

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0134.9
(1610 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
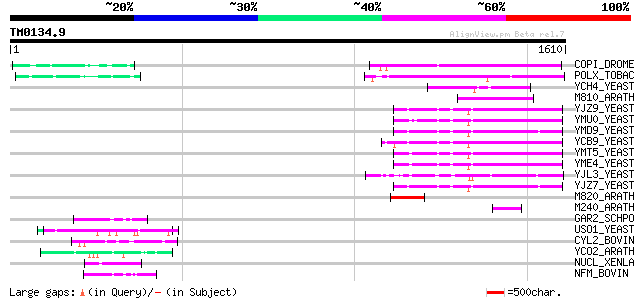
Score E
Sequences producing significant alignments: (bits) Value
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 387 e-106
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 342 6e-93
YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein 161 2e-38
M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810... 154 1e-36
YJZ9_YEAST (P47100) Transposon Ty1 protein B 117 2e-25
YMU0_YEAST (Q04670) Transposon Ty1 protein B 115 1e-24
YMD9_YEAST (Q03434) Transposon Ty1 protein B 115 1e-24
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 115 1e-24
YMT5_YEAST (Q04214) Transposon Ty1 protein B 114 2e-24
YME4_YEAST (Q04711) Transposon Ty1 protein B 114 2e-24
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 114 2e-24
YJZ7_YEAST (P47098) Transposon Ty1 protein B 113 5e-24
M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820... 100 2e-20
M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240... 66 7e-10
GAR2_SCHPO (P41891) Protein gar2 52 1e-05
USO1_YEAST (P25386) Intracellular protein transport protein USO1 52 1e-05
CYL2_BOVIN (Q28092) Cylicin II (Multiple-band polypeptide II) 51 3e-05
YCO2_ARATH (Q9LME2) Hypothetical protein At1g22260 50 4e-05
NUCL_XENLA (P20397) Nucleolin (Protein C23) 50 4e-05
NFM_BOVIN (O77788) Neurofilament triplet M protein (160 kDa neur... 50 4e-05
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 387 bits (995), Expect = e-106
Identities = 220/584 (37%), Positives = 340/584 (57%), Gaps = 33/584 (5%)
Query: 1044 EDEPEEEAGPSNSQTLKKSRITAAHPKEL------------ILGNKDEPVRTRSAFRPSE 1091
+D E G N ++S TA H KE+ I+ + E ++T+ +E
Sbjct: 813 DDHLNESKGSGNPNESRESE-TAEHLKEIGIDNPTKNDGIEIINRRSERLKTKPQISYNE 871
Query: 1092 E------TLLSLKGLVSLIEPKSIDEALQDKD---WILAMEEELNQFSKNDVWSLVKKPE 1142
E +L+ + + + P S DE D W A+ ELN N+ W++ K+PE
Sbjct: 872 EDNSLNKVVLNAHTIFNDV-PNSFDEIQYRDDKSSWEEAINTELNAHKINNTWTITKRPE 930
Query: 1143 NVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISF 1202
N +++ ++WVF K NE G+ +R KARLVA+G++Q+ IDY ETFAPVAR+ + R ++S
Sbjct: 931 NKNIVDSRWVFSVKYNELGNPIRYKARLVARGFTQKYQIDYEETFAPVARISSFRFILSL 990
Query: 1203 SVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAW 1262
+ +N+ +HQMDVK+AFLNG + EE+Y+ P G D+V KL K++YGLKQA R W
Sbjct: 991 VIQYNLKVHQMDVKTAFLNGTLKEEIYMRLPQGISCNS--DNVCKLNKAIYGLKQAARCW 1048
Query: 1263 YERLSSFLLENEFVRGKVDTTLFC--KTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMM 1320
+E L E EFV VD ++ K ++ + V +YVDD++ + + + F +
Sbjct: 1049 FEVFEQALKECEFVNSSVDRCIYILDKGNINENIYVLLYVDDVVIATGDMTRMNNFKRYL 1108
Query: 1321 QAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCIL 1380
+F M+ + E+K+F+GI+++ + Y+ QS Y K++L KFNM TP+ P+ I
Sbjct: 1109 MEKFRMTDLNEIKHFIGIRIEMQEDKIYLSQSAYVKKILSKFNMENCNAVSTPL-PSKIN 1167
Query: 1381 EKEDKSGKVCQKLYRGMIGSLLY-LTASRPDILFSVHLCARFQSDPRETHLTAVKRILRY 1439
+ S + C R +IG L+Y + +RPD+ +V++ +R+ S +KR+LRY
Sbjct: 1168 YELLNSDEDCNTPCRSLIGCLMYIMLCTRPDLTTAVNILSRYSSKNNSELWQNLKRVLRY 1227
Query: 1440 LKGTTNLGLMYKK--T*EYKLSGYCDADYAGDRTERKSTSGNC-QFLGSNLVSWASKRQS 1496
LKGT ++ L++KK E K+ GY D+D+AG +RKST+G + NL+ W +KRQ+
Sbjct: 1228 LKGTIDMKLIFKKNLAFENKIIGYVDSDWAGSEIDRKSTTGYLFKMFDFNLICWNTKRQN 1287
Query: 1497 TIALSTAEAEYISAAICSTQMLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSR 1555
++A S+ EAEY++ + LW+K L I LE+ I IY DN IS++ NP H R
Sbjct: 1288 SVAASSTEAEYMALFEAVREALWLKFLLTSINIKLENPIKIYEDNQGCISIANNPSCHKR 1347
Query: 1556 AKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRF 1599
AKHI++KYHF R+ VQ V+ L+++ T++Q ADIFTKPL RF
Sbjct: 1348 AKHIDIKYHFAREQVQNNVICLEYIPTENQLADIFTKPLPAARF 1391
Score = 46.6 bits (109), Expect = 6e-04
Identities = 76/358 (21%), Positives = 132/358 (36%), Gaps = 75/358 (20%)
Query: 9 NAKPPMFDGQRFEYWKDRLESFFLGFDADLWDIIVDGYERPVDADGKKIPRSEMTAEQKK 68
N KP FDG+++ WK R+ + + D+ ++ VD KK R
Sbjct: 7 NIKP--FDGEKYAIWKFRIRALLA--EQDVLKVVDGLMPNEVDDSWKKAERC-------- 54
Query: 69 LYSQHHKARAILLSAISYEEYQKITDREFAKGIFDSLKMSHEGNKKVKESKALSLIQKYE 128
A++ ++ +S T A+ I ++L +E + L+L ++
Sbjct: 55 -------AKSTIIEYLSDSFLNFATSDITARQILENLDAVYERKSLASQ---LALRKRLL 104
Query: 129 SFIMEPNESIEEMFSRFQLLVAGIRPLNKSYTTKDHVIRVIRCLPESWMPLVTSIELTRD 188
S + S+ F F L++ + D + ++ LP + ++T+IE T
Sbjct: 105 SLKLSSEMSLLSHFHIFDELISELLAAGAKIEEMDKISHLLITLPSCYDGIITAIE-TLS 163
Query: 189 VENMSLEELISILKCHELK-RSEMQDLRKKSIALKSKSEKAKVEKSKALQAEEEESEEAS 247
EN++L + + L E+K +++ D KK + A V +
Sbjct: 164 EENLTLAFVKNRLLDQEIKIKNDHNDTSKKVM-------NAIVHNNN------------- 203
Query: 248 EDSGEDELTLISKRLNRIWKHRQSKFKGSGKAKGKYESSGQKKSSIREVTCFECKESGHY 307
+ N ++K+R +K K K KY +V C C GH
Sbjct: 204 -----------NTYKNNLFKNRVTKPKKIFKGNSKY-----------KVKCHHCGREGHI 241
Query: 308 KSDCPKLK-----KDKKPKKHFKTKKSLMVTFDESESEDVDSDGEVQGLMAIVKDKGA 360
K DC K K+K+ +K +T S + F E + V V D GA
Sbjct: 242 KKDCFHYKRILNNKNKENEKQVQTATSHGIAFMVKEVNNT----SVMDNCGFVLDSGA 295
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 342 bits (876), Expect = 6e-93
Identities = 204/600 (34%), Positives = 336/600 (56%), Gaps = 42/600 (7%)
Query: 1030 VSDKGKAPEEVEPEEDEPEEEA--------GPSNSQTLKKSRITAAHPKELILGNKDEPV 1081
VS++G+ P EV + ++ +E G Q L++S E
Sbjct: 747 VSEQGEQPGEVIEQGEQLDEGVEEVEHPTQGEEQHQPLRRS----------------ERP 790
Query: 1082 RTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQ--DKDWIL-AMEEELNQFSKNDVWSLV 1138
R S PS E +L + EP+S+ E L +K+ ++ AM+EE+ KN + LV
Sbjct: 791 RVESRRYPSTEYVL----ISDDREPESLKEVLSHPEKNQLMKAMQEEMESLQKNGTYKLV 846
Query: 1139 KKPENVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRL 1198
+ P+ + KWVF+ K + +VR KARLV +G+ Q++GID+ E F+PV ++ +IR
Sbjct: 847 ELPKGKRPLKCKWVFKLKKDGDCKLVRYKARLVVKGFEQKKGIDFDEIFSPVVKMTSIRT 906
Query: 1199 LISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQA 1258
++S + + ++ + Q+DVK+AFL+G + EE+Y+ QP GFE K V KL KSLYGLKQA
Sbjct: 907 ILSLAASLDLEVEQLDVKTAFLHGDLEEEIYMEQPEGFEVAGKKHMVCKLNKSLYGLKQA 966
Query: 1259 PRAWYERLSSFLLENEFVRGKVDTTLFCKTY-KDDILIVQIYVDDIIFGSANQSLCKEFS 1317
PR WY + SF+ +++ D ++ K + +++ +I+ +YVDD++ ++ L +
Sbjct: 967 PRQWYMKFDSFMKSQTYLKTYSDPCVYFKRFSENNFIILLLYVDDMLIVGKDKGLIAKLK 1026
Query: 1318 EMMQAEFEMSMMGELKYFLGIQV--DQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMH 1375
+ F+M +G + LG+++ ++T ++ Q KY + +L++FNM + TP+
Sbjct: 1027 GDLSKSFDMKDLGPAQQILGMKIVRERTSRKLWLSQEKYIERVLERFNMKNAKPVSTPLA 1086
Query: 1376 PTCILEKE------DKSGKVCQKLYRGMIGSLLY-LTASRPDILFSVHLCARFQSDPRET 1428
L K+ ++ G + + Y +GSL+Y + +RPDI +V + +RF +P +
Sbjct: 1087 GHLKLSKKMCPTTVEEKGNMAKVPYSSAVGSLMYAMVCTRPDIAHAVGVVSRFLENPGKE 1146
Query: 1429 HLTAVKRILRYLKGTTNLGLMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLV 1488
H AVK ILRYL+GTT L + + + L GY DAD AGD RKS++G +
Sbjct: 1147 HWEAVKWILRYLRGTTGDCLCFGGS-DPILKGYTDADMAGDIDNRKSSTGYLFTFSGGAI 1205
Query: 1489 SWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSK 1548
SW SK Q +ALST EAEYI+A +M+W+K L++ + + +YCD+ +AI LSK
Sbjct: 1206 SWQSKLQKCVALSTTEAEYIAATETGKEMIWLKRFLQELGLHQKEYVVYCDSQSAIDLSK 1265
Query: 1549 NPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFILKNLNM 1608
N + H+R KHI+V+YH+IR+ V L + + T+ AD+ TK + ++F + + M
Sbjct: 1266 NSMYHARTKHIDVRYHWIREMVDDESLKVLKISTNENPADMLTKVVPRNKFELCKELVGM 1325
Score = 52.0 bits (123), Expect = 1e-05
Identities = 80/369 (21%), Positives = 144/369 (38%), Gaps = 68/369 (18%)
Query: 17 GQRFEYWKDRLESFFLGFDADLWDIIVD-GYERPVDADGKKIPRSEMTAEQKKLYSQHHK 75
G ++E K ++ F + + D+++ G + +D D KK T + + +
Sbjct: 3 GVKYEVAKFNGDNGFSTWQRRMRDLLIQQGLHKVLDVDSKKPD----TMKAEDWADLDER 58
Query: 76 ARAILLSAISYEEYQKITDREFAKGIF---DSLKMSHEGNKKVKESKALSLIQKYESFIM 132
A + + +S + I D + A+GI+ +SL MS K+ K L + M
Sbjct: 59 AASAIRLHLSDDVVNNIIDEDTARGIWTRLESLYMSKTLTNKLYLKKQLYALH------M 112
Query: 133 EPNESIEEMFSRFQLLVAGIRPLNKSYTTKDHVIRVIRCLPESWMPLVTSIELTRDVENM 192
+ + F L+ + L +D I ++ LP S+ L T+I +
Sbjct: 113 SEGTNFLSHLNVFNGLITQLANLGVKIEEEDKAILLLNSLPSSYDNLATTI--LHGKTTI 170
Query: 193 SLEELISILKCHELKRSEMQDLRKKSIALKSKSEKAKVEKSKALQAEEEESEEASEDSGE 252
L+++ S L +E +RKK E+ G+
Sbjct: 171 ELKDVTSALLLNE-------KMRKKP-----------------------------ENQGQ 194
Query: 253 DELTLISKRLNRIWKHRQSKFKGSGKAKGKYESSGQKKSSIREVTCFECKESGHYKSDCP 312
LI++ R ++ + + SG A+GK S + KS +R C+ C + GH+K DCP
Sbjct: 195 ---ALITEGRGRSYQRSSNNYGRSG-ARGK--SKNRSKSRVRN--CYNCNQPGHFKRDCP 246
Query: 313 KLKKDKKPKKHFKT--KKSLMVTFDESESEDVDSDGEVQGLMAIVKDKGAESKEAVDSDS 370
+K K K + MV +++ ++ + E L G ES+ VD+ +
Sbjct: 247 NPRKGKGETSGQKNDDNTAAMVQNNDNVVLFINEEEECMHL------SGPESEWVVDTAA 300
Query: 371 ESEGDPDSD 379
P D
Sbjct: 301 SHHATPVRD 309
>YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein
Length = 308
Score = 161 bits (407), Expect = 2e-38
Identities = 96/309 (31%), Positives = 166/309 (53%), Gaps = 17/309 (5%)
Query: 1213 MDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLE 1272
MDV +AFLN + E +YV QPPGF +E+ PD+V++L +YGLKQAP W E +++ L +
Sbjct: 1 MDVDTAFLNSTMDEPIYVKQPPGFVNERNPDYVWELYGGMYGLKQAPLLWNEHINNTLKK 60
Query: 1273 NEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGEL 1332
F R + + L+ ++ D + + +YVDD++ + + + + + + M +G++
Sbjct: 61 IGFCRHEGEHGLYFRSTSDGPIYIGVYVDDLLVAAPSPKIYDRVKQELTKLYSMKDLGKV 120
Query: 1333 KYFLGIQVDQTPEG-------TYIHQSKYTKELLKKFNMLESTVAKT-PMHPTCILEKED 1384
FLG+ + Q+ G YI ++ E + F + ++ + + P+ T +D
Sbjct: 121 DKFLGLNIHQSTNGDITLSLQDYIAKAASESE-INTFKLTQTPLCNSKPLFETTSPHLKD 179
Query: 1385 KSGKVCQKLYRGMIGSLLY-LTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGT 1443
+ Y+ ++G LL+ RPDI + V L +RF +PR HL + +R+LRYL T
Sbjct: 180 ITP------YQSIVGQLLFCANTGRPDISYPVSLLSRFLREPRAIHLESARRVLRYLYTT 233
Query: 1444 TNLGLMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKR-QSTIALST 1502
++ L Y+ + L+ YCDA + ST G L V+W+SK+ + I + +
Sbjct: 234 RSMCLKYRSGSQVALTVYCDASHGAIHDLPHSTGGYVTLLAGAPVTWSSKKLKGVIPVPS 293
Query: 1503 AEAEYISAA 1511
EAEYI+A+
Sbjct: 294 TEAEYITAS 302
>M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810
(ORF240b)
Length = 240
Score = 154 bits (390), Expect = 1e-36
Identities = 75/222 (33%), Positives = 129/222 (57%), Gaps = 1/222 (0%)
Query: 1298 IYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKE 1357
+YVDDI+ ++ +L + + F M +G + YFLGIQ+ P G ++ Q+KY ++
Sbjct: 5 LYVDDILLTGSSNTLLNMLIFQLSSTFSMKDLGPVHYFLGIQIKTHPSGLFLSQTKYAEQ 64
Query: 1358 LLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHL 1417
+L ML+ TP+ P + + +R ++G+L YLT +RPDI ++V++
Sbjct: 65 ILNNAGMLDCKPMSTPL-PLKLNSSVSTAKYPDPSDFRSIVGALQYLTLTRPDISYAVNI 123
Query: 1418 CARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKT*EYKLSGYCDADYAGDRTERKSTS 1477
+ +P +KR+LRY+KGT GL K + + +CD+D+AG + R+ST+
Sbjct: 124 VCQRMHEPTLADFDLLKRVLRYVKGTIFHGLYIHKNSKLNVQAFCDSDWAGCTSTRRSTT 183
Query: 1478 GNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLW 1519
G C FLG N++SW++KRQ T++ S+ E EY + A+ + ++ W
Sbjct: 184 GFCTFLGCNIISWSAKRQPTVSRSSTETEYRALALTAAELTW 225
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 117 bits (294), Expect = 2e-25
Identities = 131/521 (25%), Positives = 233/521 (44%), Gaps = 49/521 (9%)
Query: 1114 QDKDWILAMEEELNQFSKNDVWSLV-----KKPENVHVIGTKWVFRNKLNEKGDVVRNKA 1168
+ + +I A +E+NQ K W K+ + VI + ++F N+K D +KA
Sbjct: 1246 EKEKYIEAYHKEVNQLLKMKTWDTDEYYDRKEIDPKRVINSMFIF----NKKRDGT-HKA 1300
Query: 1169 RLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEV 1228
R VA+G Q + + A+ +S ++++N + Q+D+ SA+L I EE+
Sbjct: 1301 RFVARGDIQHPDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISSAYLYADIKEEL 1360
Query: 1229 YVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLEN---EFVRGKVDTTLF 1285
Y+ PP D + +LKKSLYGLKQ+ WYE + S+L++ E VRG +
Sbjct: 1361 YIRPPPHLGMN---DKLIRLKKSLYGLKQSGANWYETIKSYLIQQCGMEEVRG------W 1411
Query: 1286 CKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMG------ELKY-FLGI 1338
+K+ + + ++VDD++ S N + K E ++ +++ ++ E++Y LG+
Sbjct: 1412 SCVFKNSQVTICLFVDDMVLFSKNLNSNKRIIEKLKMQYDTKIINLGESDEEIQYDILGL 1471
Query: 1339 QVDQTPEGTY--IHQSKYTKELLKKFNM---LESTVAKTPMHPTCI-----LEKEDKSGK 1388
++ + G Y + E + K N+ + P P LE E+ K
Sbjct: 1472 EI-KYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYK 1530
Query: 1389 VCQKLYRGMIGSLLYLTAS-RPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLG 1447
+ + +IG Y+ R D+L+ ++ A+ P + L +++++ T +
Sbjct: 1531 MKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWNTRDKQ 1590
Query: 1448 LMYKKT*EY----KLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTA 1503
L++ K+ KL DA Y G++ KS GN L ++ S + S ST
Sbjct: 1591 LIWHKSKPVKPTNKLVVISDASY-GNQPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTT 1649
Query: 1504 EAEY--ISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEV 1561
EAE IS ++ L Q D + + + +T +I +S N R +
Sbjct: 1650 EAEIHAISESVPLLNNLSYLIQELDKKPITKGLLTDSKSTISIIISNNE-EKFRNRFFGT 1708
Query: 1562 KYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFI 1602
K +RD V L + +++T AD+ TKPL F +
Sbjct: 1709 KAMRLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLL 1749
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 115 bits (287), Expect = 1e-24
Identities = 126/525 (24%), Positives = 234/525 (44%), Gaps = 57/525 (10%)
Query: 1114 QDKDWILAMEEELNQFSKNDVWSLV-----KKPENVHVIGTKWVFRNKLNEKGDVVRNKA 1168
+ + +I A +E+NQ K W K+ + VI + ++F K + +KA
Sbjct: 819 EKEKYIQAYHKEVNQLLKMKTWDTDRYYDRKEIDPKRVINSMFIFNRKRDGT-----HKA 873
Query: 1169 RLVAQGYSQQEGIDYTETFAPVARLE-----AIRLLISFSVNHNIVLHQMDVKSAFLNGY 1223
R VA+G I + +T+ P + A+ +S ++++N + Q+D+ SA+L
Sbjct: 874 RFVARG-----DIQHPDTYDPGMQSNTVHHYALMTSLSLALDNNYYITQLDISSAYLYAD 928
Query: 1224 ISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLEN---EFVRGKV 1280
I EE+Y+ PP D + +LKKSLYGLKQ+ WYE + S+L++ E VRG
Sbjct: 929 IKEELYIRPPPHLGMN---DKLIRLKKSLYGLKQSGANWYETIKSYLIKQCGMEEVRG-- 983
Query: 1281 DTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMM------GELKY 1334
+ +K+ + + ++VDD+I S + + K+ ++ +++ ++ E++Y
Sbjct: 984 ----WSCVFKNSQVTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQY 1039
Query: 1335 -FLGIQVDQTPEGTY--IHQSKYTKELLKKFNM---LESTVAKTPMHPTCI-----LEKE 1383
LG+++ + G Y + E + K N+ + P P LE E
Sbjct: 1040 DILGLEI-KYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELE 1098
Query: 1384 DKSGKVCQKLYRGMIGSLLYLTAS-RPDILFSVHLCARFQSDPRETHLTAVKRILRYLKG 1442
+ K+ + +IG Y+ R D+L+ ++ A+ P + L +++++
Sbjct: 1099 EDDYKMKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWN 1158
Query: 1443 TTNLGLMYKKT*EY----KLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTI 1498
T + L++ K+ KL DA Y G++ KS GN L ++ S + S
Sbjct: 1159 TRDKQLIWHKSKPVKPTNKLVVISDASY-GNQPYYKSQIGNIYLLNGKVIGGKSTKASLT 1217
Query: 1499 ALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAIS-LSKNPILHSRAK 1557
ST EAE + + + + H +++ + D+ + IS + N R +
Sbjct: 1218 CTSTTEAEIHAISESVPLLNNLSHLVQELNKKPITKGLLTDSKSTISIIISNNEEKFRNR 1277
Query: 1558 HIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFI 1602
K +RD V L + +++T AD+ TKPL F +
Sbjct: 1278 FFGTKAMRLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLL 1322
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 115 bits (287), Expect = 1e-24
Identities = 126/523 (24%), Positives = 240/523 (45%), Gaps = 53/523 (10%)
Query: 1114 QDKDWILAMEEELNQFSKNDVWSLV-----KKPENVHVIGTKWVFRNKLNEKGDVVRNKA 1168
+ + +I A +E+NQ K W K+ + VI + ++F N+K D +KA
Sbjct: 819 EKEKYIEAYHKEVNQLLKMKTWDTDEYYDRKEIDPKRVINSMFIF----NKKRDGT-HKA 873
Query: 1169 RLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEV 1228
R VA+G Q + + A+ +S ++++N + Q+D+ SA+L I EE+
Sbjct: 874 RFVARGDIQHPDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISSAYLYADIKEEL 933
Query: 1229 YVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLEN---EFVRGKVDTTLF 1285
Y+ PP D + +LKKSLYGLKQ+ WYE + S+L++ E VRG +
Sbjct: 934 YIRPPPHLGMN---DKLIRLKKSLYGLKQSGANWYETIKSYLIKQCGMEEVRG------W 984
Query: 1286 CKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMM------GELKY-FLGI 1338
+K+ + + ++VDD+I S + + K+ ++ +++ ++ E++Y LG+
Sbjct: 985 SCVFKNSQVTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGL 1044
Query: 1339 QVDQTPEGTY--IHQSKYTKELLKKFNM---LESTVAKTPMHPTCILEKED---KSGKVC 1390
++ + G Y + E + K N+ + P P +++++ +
Sbjct: 1045 EI-KYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQDELEIDEDEYK 1103
Query: 1391 QKLY--RGMIGSLLYLTAS-RPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLG 1447
+K++ + +IG Y+ R D+L+ ++ A+ P L +++++ T +
Sbjct: 1104 EKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQ 1163
Query: 1448 LMYKKT----*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTA 1503
L++ K + KL DA Y G++ KS GN L ++ S + S ST
Sbjct: 1164 LIWHKNKPTEPDNKLVAISDASY-GNQPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTT 1222
Query: 1504 EAEY--ISAAI-CSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHS-RAKHI 1559
EAE IS ++ + ++ +L I++ + D+ + IS+ K+ R +
Sbjct: 1223 EAEIHAISESVPLLNNLSYLIQELNKKPIIKG---LLTDSRSTISIIKSTNEEKFRNRFF 1279
Query: 1560 EVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFI 1602
K +RD V L + +++T AD+ TKPL F +
Sbjct: 1280 GTKAMRLRDEVSGNNLYVYYIETKKNIADVMTKPLPIKTFKLL 1322
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 115 bits (287), Expect = 1e-24
Identities = 132/566 (23%), Positives = 261/566 (45%), Gaps = 63/566 (11%)
Query: 1079 EPVRTRSAFRPSEETLLSLKGLVSLIEPKSI---DEAL------QDKD-WILAMEEELNQ 1128
EP R++ + ++KG+ S+ ++ DEA+ ++KD ++ A +E++Q
Sbjct: 1220 EPPRSKKRIN----LIAAIKGVKSIKPVRTTLRYDEAITYNKDNKEKDRYVEAYHKEISQ 1275
Query: 1129 FSKNDVWSLVKKPEN-----VHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDY 1183
K + W K + VI + ++F N+K D +KAR VA+G Q
Sbjct: 1276 LLKMNTWDTNKYYDRNDIDPKKVINSMFIF----NKKRDGT-HKARFVARGDIQHPDTYD 1330
Query: 1184 TETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPP--GFEDEKK 1241
++ + A+ +S +++++ + Q+D+ SA+L I EE+Y+ PP G D+
Sbjct: 1331 SDMQSNTVHHYALMTSLSIALDNDYYITQLDISSAYLYADIKEELYIRPPPHLGLNDK-- 1388
Query: 1242 PDHVFKLKKSLYGLKQAPRAWYERLSSFLL---ENEFVRGKVDTTLFCKTYKDDILIVQI 1298
+ +L+KSLYGLKQ+ WYE + S+L+ + + VRG + +K+ + + +
Sbjct: 1389 ---LLRLRKSLYGLKQSGANWYETIKSYLINCCDMQEVRG------WSCVFKNSQVTICL 1439
Query: 1299 YVDDIIFGSANQSLCKEFSEMMQAEFEMSMM------GELKY-FLGIQVD-QTPEGTYIH 1350
+VDD+I S + + K+ ++ +++ ++ E++Y LG+++ Q + +
Sbjct: 1440 FVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRSKYMKLG 1499
Query: 1351 QSKYTKELLKKFNMLESTVAK---TPMHPTCILEKED---KSGKVCQKLY--RGMIGSLL 1402
K E L K N+ + K P P +++++ + +K++ + +IG
Sbjct: 1500 MEKSLTEKLPKLNVPLNPKGKKLRAPGQPGHYIDQDELEIDEDEYKEKVHEMQKLIGLAS 1559
Query: 1403 YLTAS-RPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKT----*EYK 1457
Y+ R D+L+ ++ A+ P L +++++ T + L++ K + K
Sbjct: 1560 YVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHKNKPTKPDNK 1619
Query: 1458 LSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQM 1517
L DA Y G++ KS GN L ++ S + S ST EAE + + +
Sbjct: 1620 LVAISDASY-GNQPYYKSQIGNIFLLNGKVIGGKSTKASLTCTSTTEAEIHAVSEAIPLL 1678
Query: 1518 LWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHS-RAKHIEVKYHFIRDYVQKGVLL 1576
+ H +++ + D+ + IS+ K+ R + K +RD V L
Sbjct: 1679 NNLSHLVQELNKKPIIKGLLTDSRSTISIIKSTNEEKFRNRFFGTKAMRLRDEVSGNNLY 1738
Query: 1577 LKFVDTDHQWADIFTKPLAEDRFNFI 1602
+ +++T AD+ TKPL F +
Sbjct: 1739 VYYIETKKNIADVMTKPLPIKTFKLL 1764
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 114 bits (285), Expect = 2e-24
Identities = 127/521 (24%), Positives = 231/521 (43%), Gaps = 49/521 (9%)
Query: 1114 QDKDWILAMEEELNQFSKNDVWSLVK-----KPENVHVIGTKWVFRNKLNEKGDVVRNKA 1168
+ + +I A +E+NQ K W K + + VI + ++F K + +KA
Sbjct: 819 EKEKYIEAYHKEVNQLLKMKTWDTDKYYDRKEIDPKRVINSMFIFNRKRDGT-----HKA 873
Query: 1169 RLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEV 1228
R VA+G Q + + A+ +S ++++N + Q+D+ SA+L I EE+
Sbjct: 874 RFVARGDIQHPDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISSAYLYADIKEEL 933
Query: 1229 YVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLEN---EFVRGKVDTTLF 1285
Y+ PP D + +LKKSLYGLKQ+ WYE + S+L++ E VRG +
Sbjct: 934 YIRPPPHLGMN---DKLIRLKKSLYGLKQSGANWYETIKSYLIKQCGMEEVRG------W 984
Query: 1286 CKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMG------ELKY-FLGI 1338
+++ + + ++VDD++ S N + K + ++ +++ ++ E++Y LG+
Sbjct: 985 SCVFENSQVTICLFVDDMVLFSKNLNSNKRIIDKLKMQYDTKIINLGESDEEIQYDILGL 1044
Query: 1339 QVDQTPEGTY--IHQSKYTKELLKKFNM---LESTVAKTPMHPTCI-----LEKEDKSGK 1388
++ + G Y + E + K N+ + P P LE E+ K
Sbjct: 1045 EI-KYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYK 1103
Query: 1389 VCQKLYRGMIGSLLYLTAS-RPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLG 1447
+ + +IG Y+ R D+L+ ++ A+ P + L +++++ T +
Sbjct: 1104 MKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWNTRDKQ 1163
Query: 1448 LMYKKT*EY----KLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTA 1503
L++ K+ KL DA Y G++ KS GN L ++ S + S ST
Sbjct: 1164 LIWHKSKPVKPTNKLVVISDASY-GNQPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTT 1222
Query: 1504 EAEY--ISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEV 1561
EAE IS ++ L Q D + + + +T +I +S N R +
Sbjct: 1223 EAEIHAISESVPLLNNLSYLIQELDKKPITKGLLTDSKSTISIIISNNE-EKFRNRFFGT 1281
Query: 1562 KYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFI 1602
K +RD V L + +++T AD+ TKPL F +
Sbjct: 1282 KAMRLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLL 1322
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 114 bits (285), Expect = 2e-24
Identities = 121/520 (23%), Positives = 234/520 (44%), Gaps = 47/520 (9%)
Query: 1114 QDKDWILAMEEELNQFSKNDVWSLVK-----KPENVHVIGTKWVFRNKLNEKGDVVRNKA 1168
+ + +I A +E+NQ K + W K + + VI + ++F K + +KA
Sbjct: 819 EKEKYIEAYHKEVNQLLKMNTWDTDKYYDRKEIDPKRVINSMFIFNRKRDGT-----HKA 873
Query: 1169 RLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEV 1228
R VA+G Q + + A+ +S ++++N + Q+D+ SA+L I EE+
Sbjct: 874 RFVARGDIQHPDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISSAYLYADIKEEL 933
Query: 1229 YVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLEN---EFVRGKVDTTLF 1285
Y+ PP D + +LKKSLYGLKQ+ WYE + S+L++ E VRG +
Sbjct: 934 YIRPPPHLGMN---DKLIRLKKSLYGLKQSGANWYETIKSYLIKQCGMEEVRG------W 984
Query: 1286 CKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMM------GELKY-FLGI 1338
+K+ + + ++VDD+I S + + K+ ++ +++ ++ E++Y LG+
Sbjct: 985 SCVFKNSQVTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGL 1044
Query: 1339 QVDQTPEGTY--IHQSKYTKELLKKFNM---LESTVAKTPMHPTCILEKED---KSGKVC 1390
++ + G Y + E + K N+ + P P +++++ +
Sbjct: 1045 EI-KYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQDELEIDEDEYK 1103
Query: 1391 QKLY--RGMIGSLLYLTAS-RPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLG 1447
+K++ + +IG Y+ R D+L+ ++ A+ P L +++++ T +
Sbjct: 1104 EKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQ 1163
Query: 1448 LMYKKT----*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTA 1503
L++ K + KL DA Y G++ KS GN L ++ S + S ST
Sbjct: 1164 LIWHKNKPTEPDNKLVAISDASY-GNQPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTT 1222
Query: 1504 EAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAIS-LSKNPILHSRAKHIEVK 1562
EAE + + + + H +++ + D+ + IS + N R + K
Sbjct: 1223 EAEIHAISESVPLLNNLSHLVQELNKKPITKGLLTDSKSTISIIISNNEEKFRNRFFGTK 1282
Query: 1563 YHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFI 1602
+RD V L + +++T AD+ TKPL F +
Sbjct: 1283 AMRLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLL 1322
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 114 bits (285), Expect = 2e-24
Identities = 145/614 (23%), Positives = 268/614 (43%), Gaps = 94/614 (15%)
Query: 1032 DKGKAPEEVEPEEDEPEEEAG-----PSNSQTLKKSRITAAHPKELILGNKDEPVRTRSA 1086
DK + E E D+ + P N +T+ +I A + E I N D ++ +
Sbjct: 1231 DKNNSLTSYELERDKKRSKKNRVKLIPDNMETVSAPKIRAIYYNEAISKNPD--LKEKHE 1288
Query: 1087 FRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVHV 1146
++ + K L +L + K D ++ +S++++ P+N+ +
Sbjct: 1289 YKQAYH-----KELQNLKDMKVFDVDVK--------------YSRSEI------PDNL-I 1322
Query: 1147 IGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNH 1206
+ T +F K N KAR+V +G +Q Y+ I++ + + N
Sbjct: 1323 VPTNTIFTKKRNGI-----YKARIVCRGDTQSPDT-YSVITTESLNHNHIKIFLMIANNR 1376
Query: 1207 NIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERL 1266
N+ + +D+ AFL + EE+Y+ P D + V KL K+LYGLKQ+P+ W + L
Sbjct: 1377 NMFMKTLDINHAFLYAKLEEEIYIPHP---HDRRC---VVKLNKALYGLKQSPKEWNDHL 1430
Query: 1267 SSFL-----LENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQ 1321
+L +N + G T +D L++ +YVDD + ++N+ EF ++
Sbjct: 1431 RQYLNGIGLKDNSYTPGLYQT-------EDKNLMIAVYVDDCVIAASNEQRLDEFINKLK 1483
Query: 1322 AEFEMSMMGEL------KYFLGIQ---------VDQTPEGTYIHQ--SKYTKEL--LKKF 1362
+ FE+ + G L LG+ +D T + ++I++ KY +EL ++K
Sbjct: 1484 SNFELKITGTLIDDVLDTDILGMDLVYNKRLGTIDLTLK-SFINRMDKKYNEELKKIRKS 1542
Query: 1363 NMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTAS-RPDILFSVHLCARF 1421
++ + K + E++ + KL + ++G L Y+ R DI F+V AR
Sbjct: 1543 SIPHMSTYKIDPKKDVLQMSEEEFRQGVLKLQQ-LLGELNYVRHKCRYDIEFAVKKVARL 1601
Query: 1422 QSDPRETHLTAVKRILRYLKGTTNLGLMYKK--T*EYKLSGYCDADYAGDRTERKSTSGN 1479
+ P E + +I++YL ++G+ Y + + K+ DA G + +S G
Sbjct: 1602 VNYPHERVFYMIYKIIQYLVRYKDIGIHYDRDCNKDKKVIAITDAS-VGSEYDAQSRIGV 1660
Query: 1480 CQFLGSNLVSWASKRQSTIALSTAEAE----YISAAICSTQMLWMKHQLEDYQILESNIP 1535
+ G N+ + S + + +S+ EAE Y A T + +K E ++I
Sbjct: 1661 ILWYGMNIFNVYSNKSTNRCVSSTEAELHAIYEGYADSETLKVTLKELGEGDN---NDIV 1717
Query: 1536 IYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV-QKGVLLLKFVDTDHQWADIFTKPL 1594
+ D+ AI + K +K I++ + +K + LLK + AD+ TKP+
Sbjct: 1718 MITDSKPAIQGLNRSYQQPKEKFTWIKTEIIKEKIKEKSIKLLKITGKGNI-ADLLTKPV 1776
Query: 1595 AED---RFNFILKN 1605
+ RF +LKN
Sbjct: 1777 SASDFKRFIQVLKN 1790
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 113 bits (282), Expect = 5e-24
Identities = 126/523 (24%), Positives = 239/523 (45%), Gaps = 53/523 (10%)
Query: 1114 QDKDWILAMEEELNQFSKNDVWSLV-----KKPENVHVIGTKWVFRNKLNEKGDVVRNKA 1168
+ + +I A +E+NQ K W K+ + VI + ++F N+K D +KA
Sbjct: 1246 EKEKYIEAYHKEVNQLLKMKTWDTDEYYDRKEIDPKRVINSMFIF----NKKRDGT-HKA 1300
Query: 1169 RLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEV 1228
R VA+G Q T + A+ +S ++++N + Q+D+ SA+L I EE+
Sbjct: 1301 RFVARGDIQHPDTYDTGMQSNTVHHYALMTSLSLALDNNYYITQLDISSAYLYADIKEEL 1360
Query: 1229 YVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLEN---EFVRGKVDTTLF 1285
Y+ PP D + +LKKS YGLKQ+ WYE + S+L++ E VRG +
Sbjct: 1361 YIRPPPHLGMN---DKLIRLKKSHYGLKQSGANWYETIKSYLIKQCGMEEVRG------W 1411
Query: 1286 CKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMM------GELKY-FLGI 1338
+K+ + + ++VDD+I S + + K+ ++ +++ ++ E++Y LG+
Sbjct: 1412 SCVFKNSQVTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGL 1471
Query: 1339 QVDQTPEGTY--IHQSKYTKELLKKFNM---LESTVAKTPMHPTCILEKED---KSGKVC 1390
++ + G Y + E + K N+ + P P +++++ +
Sbjct: 1472 EI-KYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQDELEIDEDEYK 1530
Query: 1391 QKLY--RGMIGSLLYLTAS-RPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLG 1447
+K++ + +IG Y+ R D+L+ ++ A+ P L +++++ T +
Sbjct: 1531 EKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQ 1590
Query: 1448 LMYKKT----*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTA 1503
L++ K + KL DA Y G++ KS GN L ++ S + S ST
Sbjct: 1591 LIWHKNKPTEPDNKLVAISDASY-GNQPYYKSQIGNIFLLNGKVIGGKSTKASLTCTSTT 1649
Query: 1504 EAEY--ISAAI-CSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHS-RAKHI 1559
EAE IS ++ + ++ +L I++ + D+ + IS+ K+ R +
Sbjct: 1650 EAEIHAISESVPLLNNLSYLIQELNKKPIIKG---LLTDSRSTISIIKSTNEEKFRNRFF 1706
Query: 1560 EVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFI 1602
K +RD V L + +++T AD+ TKPL F +
Sbjct: 1707 GTKAMRLRDEVSGNNLYVYYIETKKNIADVMTKPLPIKTFKLL 1749
>M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820
(ORF170)
Length = 170
Score = 100 bits (250), Expect = 2e-20
Identities = 47/97 (48%), Positives = 67/97 (68%)
Query: 1105 EPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVHVIGTKWVFRNKLNEKGDVV 1164
EPKS+ AL+D W AM+EEL+ S+N W LV P N +++G KWVF+ KL+ G +
Sbjct: 27 EPKSVIFALKDPGWCQAMQEELDALSRNKTWILVPPPVNQNILGCKWVFKTKLHSDGTLD 86
Query: 1165 RNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLIS 1201
R KARLVA+G+ Q+EGI + ET++PV R IR +++
Sbjct: 87 RLKARLVAKGFHQEEGIYFVETYSPVVRTATIRTILN 123
>M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240
(ORF111a)
Length = 111
Score = 66.2 bits (160), Expect = 7e-10
Identities = 30/82 (36%), Positives = 49/82 (59%)
Query: 1402 LYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKT*EYKLSGY 1461
+YLT +RPD+ F+V+ ++F S R + AV ++L Y+KGT GL Y T + +L +
Sbjct: 1 MYLTITRPDLTFAVNRLSQFSSASRTAQMQAVYKVLHYVKGTVGQGLFYSATSDLQLKAF 60
Query: 1462 CDADYAGDRTERKSTSGNCQFL 1483
D+D+A R+S +G C +
Sbjct: 61 ADSDWASCPDTRRSVTGFCSLV 82
>GAR2_SCHPO (P41891) Protein gar2
Length = 500
Score = 52.4 bits (124), Expect = 1e-05
Identities = 59/218 (27%), Positives = 96/218 (43%), Gaps = 19/218 (8%)
Query: 184 ELTRDVENMSLEELISILKCH-ELKRSEMQDLRKKSIALKSKSEKAKVEKSKALQAEEEE 242
E +++ S + +S K E KR+ + KKS+ + KS+K K E S ++E
Sbjct: 35 EAAKEIAKQSSKTDVSPKKSKKEAKRASSPEPSKKSVKKQKKSKK-KEESSSESESESSS 93
Query: 243 SEEASEDSGEDELTLISKRLNRIWKHRQSKFKGSGKAKGKYESSGQKKSSIREVTCFECK 302
SE S S + + S+ + +S+ + K + K ESS + SS
Sbjct: 94 SESESSSSESESSSSESESSSSESSSSESEEEVIVKTEEKKESSSESSSS---------S 144
Query: 303 ESGHYKSDCPKLKKDKKPKKHFKTKKSLMVTFDESESEDVDSDGEVQGLMAIVKDKGAES 362
ES + K+++ K+ ++ S ESESE S+ E + V +K E
Sbjct: 145 ESEEEEEAVVKIEEKKESSSDSSSESSS----SESESESSSSESEEE---EEVVEKTEEK 197
Query: 363 KE-AVDSDSESEGDPDSDDENEVFASFSTSELKHALSD 399
KE + +S S+SE DS E+ S S SE + + D
Sbjct: 198 KEGSSESSSDSESSSDSSSESGDSDSSSDSESESSSED 235
Score = 40.8 bits (94), Expect = 0.030
Identities = 47/206 (22%), Positives = 80/206 (38%), Gaps = 9/206 (4%)
Query: 274 KGSGKAKGKYESSGQKKSSIREVTCFECKESGHYKSDCPKLKKD-KKPKKHFKTKKSLMV 332
K + KG E + K +E K+S K KK+ K+ +KKS+
Sbjct: 14 KETASKKGAIEKPSKSKKITKEAAKEIAKQSSKTDVSPKKSKKEAKRASSPEPSKKSVKK 73
Query: 333 TFDESESEDVDSDGEVQGLMAIVKDKGAESKEAVDSDSESEGDPDSDDENEVFASFSTSE 392
+ E+ S+ E + + + +ES E+ S+SES S E+E T E
Sbjct: 74 QKKSKKKEESSSESESESSSSESESSSSES-ESSSSESESSSSESSSSESEEEVIVKTEE 132
Query: 393 LKHALSDIMDKYNS-------LLSKHKKLKKNLSAVSKTPSEHEKIISDLKNDNHALVNS 445
K + S+ S + + KK + S+ + SE E S +++ V
Sbjct: 133 KKESSSESSSSSESEEEEEAVVKIEEKKESSSDSSSESSSSESESESSSSESEEEEEVVE 192
Query: 446 NSVLKNQIAKLEEVIACDASDSKHES 471
+ K + + + +SDS ES
Sbjct: 193 KTEEKKEGSSESSSDSESSSDSSSES 218
Score = 35.8 bits (81), Expect = 0.98
Identities = 41/198 (20%), Positives = 73/198 (36%), Gaps = 9/198 (4%)
Query: 199 SILKCHELKRSEMQDLRKKSIALKSKSEKAKVEKSKALQAEEEESEEASEDSGEDELTLI 258
S+ K + K+ E +S + S+SE + E + E S E+S E+E+ +
Sbjct: 70 SVKKQKKSKKKEESSSESESESSSSESESSSSESESSSSESESSSSESSSSESEEEVIVK 129
Query: 259 SKRLNRIWKHRQSKFKGSG------KAKGKYESSGQK--KSSIREVTCFECKESGHYKSD 310
++ S + K + K ESS +SS E + +
Sbjct: 130 TEEKKESSSESSSSSESEEEEEAVVKIEEKKESSSDSSSESSSSESESESSSSESEEEEE 189
Query: 311 CPKLKKDKKPKKHFKTKKSLMVTFDESESEDVDSDGEVQGLMAIVKDKGAESKEAVDSDS 370
+ ++KK + S + SES D DS + + + +K +++ A +
Sbjct: 190 VVEKTEEKKEGSSESSSDSESSSDSSSESGDSDSSSDSESESSSEDEKKRKAEPASEERP 249
Query: 371 ESEGDPDSDDENEVFASF 388
P S D NE F
Sbjct: 250 AKITKP-SQDSNETCTVF 266
>USO1_YEAST (P25386) Intracellular protein transport protein USO1
Length = 1790
Score = 52.0 bits (123), Expect = 1e-05
Identities = 85/432 (19%), Positives = 179/432 (40%), Gaps = 47/432 (10%)
Query: 99 KGIFDSLKMSHEGNKKVKESKALSLIQKYESFIMEPNESIEEMFSRFQLLVAGIRPL-NK 157
K D+L + + KK E+ SL++ +S E I+E+ + L +K
Sbjct: 1222 KSEIDALNLQIKELKKKNETNEASLLESIKSVESE-TVKIKELQDECNFKEKEVSELEDK 1280
Query: 158 SYTTKDHVIRVIRCLPESWMPLVTSIELTRDVENMSLEELISILKCHELKRSEMQDLRKK 217
++D + + ES + ++ + LE++ ++ K E SE+ L+K
Sbjct: 1281 LKASEDKNSKYLELQKES-EKIKEELDAKTTELKIQLEKITNLSKAKEKSESELSRLKKT 1339
Query: 218 SIALKSKS----EKAKVEKSKALQAEEEESEEASEDSG------EDELTLISKRLNRIWK 267
S + + EK K E QA E+E + +E S +++ + L R+
Sbjct: 1340 SSEERKNAEEQLEKLKNEIQIKNQAFEKERKLLNEGSSTITQEYSEKINTLEDELIRLQN 1399
Query: 268 HRQSKFKGSGKAKGKYESSG--------QKKSSIR----EVTCFECKESGH--------- 306
+ K K + + E +K+++I+ E+ ++ K + +
Sbjct: 1400 ENELKAKEIDNTRSELEKVSLSNDELLEEKQNTIKSLQDEILSYKDKITRNDEKLLSIER 1459
Query: 307 -YKSDCPKLKKDKKPKKHFKTK--KSLMVTFDESESEDVD---SDGEVQGLMAIVKDKGA 360
K D LK+ + + K K + L +ES E + S ++ L + ++
Sbjct: 1460 DNKRDLESLKEQLRAAQESKAKVEEGLKKLEEESSKEKAELEKSKEMMKKLESTIESNET 1519
Query: 361 ESKEAVDSDSESEGDPDSDDENEVFASFSTSELKHALSDIMDKYNSLLSKHKKLKKNLSA 420
E K ++++ +S+ + ++++ A L+H SD++ + N ++LK L
Sbjct: 1520 ELKSSMETIRKSD---EKLEQSKKSAEEDIKNLQHEKSDLISRINESEKDIEELKSKLRI 1576
Query: 421 VSKTPSEHEKIISDLKNDNHAL---VNSNSVLKNQIAKLEEVIACDASDSKHESKYEKSF 477
+K+ SE E + +L N + N+VLK+++ +E + ++ K ++ EK
Sbjct: 1577 EAKSGSELETVKQELNNAQEKIRINAEENTVLKSKLEDIERELKDKQAEIK-SNQEEKEL 1635
Query: 478 QRFLAKSVDRSL 489
K +++ L
Sbjct: 1636 LTSRLKELEQEL 1647
Score = 50.1 bits (118), Expect = 5e-05
Identities = 95/420 (22%), Positives = 162/420 (37%), Gaps = 64/420 (15%)
Query: 82 SAISYEEYQKITDREFAKGIFDSLKMSHEGNKKVKESKALSLIQKYESFIMEPNESIEEM 141
S I+ E +KI E + +++ +E K KE I S + + + S +E+
Sbjct: 1377 STITQEYSEKINTLED-----ELIRLQNENELKAKE------IDNTRSELEKVSLSNDEL 1425
Query: 142 FSRFQLLVAGIRPLNKSYTTKDHVIRVIRCLPESWMPLVTSIELTRDVENMSLEELISIL 201
Q + ++ SY KD + R L + + RD LE L L
Sbjct: 1426 LEEKQNTIKSLQDEILSY--KDKITRNDEKL----------LSIERD-NKRDLESLKEQL 1472
Query: 202 KCHELKRSEMQDLRKKSIALKSKSEKAKVEKSKALQAEEEESEEASEDSGEDELTLISKR 261
+ + ++++++ KK + +S EKA++EKSK + + E + E++E + + I K
Sbjct: 1473 RAAQESKAKVEEGLKK-LEEESSKEKAELEKSKEMMKKLESTIESNETELKSSMETIRKS 1531
Query: 262 LNRIWKHRQSKFKGSGKAKGKYESSGQKKSSIREVTCFECKESGHYKSDCPKLKKDKKPK 321
++ QSK K S I E K+ KS KL+ + K
Sbjct: 1532 DEKL---EQSKKSAEEDIKNLQHEKSDLISRINESE----KDIEELKS---KLRIEAKSG 1581
Query: 322 KHFKTKKSLMVTFDE----SESEDVDSDGEVQGLMAIVKDKGA------ESKEAVDS--- 368
+T K + E + E+ +++ + +KDK A E KE + S
Sbjct: 1582 SELETVKQELNNAQEKIRINAEENTVLKSKLEDIERELKDKQAEIKSNQEEKELLTSRLK 1641
Query: 369 ------DSESEGDPDSDDENEVFA---SFSTSELKHALSDIMDKYNSLLSKHKKLKKNLS 419
DS + S++E S+L + KYN L++K + K++
Sbjct: 1642 ELEQELDSTQQKAQKSEEERRAEVRKFQVEKSQLDEKAMLLETKYNDLVNKEQAWKRDED 1701
Query: 420 AVSKTPSEHEKIISDLKNDNHALVNSNSVLKNQIAKLEE-------VIACDASDSKHESK 472
V KT + I L + L NS LK E V D ++K+ SK
Sbjct: 1702 TVKKTTDSQRQEIEKLAKELDNLKAENSKLKEANEDRSEIDDLMLLVTDLDEKNAKYRSK 1761
Score = 45.4 bits (106), Expect = 0.001
Identities = 92/460 (20%), Positives = 174/460 (37%), Gaps = 77/460 (16%)
Query: 56 KIPRSEMTAEQKKLYSQHHKARAILLSAISYEEYQKITDREFAKGIFDSLKMSHEGNKKV 115
K+ SE ++ K +H K I L + E Q++ +SL+ + E +K
Sbjct: 1104 KLETSEKALKEVKENEEHLKEEKIQLEKEATETKQQL----------NSLRANLESLEKE 1153
Query: 116 KESKALSLIQKYESFIMEPNESIEEMFSRFQLLVAGIRPLNKSYTTKDHVIRVIRCLPES 175
E A L +KYE I E S+ + + N+S K+
Sbjct: 1154 HEDLAAQL-KKYEEQIANKERQYNEEISQLNDEITSTQQENESIKKKND----------- 1201
Query: 176 WMPLVTSIELTRDVENM-SLEELISILKCHELKRSEMQDLRKKSIALKSKSEKAKVEKSK 234
EL +V+ M S E S LK+SE+ L + LK K+E + +
Sbjct: 1202 --------ELEGEVKAMKSTSEEQS-----NLKKSEIDALNLQIKELKKKNETNEASLLE 1248
Query: 235 ALQAEEEESEEASEDSGEDELTLISKRLNRIWKHRQSKFKGSGKAKGKYESSGQKKSSIR 294
++++ E E+ + E +DE K ++ + + K K S KY ++ I+
Sbjct: 1249 SIKSVESETVKIKE--LQDECNFKEKEVSEL----EDKLKASEDKNSKYLELQKESEKIK 1302
Query: 295 EVTCFECKESGHYKSDCPKLKKDKKPKKHFKTKKSLMVTFDESESEDVDSDGEVQGLMAI 354
E + E K K+ K K+ +++ S + E ++ + E
Sbjct: 1303 EELDAKTTE---LKIQLEKITNLSKAKEKSESELSRLKKTSSEERKNAEEQLEKLKNEIQ 1359
Query: 355 VKDKGAESKEAVDSDSESEGDPDSDD--------------ENEVFA-------------S 387
+K++ E + + ++ S + + ENE+ A S
Sbjct: 1360 IKNQAFEKERKLLNEGSSTITQEYSEKINTLEDELIRLQNENELKAKEIDNTRSELEKVS 1419
Query: 388 FSTSE-LKHALSDIMDKYNSLLSKHKKLKKNLSAVSKTPSEHEKIISDLKNDNHALVNSN 446
S E L+ + I + +LS K+ +N + ++++ + LK A S
Sbjct: 1420 LSNDELLEEKQNTIKSLQDEILSYKDKITRNDEKLLSIERDNKRDLESLKEQLRAAQESK 1479
Query: 447 SVLKNQIAKLEEVIACDASDSKHESKYEKSFQRFLAKSVD 486
+ ++ + KLEE ++S K E + K + L +++
Sbjct: 1480 AKVEEGLKKLEE----ESSKEKAELEKSKEMMKKLESTIE 1515
Score = 45.4 bits (106), Expect = 0.001
Identities = 63/310 (20%), Positives = 134/310 (42%), Gaps = 29/310 (9%)
Query: 191 NMSLEELISILKCHELKRSEMQDLRKKSIALKSKSE-------------KAKVEKSKALQ 237
N +LEE+ ++C+ L + E + + K+ + KS+ + K+ K +Q
Sbjct: 919 NENLEEMK--IQCNNLSK-EKEHISKELVEYKSRFQSHDNLVAKLTEKLKSLANNYKDMQ 975
Query: 238 AEEEESEEASEDSGEDELTLISKRLNRIWKHRQSKFKGSGKAKGKYESSGQKKSSIREVT 297
AE E +A E+S + +S N+I Q K + ++ Q K +I ++
Sbjct: 976 AENESLIKAVEESKNESSIQLSNLQNKIDSMSQEKENFQIERGSIEKNIEQLKKTISDLE 1035
Query: 298 CFECKESGHYKSDCPKLKKDKKPKKHFKTKKSLMVTFDESE---SEDVDSDGEVQGLMAI 354
+ KE KSD K + + + + ++ DE+ SE + E++ +A
Sbjct: 1036 --QTKEEIISKSDSSKDEYESQISLLKEKLETATTANDENVNKISELTKTREELEAELAA 1093
Query: 355 VKDKGAESKEAVDSDSESEGDPDSDDEN----EVFASFSTSELKHALSDIMDKYNSLLSK 410
K+ E + +++ ++ + ++E+ ++ +E K L+ + SL +
Sbjct: 1094 YKNLKNELETKLETSEKALKEVKENEEHLKEEKIQLEKEATETKQQLNSLRANLESLEKE 1153
Query: 411 HK----KLKKNLSAVSKTPSEHEKIISDLKNDNHALVNSNSVLKNQIAKLEEVIACDASD 466
H+ +LKK ++ ++ + IS L ++ + N +K + +LE + S
Sbjct: 1154 HEDLAAQLKKYEEQIANKERQYNEEISQLNDEITSTQQENESIKKKNDELEGEVKAMKST 1213
Query: 467 SKHESKYEKS 476
S+ +S +KS
Sbjct: 1214 SEEQSNLKKS 1223
Score = 43.9 bits (102), Expect = 0.004
Identities = 74/306 (24%), Positives = 137/306 (44%), Gaps = 35/306 (11%)
Query: 106 KMSHEGNKKVKE-SKALSLIQKYESFIMEPNESIEEMFSRFQLLVAGIRPLNKSYTTKDH 164
K+ E +K+ E K+ +++K ES I E NE+ E+ S + + L +S K
Sbjct: 1488 KLEEESSKEKAELEKSKEMMKKLESTI-ESNET--ELKSSMETIRKSDEKLEQS---KKS 1541
Query: 165 VIRVIRCLPESWMPLVTSI-ELTRDVENMSLEELISILKCHELKRSEMQDLRKKSIALKS 223
I+ L L++ I E +D+E + + I EL+ + Q+L ++
Sbjct: 1542 AEEDIKNLQHEKSDLISRINESEKDIEELKSKLRIEAKSGSELETVK-QELNNAQEKIRI 1600
Query: 224 KSEKAKVEKSKALQAEEEESEEASE-DSGEDELTLISKRLNRIWKHRQSKFKGSGKAKGK 282
+E+ V KSK E E ++ +E S ++E L++ RL + + S + + K++ +
Sbjct: 1601 NAEENTVLKSKLEDIERELKDKQAEIKSNQEEKELLTSRLKELEQELDSTQQKAQKSEEE 1660
Query: 283 YESSGQK----KSSIRE-VTCFECK------ESGHYKSDCPKLKK--DKKPKKHFKTKKS 329
+ +K KS + E E K + +K D +KK D + ++ K K
Sbjct: 1661 RRAEVRKFQVEKSQLDEKAMLLETKYNDLVNKEQAWKRDEDTVKKTTDSQRQEIEKLAKE 1720
Query: 330 L-MVTFDESESEDVDSD-GEVQGLMAIVKD---KGAESKEA-------VDSDSESEGDPD 377
L + + S+ ++ + D E+ LM +V D K A+ + + SD E + + D
Sbjct: 1721 LDNLKAENSKLKEANEDRSEIDDLMLLVTDLDEKNAKYRSKLKDLGVEISSDEEDDEEDD 1780
Query: 378 SDDENE 383
+DE E
Sbjct: 1781 EEDEEE 1786
>CYL2_BOVIN (Q28092) Cylicin II (Multiple-band polypeptide II)
Length = 488
Score = 50.8 bits (120), Expect = 3e-05
Identities = 79/332 (23%), Positives = 139/332 (41%), Gaps = 48/332 (14%)
Query: 178 PLVTSIELTRDVENMSLEELIS------ILKCHELKRSEMQDLRK--------KSIALKS 223
P V + +D + E ++S + K E K E +DL+K K A +S
Sbjct: 119 PKVKKSKEDKDKSDSEAESIVSKEKPRKLSKAKEEKPDEKKDLKKERKDSKKGKESATES 178
Query: 224 KSEKAKVEK-SKALQAEEEESEEASEDSGED--ELTLISKRLNRIWKHRQSKFKGSGKAK 280
+ EKA EK +K + ++ +E DSG + + SK+ + K ++S + G+ K
Sbjct: 179 EDEKAGAEKGAKKDRKGSKKGKETPSDSGSEKGDAKKDSKKSKKDSKGKESATESEGE-K 237
Query: 281 G---KYESSGQKKSSIREVTCFECK-ESGHYKSDCPKLKKDKKPKKHFKTKKSLMVTFDE 336
G K + G+K S + + E + E G K D DKK KK K K T E
Sbjct: 238 GDAKKDDKKGKKGSKKGKESATESEGEKGDAKKD------DKKGKKGSKKGKE-SATESE 290
Query: 337 SESEDVDSDGEVQGLMAIVKDKGAE-SKEAVDSDSESEGDPDSDDENEVFASFSTSELKH 395
E D D + KG + SK+ +S +ESEG+ +++ + + K
Sbjct: 291 GEKGDAKKDDK----------KGKKGSKKGKESATESEGEKGDAKKDDKKGKKGSKKGKE 340
Query: 396 ALSDIMDKYNSLLSKHKKLKKNLSAVSKTPSEHEKIISDLKNDNHALVNSNSVLKNQIAK 455
+ ++ + KK KK ++ S+ E D K D+ + +K
Sbjct: 341 SATESEGEKGDAKKDDKKGKKGSKKGKESDSKAEGDKGDAKKDDKK--------DKKGSK 392
Query: 456 LEEVIACDASDSKHESKYEKSFQRFLAKSVDR 487
+ A ++ K +SK +K+ ++ K+ ++
Sbjct: 393 KGKESATESEGEKKDSKKDKAGKKDPTKAGEK 424
Score = 40.8 bits (94), Expect = 0.030
Identities = 47/189 (24%), Positives = 77/189 (39%), Gaps = 28/189 (14%)
Query: 207 KRSEMQDLRKKSIALKSKSEK--AKVEKSKALQAEEEESEEASEDSGED----ELTLISK 260
K+ + + K A +S+ EK AK + K + ++ E A+E GE + K
Sbjct: 301 KKGKKGSKKGKESATESEGEKGDAKKDDKKGKKGSKKGKESATESEGEKGDAKKDDKKGK 360
Query: 261 RLNRIWKHRQSKFKGS-------------GKAKGKY---ESSGQKKSSIREVTCFECKES 304
+ ++ K SK +G G KGK ES G+KK S ++ +
Sbjct: 361 KGSKKGKESDSKAEGDKGDAKKDDKKDKKGSKKGKESATESEGEKKDSKKDKAGKKDPTK 420
Query: 305 GHYKSDCPKLKKDKKPKKHFKTKKSLMVTFDESESEDVDSDGEVQGLMAIVKDKGAESKE 364
K D K KKD K K K KK DE + + +S+ + KD ++K+
Sbjct: 421 AGEKGDESKDKKDAKKKDSKKEKK------DEKKPGEAESEPKDSAKKDAKKDAKKDAKK 474
Query: 365 AVDSDSESE 373
D++ +
Sbjct: 475 DAKKDAKKD 483
>YCO2_ARATH (Q9LME2) Hypothetical protein At1g22260
Length = 871
Score = 50.4 bits (119), Expect = 4e-05
Identities = 95/455 (20%), Positives = 183/455 (39%), Gaps = 90/455 (19%)
Query: 90 QKITDREFAKGIFDSLKMSHEGNKKVKESKALSLIQKYESFIMEPNESIEEMFSRFQLLV 149
++IT R+ K E + + SLI+K ++ I + S E + L
Sbjct: 172 EEITSRDKELEELKLEKQQKEMFYQTERCGTASLIEKKDAVITKLEASAAERKLNIENLN 231
Query: 150 AGIRPLNKSYTTKDHVIRVIRCLPESWMPLVTSIELTRDVENMSLEELIS----ILKCHE 205
+ + ++ TTK+ ++ + + E TS++L+ D E+L+S + K E
Sbjct: 232 SQLEKVHLELTTKEDEVKDLVSIQEKLEKEKTSVQLSAD---NCFEKLVSSEQEVKKLDE 288
Query: 206 LKR---SEMQDLRKKSIALKSKSEKAK---------VEKSKALQAEEEE----------- 242
L + +E+ +L KK++ K K +K ++K + L + +
Sbjct: 289 LVQYLVAELTELDKKNLTFKEKFDKLSGLYDTHIMLLQKDRDLALDRAQRSFDNLQGELF 348
Query: 243 ----SEEASEDSG-----------EDELTLISKRLNRIWKHRQSKFKGSGKAKGKYESSG 287
++EA E +G D+ +LIS+ Q+ K +AKG
Sbjct: 349 RVAATKEALESAGNELNEKIVELQNDKESLISQLSGLRCSTSQTIDKLESEAKGLVSKHA 408
Query: 288 QKKSSIREVTCFECKESGHYKSDCPKLKKDKKPKKHFKTK-------------------- 327
+S+I ++ KE + K +DKK + K
Sbjct: 409 DAESAISQL-----KEEMETLLESVKTSEDKKQELSLKLSSLEMESKEKCEKLQADAQRQ 463
Query: 328 -KSLMVTFDESESEDVDSD---GEVQGLMAIVKDKGAESKEAVDSDSESEGDPDSDDENE 383
+ L ESES + +D EV L ++++KG + +++ + D E
Sbjct: 464 VEELETLQKESESHQLQADLLAKEVNQLQTVIEEKGHVILQCNENEKQLNQQIIKDKE-- 521
Query: 384 VFASFSTSELKHALSDIMDKYNSLL-SKHKKLKKNLSAVSKTPSEHEKIISDLKN----D 438
+T+E K L++ +Y+ +L SK +L ++L +S+ +++ I++++ +
Sbjct: 522 ---LLATAETK--LAEAKKQYDLMLESKQLELSRHLKELSQ---RNDQAINEIRRKYDVE 573
Query: 439 NHALVNSNSVLKNQIAK-LEEVIACDASDSKHESK 472
H ++NS +I K L + SD K ESK
Sbjct: 574 KHEIINSEKDKVEKIIKDLSNKFDKELSDCKEESK 608
>NUCL_XENLA (P20397) Nucleolin (Protein C23)
Length = 650
Score = 50.4 bits (119), Expect = 4e-05
Identities = 43/167 (25%), Positives = 74/167 (43%), Gaps = 8/167 (4%)
Query: 217 KSIALKSKSEKAKVEKSKALQAEEEESEEASEDSGEDELTLISKRLNRIWKHRQSKFKGS 276
K ++K +KA K ++ EEE + +DS ++E+ + K K K
Sbjct: 5 KGAKTQAKPKKAAPPPPKDMEDSEEEEDMEEDDSSDEEVEVPVK------KTPAKKTATP 58
Query: 277 GKAK-GKYESSGQKKSS-IREVTCFECKESGHYKSDCPKLKKDKKPKKHFKTKKSLMVTF 334
KA GK + G+K ++ + + +ES + D + +D+KP K+ K +
Sbjct: 59 AKATPGKAATPGKKGATPAKNGKQAKKQESEEEEDDSDEEAEDQKPIKNKPVAKKAVAKK 118
Query: 335 DESESEDVDSDGEVQGLMAIVKDKGAESKEAVDSDSESEGDPDSDDE 381
+ESE +D D D + K A+ +SE E D +S+DE
Sbjct: 119 EESEEDDDDEDESEEEKAVAKKPTPAKKPAGKKQESEEEDDEESEDE 165
>NFM_BOVIN (O77788) Neurofilament triplet M protein (160 kDa
neurofilament protein) (Neurofilament medium
polypeptide) (NF-M) (Fragment)
Length = 810
Score = 50.4 bits (119), Expect = 4e-05
Identities = 53/215 (24%), Positives = 100/215 (45%), Gaps = 17/215 (7%)
Query: 215 RKKSIALKSKSEKAKVEKSKALQAEEEESEEASEDSGEDELTLISKRLNRIWKHRQSKFK 274
R+ SIA+ SK +K KVE K + E E EDE + + + L I + K
Sbjct: 311 RQPSIAISSKIQKTKVEAPKLKVQHKFVEEIIEETKVEDEKSEMEEALTAITEELAVSVK 370
Query: 275 GSGKAKGKYESSGQKKSSIREVTCFECKESGHYKSDCPKLKKDKKPKKHFKTKKSLMVTF 334
K + + E +K+ + EV + K+ P+LK+++ K+ + ++
Sbjct: 371 EEVKEE-EAEEKEEKEEAEEEVV---AAKKSPVKATAPELKEEEGEKEEEEGQEE----- 421
Query: 335 DESESEDVDSDGEVQGLMAIVKDKGAESKEAVDSDSESEGDPDSDDENEVFASFSTSE-- 392
+E E E SD +G + +G+ KE + + E EG+ +++ E E A+ + E
Sbjct: 422 EEEEEEAAKSDQAEEGGS---EKEGSSEKE--EGEQEEEGETEAEGEGEEAAAEAKEEKK 476
Query: 393 LKHALSDIMDKYN-SLLSKHKKLKKNLSAVSKTPS 426
++ ++ K + +K +K +K S V+K+P+
Sbjct: 477 MEEKAEEVAPKEELAAEAKVEKPEKAKSPVAKSPT 511
Score = 38.9 bits (89), Expect = 0.12
Identities = 63/260 (24%), Positives = 108/260 (41%), Gaps = 58/260 (22%)
Query: 208 RSEMQDLRKKSIALKSKSE-KAKVEKSKALQAEEEESEEASEDSGEDELTLISKRLNRIW 266
+S +++++ K+ A K E K KVE+ K +A+E EE +E E + K
Sbjct: 578 KSPVEEVKPKAEAGAEKGEQKEKVEEEKK-EAKESPKEEKAEKKEEKPKDVPEK------ 630
Query: 267 KHRQSKFKGSGKAKGKYESSGQKKSSIREVTCFECKESGHYKSDCPKLKKDKKPKKHFKT 326
K +S K K + + K S +E K+++KP++ K
Sbjct: 631 KKAESPVKAESPVKEEVPAKPVKVSPEKEA------------------KEEEKPQEKEKE 672
Query: 327 KKSLMVT-------FDESESEDVDSDGEVQGLMAIVKDKGAESKEAVDSDSESEG----- 374
K+ + ES ED+ +GEV+G K++ E+KE E +G
Sbjct: 673 KEKVEEVGGKEEGGLKESRKEDIAINGEVEG-----KEEEQETKEKGSGGEEEKGVVTNG 727
Query: 375 -DPDSDDENEVFASFSTSELKHALSDIMDKYNSL--LSKHKKLKKNLSAVSKTPSEH--- 428
D DE + SE K ++ +++K S K + K+++ K EH
Sbjct: 728 LDVSPGDEKK---GGDKSEEKVVVTKMVEKITSEGGDGATKYITKSVTVTQKV-EEHEET 783
Query: 429 --EKIISDLKND---NHALV 443
EK++S K + +HA+V
Sbjct: 784 FEEKLVSTKKVEKVTSHAIV 803
Score = 33.1 bits (74), Expect = 6.3
Identities = 42/206 (20%), Positives = 80/206 (38%), Gaps = 22/206 (10%)
Query: 185 LTRDVENMSLEELISILKCHELKRSEMQDLRKKSIALKSKSEKAKVEKSKALQAEEEESE 244
LT E +++ + + ++ E ++ ++ +A K KA + K + E+EE E
Sbjct: 358 LTAITEELAVSVKEEVKEEEAEEKEEKEEAEEEVVAAKKSPVKATAPELKEEEGEKEEEE 417
Query: 245 EASEDSGEDELTLISKRLNRIWKHRQSKFKGSGKAKGKYESSGQKKSSIR-EVTCFECKE 303
E+ E+E + + ++ + K S K +G+ E G+ ++ E E KE
Sbjct: 418 GQEEEEEEEE----AAKSDQAEEGGSEKEGSSEKEEGEQEEEGETEAEGEGEEAAAEAKE 473
Query: 304 SGHY---------------KSDCPKLKKDKKPKKHFKTKKSLMVTFDESESEDVDSD--G 346
++ K +K K P T KS E++S + S
Sbjct: 474 EKKMEEKAEEVAPKEELAAEAKVEKPEKAKSPVAKSPTTKSPTAKSPEAKSPEAKSPTAK 533
Query: 347 EVQGLMAIVKDKGAESKEAVDSDSES 372
+ K A+S EA +++S
Sbjct: 534 SPTAKSPVAKSPTAKSPEAKSPEAKS 559
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.335 0.145 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 175,070,355
Number of Sequences: 164201
Number of extensions: 7206862
Number of successful extensions: 48856
Number of sequences better than 10.0: 583
Number of HSP's better than 10.0 without gapping: 75
Number of HSP's successfully gapped in prelim test: 532
Number of HSP's that attempted gapping in prelim test: 44781
Number of HSP's gapped (non-prelim): 2848
length of query: 1610
length of database: 59,974,054
effective HSP length: 124
effective length of query: 1486
effective length of database: 39,613,130
effective search space: 58865111180
effective search space used: 58865111180
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0134.9


