K P+P PE
Sbjct: 356 EVEGPLLSDTHVTFKLTMPSPMPE 379
>TR4_MOUSE (P49117) Orphan nuclear receptor TR4 (Orphan nuclear
receptor TAK1)
Length = 596
Score = 31.2 bits (69), Expect = 2.4
Identities = 36/144 (25%), Positives = 58/144 (40%), Gaps = 35/144 (24%)
Query: 57 QTTQVATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVS 116
+T+Q A A ++ S A L+ N +A S ++P DQ +E+ +
Sbjct: 268 ETSQGALGTLANVVTSLANLSESLNNGDA-SEMQP------EDQSASEITRA-------- 312
Query: 117 MSNFDTLSNPVRTQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVY---SDNQLAVY 173
FDTL+ + T +P LA + +SG GGS++ D +
Sbjct: 313 ---FDTLAKALNTTDSASPPSLADGIDASG--------------GGSIHVISRDQSTPII 355
Query: 174 QVDKVLLPRDFFVAKPPAPAPAPE 197
+V+ LL K P+P PE
Sbjct: 356 EVEGPLLSDTHVTFKLTMPSPMPE 379
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.130 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,479,313
Number of Sequences: 164201
Number of extensions: 1133276
Number of successful extensions: 6462
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 6399
Number of HSP's gapped (non-prelim): 73
length of query: 245
length of database: 59,974,054
effective HSP length: 107
effective length of query: 138
effective length of database: 42,404,547
effective search space: 5851827486
effective search space used: 5851827486
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0136.12


