

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0134.4
(311 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
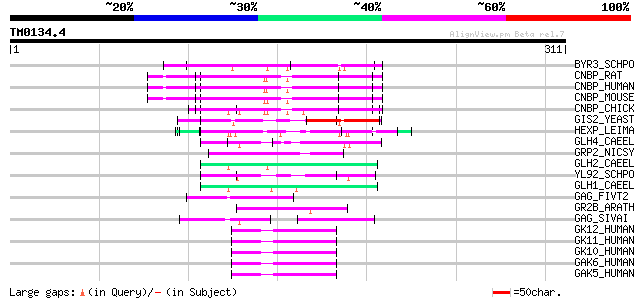
Score E
Sequences producing significant alignments: (bits) Value
BYR3_SCHPO (P36627) Cellular nucleic acid binding protein homolog 78 3e-14
CNBP_RAT (P62634) Cellular nucleic acid binding protein (CNBP) (... 77 4e-14
CNBP_HUMAN (P62633) Cellular nucleic acid binding protein (CNBP)... 77 4e-14
CNBP_MOUSE (P53996) Cellular nucleic acid binding protein (CNBP)... 76 1e-13
CNBP_CHICK (O42395) Cellular nucleic acid binding protein (CNBP) 73 1e-12
GIS2_YEAST (P53849) Zinc-finger protein GIS2 70 5e-12
HEXP_LEIMA (Q04832) DNA-binding protein HEXBP (Hexamer-binding p... 62 1e-09
GLH4_CAEEL (O76743) ATP-dependent RNA helicase glh-4 (EC 3.6.1.-... 55 2e-07
GRP2_NICSY (P27484) Glycine-rich protein 2 51 4e-06
GLH2_CAEEL (Q966L9) ATP-dependent RNA helicase glh-2 (EC 3.6.1.-... 51 4e-06
YL92_SCHPO (Q9HFF2) Hypothetical protein C683.02c in chromosome I 49 2e-05
GLH1_CAEEL (P34689) ATP-dependent RNA helicase glh-1 (EC 3.6.1.-... 48 4e-05
GAG_FIVT2 (P31821) Gag polyprotein [Contains: Core protein p15; ... 48 4e-05
GR2B_ARATH (Q38896) Glycine-rich protein 2b (AtGRP2b) 47 8e-05
GAG_SIVAI (Q02843) Gag polyprotein [Contains: Core protein p17; ... 47 8e-05
GK12_HUMAN (P63130) HERV-K_1q22 provirus ancestral Gag polyprote... 45 2e-04
GK11_HUMAN (P63145) HERV-K_22q11.21 provirus ancestral Gag polyp... 45 2e-04
GK10_HUMAN (P87889) HERV-K_5q33.3 provirus ancestral Gag polypro... 45 2e-04
GAK6_HUMAN (P62685) HERV-K_8p23.1 provirus ancestral Gag polypro... 45 2e-04
GAK5_HUMAN (P62684) HERV-K_19p13.11 provirus ancestral Gag polyp... 45 2e-04
>BYR3_SCHPO (P36627) Cellular nucleic acid binding protein homolog
Length = 179
Score = 77.8 bits (190), Expect = 3e-14
Identities = 42/140 (30%), Positives = 64/140 (45%), Gaps = 17/140 (12%)
Query: 87 FKSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLG-----QGPRCFNCNQIGHLAVNC 141
+ + T + E TC+ CG GH C QG C+ C ++GH+A +C
Sbjct: 39 YNCNQTGHKASECTEPQQEKTCYACGTAGHLVRDCPSSPNPRQGAECYKCGRVGHIARDC 98
Query: 142 KT------FQAGPSDNKMK----GKFPAIARGTEKQVSCYRCGEIGHFASRC--SATGPR 189
+T + G + M G + AR V CY CG+IGH + C ++ G
Sbjct: 99 RTNGQQSGGRFGGHRSNMNCYACGSYGHQARDCTMGVKCYSCGKIGHRSFECQQASDGQL 158
Query: 190 CFNCHRVGHLAVVCKKPKVE 209
C+ C++ GH+AV C P +E
Sbjct: 159 CYKCNQPGHIAVNCTSPVIE 178
Score = 73.2 bits (178), Expect = 8e-13
Identities = 36/110 (32%), Positives = 50/110 (44%), Gaps = 26/110 (23%)
Query: 100 QSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPA 159
Q+ P C+ CG+NGH A C +G C+NCNQ GH A C
Sbjct: 11 QTTRPGPRCYNCGENGHQARECT-KGSICYNCNQTGHKASECTE---------------- 53
Query: 160 IARGTEKQVSCYRCGEIGHFASRCSAT-----GPRCFNCHRVGHLAVVCK 204
+++ +CY CG GH C ++ G C+ C RVGH+A C+
Sbjct: 54 ----PQQEKTCYACGTAGHLVRDCPSSPNPRQGAECYKCGRVGHIARDCR 99
Score = 48.5 bits (114), Expect = 2e-05
Identities = 21/52 (40%), Positives = 29/52 (55%), Gaps = 1/52 (1%)
Query: 158 PAIARGTEKQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVE 209
P + + T CY CGE GH A C+ G C+NC++ GH A C +P+ E
Sbjct: 7 PTVPQTTRPGPRCYNCGENGHQARECT-KGSICYNCNQTGHKASECTEPQQE 57
>CNBP_RAT (P62634) Cellular nucleic acid binding protein (CNBP)
(Zinc finger protein 9)
Length = 177
Score = 77.4 bits (189), Expect = 4e-14
Identities = 49/149 (32%), Positives = 67/149 (44%), Gaps = 25/149 (16%)
Query: 78 KGRGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHL 137
+GRG R + T+ G + S+ C+RCG++GH A C Q C+NC + GH+
Sbjct: 25 RGRGMRSRG-RGGFTSDRGFQFVSSSLPDICYRCGESGHLAKDCDLQEDACYNCGRGGHI 83
Query: 138 AVNCK---------TFQAGPSDNKMK-------------GKFPAIARGTEKQVSCYRCGE 175
A +CK + G + + G+F I + K V CYRCGE
Sbjct: 84 AKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTK-VKCYRCGE 142
Query: 176 IGHFASRCSATGP-RCFNCHRVGHLAVVC 203
GH A CS T C+ C GHLA C
Sbjct: 143 TGHVAINCSKTSEVNCYRCGESGHLAREC 171
Score = 65.1 bits (157), Expect = 2e-10
Identities = 36/102 (35%), Positives = 45/102 (43%), Gaps = 14/102 (13%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
CF+CG++GH+A C G R G F + + P I
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGR-----GGFTSDRGFQFVSSSLPDI------- 53
Query: 168 VSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVE 209
CYRCGE GH A C C+NC R GH+A CK+PK E
Sbjct: 54 --CYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPKRE 93
Score = 59.3 bits (142), Expect = 1e-08
Identities = 29/84 (34%), Positives = 43/84 (50%), Gaps = 10/84 (11%)
Query: 105 EVTCFRCGKNGHYANAC-LGQGPRCFNCNQIGHLAVNC---KTFQAGPSDNKMKGKFPAI 160
E C+ CGK GH A C +C++C + GH+ +C K ++ G + + AI
Sbjct: 95 EQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTKVKCYRCGETGHV------AI 148
Query: 161 ARGTEKQVSCYRCGEIGHFASRCS 184
+V+CYRCGE GH A C+
Sbjct: 149 NCSKTSEVNCYRCGESGHLARECT 172
>CNBP_HUMAN (P62633) Cellular nucleic acid binding protein (CNBP)
(Zinc finger protein 9)
Length = 177
Score = 77.4 bits (189), Expect = 4e-14
Identities = 49/149 (32%), Positives = 67/149 (44%), Gaps = 25/149 (16%)
Query: 78 KGRGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHL 137
+GRG R + T+ G + S+ C+RCG++GH A C Q C+NC + GH+
Sbjct: 25 RGRGMRSRG-RGGFTSDRGFQFVSSSLPDICYRCGESGHLAKDCDLQEDACYNCGRGGHI 83
Query: 138 AVNCK---------TFQAGPSDNKMK-------------GKFPAIARGTEKQVSCYRCGE 175
A +CK + G + + G+F I + K V CYRCGE
Sbjct: 84 AKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTK-VKCYRCGE 142
Query: 176 IGHFASRCSATGP-RCFNCHRVGHLAVVC 203
GH A CS T C+ C GHLA C
Sbjct: 143 TGHVAINCSKTSEVNCYRCGESGHLAREC 171
Score = 65.1 bits (157), Expect = 2e-10
Identities = 36/102 (35%), Positives = 45/102 (43%), Gaps = 14/102 (13%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
CF+CG++GH+A C G R G F + + P I
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGR-----GGFTSDRGFQFVSSSLPDI------- 53
Query: 168 VSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVE 209
CYRCGE GH A C C+NC R GH+A CK+PK E
Sbjct: 54 --CYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPKRE 93
Score = 59.3 bits (142), Expect = 1e-08
Identities = 29/84 (34%), Positives = 43/84 (50%), Gaps = 10/84 (11%)
Query: 105 EVTCFRCGKNGHYANAC-LGQGPRCFNCNQIGHLAVNC---KTFQAGPSDNKMKGKFPAI 160
E C+ CGK GH A C +C++C + GH+ +C K ++ G + + AI
Sbjct: 95 EQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTKVKCYRCGETGHV------AI 148
Query: 161 ARGTEKQVSCYRCGEIGHFASRCS 184
+V+CYRCGE GH A C+
Sbjct: 149 NCSKTSEVNCYRCGESGHLARECT 172
>CNBP_MOUSE (P53996) Cellular nucleic acid binding protein (CNBP)
(Zinc finger protein 9)
Length = 178
Score = 75.9 bits (185), Expect = 1e-13
Identities = 49/150 (32%), Positives = 68/150 (44%), Gaps = 26/150 (17%)
Query: 78 KGRGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANAC-LGQGPRCFNCNQIGH 136
+GRG R + T+ G + S+ C+RCG++GH A C L + C+NC + GH
Sbjct: 25 RGRGMRSRG-RGGFTSDRGFQFVSSSLPDICYRCGESGHLAKDCDLQEDEACYNCGRGGH 83
Query: 137 LAVNCKT---------FQAGPSDNKMK-------------GKFPAIARGTEKQVSCYRCG 174
+A +CK + G + + G+F I + K V CYRCG
Sbjct: 84 IAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTK-VKCYRCG 142
Query: 175 EIGHFASRCSATGP-RCFNCHRVGHLAVVC 203
E GH A CS T C+ C GHLA C
Sbjct: 143 ETGHVAINCSKTSEVNCYRCGESGHLAREC 172
Score = 60.8 bits (146), Expect = 4e-09
Identities = 36/103 (34%), Positives = 45/103 (42%), Gaps = 15/103 (14%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
CF+CG++GH+A C G R G F + + P I
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGR-----GGFTSDRGFQFVSSSLPDI------- 53
Query: 168 VSCYRCGEIGHFASRCS-ATGPRCFNCHRVGHLAVVCKKPKVE 209
CYRCGE GH A C C+NC R GH+A CK+PK E
Sbjct: 54 --CYRCGESGHLAKDCDLQEDEACYNCGRGGHIAKDCKEPKRE 94
Score = 59.3 bits (142), Expect = 1e-08
Identities = 29/84 (34%), Positives = 43/84 (50%), Gaps = 10/84 (11%)
Query: 105 EVTCFRCGKNGHYANAC-LGQGPRCFNCNQIGHLAVNC---KTFQAGPSDNKMKGKFPAI 160
E C+ CGK GH A C +C++C + GH+ +C K ++ G + + AI
Sbjct: 96 EQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTKVKCYRCGETGHV------AI 149
Query: 161 ARGTEKQVSCYRCGEIGHFASRCS 184
+V+CYRCGE GH A C+
Sbjct: 150 NCSKTSEVNCYRCGESGHLARECT 173
>CNBP_CHICK (O42395) Cellular nucleic acid binding protein (CNBP)
Length = 172
Score = 72.8 bits (177), Expect = 1e-12
Identities = 44/127 (34%), Positives = 61/127 (47%), Gaps = 26/127 (20%)
Query: 101 SAHPEVTCFRCGKNGHYANAC-LGQGPRCFNCNQIGHLAVNCKT---------FQAGPSD 150
S+ P++ C+RCG++GH A C L + C+NC + GH+A +CK + G
Sbjct: 42 SSLPDI-CYRCGESGHLAKDCDLQEDKACYNCGRGGHIAKDCKEPKREREQCCYNCGKPG 100
Query: 151 NKMK-------------GKFPAIARGTEKQVSCYRCGEIGHFASRCSATGP-RCFNCHRV 196
+ + G+F I + K V CYRCGE GH A CS T C+ C
Sbjct: 101 HLARDCDHADEQKCYSCGEFGHIQKDCTK-VKCYRCGETGHVAINCSKTSEVNCYRCGES 159
Query: 197 GHLAVVC 203
GHLA C
Sbjct: 160 GHLAREC 166
Score = 62.0 bits (149), Expect = 2e-09
Identities = 39/128 (30%), Positives = 56/128 (43%), Gaps = 30/128 (23%)
Query: 108 CFRCGKNGHYANAC-----LGQGPR-----------------CFNCNQIGHLAVNCKTFQ 145
CF+CG+ GH+A C G+G R C+ C + GHLA +C +
Sbjct: 6 CFKCGRTGHWARECPTGIGRGRGMRSRGRAGFQFMSSSLPDICYRCGESGHLAKDCDLQE 65
Query: 146 AGPSDNKMKGKFPAIARG-----TEKQVSCYRCGEIGHFASRCS-ATGPRCFNCHRVGHL 199
N +G IA+ E++ CY CG+ GH A C A +C++C GH+
Sbjct: 66 DKACYNCGRGGH--IAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHI 123
Query: 200 AVVCKKPK 207
C K K
Sbjct: 124 QKDCTKVK 131
Score = 61.6 bits (148), Expect = 2e-09
Identities = 31/83 (37%), Positives = 40/83 (47%), Gaps = 1/83 (1%)
Query: 128 CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATG 187
CF C + GH A C T + +G+ + CYRCGE GH A C
Sbjct: 6 CFKCGRTGHWARECPTGIGRGRGMRSRGRAGFQFMSSSLPDICYRCGESGHLAKDCDLQE 65
Query: 188 PR-CFNCHRVGHLAVVCKKPKVE 209
+ C+NC R GH+A CK+PK E
Sbjct: 66 DKACYNCGRGGHIAKDCKEPKRE 88
Score = 59.3 bits (142), Expect = 1e-08
Identities = 29/84 (34%), Positives = 43/84 (50%), Gaps = 10/84 (11%)
Query: 105 EVTCFRCGKNGHYANAC-LGQGPRCFNCNQIGHLAVNC---KTFQAGPSDNKMKGKFPAI 160
E C+ CGK GH A C +C++C + GH+ +C K ++ G + + AI
Sbjct: 90 EQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTKVKCYRCGETGHV------AI 143
Query: 161 ARGTEKQVSCYRCGEIGHFASRCS 184
+V+CYRCGE GH A C+
Sbjct: 144 NCSKTSEVNCYRCGESGHLARECT 167
Score = 34.7 bits (78), Expect = 0.31
Identities = 20/74 (27%), Positives = 27/74 (36%), Gaps = 22/74 (29%)
Query: 170 CYRCGEIGHFASRC----------------------SATGPRCFNCHRVGHLAVVCKKPK 207
C++CG GH+A C S+ C+ C GHLA C +
Sbjct: 6 CFKCGRTGHWARECPTGIGRGRGMRSRGRAGFQFMSSSLPDICYRCGESGHLAKDCDLQE 65
Query: 208 VEPSVNTARGKHPA 221
+ N RG H A
Sbjct: 66 DKACYNCGRGGHIA 79
>GIS2_YEAST (P53849) Zinc-finger protein GIS2
Length = 153
Score = 70.5 bits (171), Expect = 5e-12
Identities = 37/100 (37%), Positives = 49/100 (49%), Gaps = 21/100 (21%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
C+ CGK GH A C + C+NCN+ GH+ +C P + K
Sbjct: 6 CYVCGKIGHLAEDCDSER-LCYNCNKPGHVQTDCTM----PRTVEFK------------- 47
Query: 168 VSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPK 207
CY CGE GH S C T RCFNC++ GH++ C +PK
Sbjct: 48 -QCYNCGETGHVRSEC--TVQRCFNCNQTGHISRECPEPK 84
Score = 68.9 bits (167), Expect = 1e-11
Identities = 34/105 (32%), Positives = 52/105 (49%), Gaps = 21/105 (20%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
C+ CG+ GH + C Q RCFNCNQ GH++ C + K +F +
Sbjct: 49 CYNCGETGHVRSECTVQ--RCFNCNQTGHISRECP-------EPKKTSRF--------SK 91
Query: 168 VSCYRCGEIGHFASRC----SATGPRCFNCHRVGHLAVVCKKPKV 208
VSCY+CG H A C +G +C+ C + GH++ C+ ++
Sbjct: 92 VSCYKCGGPNHMAKDCMKEDGISGLKCYTCGQAGHMSRDCQNDRL 136
Score = 47.4 bits (111), Expect = 5e-05
Identities = 23/93 (24%), Positives = 38/93 (40%), Gaps = 27/93 (29%)
Query: 95 PGARSQSAHPEVTCFRCGKNGHYANACLGQ----GPRCFNCNQIGHLAVNCKTFQAGPSD 150
P + S +V+C++CG H A C+ + G +C+ C Q GH++ +C+ +
Sbjct: 81 PEPKKTSRFSKVSCYKCGGPNHMAKDCMKEDGISGLKCYTCGQAGHMSRDCQNDRL---- 136
Query: 151 NKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC 183
CY C E GH + C
Sbjct: 137 -------------------CYNCNETGHISKDC 150
Score = 43.5 bits (101), Expect = 7e-04
Identities = 17/41 (41%), Positives = 25/41 (60%), Gaps = 1/41 (2%)
Query: 167 QVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPK 207
Q +CY CG+IGH A C + C+NC++ GH+ C P+
Sbjct: 3 QKACYVCGKIGHLAEDCDSE-RLCYNCNKPGHVQTDCTMPR 42
>HEXP_LEIMA (Q04832) DNA-binding protein HEXBP (Hexamer-binding
protein)
Length = 271
Score = 62.4 bits (150), Expect = 1e-09
Identities = 38/125 (30%), Positives = 54/125 (42%), Gaps = 30/125 (24%)
Query: 107 TCFRCGKNGHYANAC-------LGQGPR-CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFP 158
TC++CG GH + C G G R C+ C GH++ +C Q G S
Sbjct: 141 TCYKCGDAGHISRDCPNGQGGYSGAGDRTCYKCGDAGHISRDCPNGQGGYS--------- 191
Query: 159 AIARGTEKQVSCYRCGEIGHFASRCSATGP------RCFNCHRVGHLAVVCKKPKVEPSV 212
G K CY+CGE GH + C + G C+ C + GH++ C P+ S
Sbjct: 192 --GAGDRK---CYKCGESGHMSRECPSAGSTGSGDRACYKCGKPGHISREC--PEAGGSY 244
Query: 213 NTARG 217
+RG
Sbjct: 245 GGSRG 249
Score = 61.6 bits (148), Expect = 2e-09
Identities = 39/146 (26%), Positives = 59/146 (39%), Gaps = 34/146 (23%)
Query: 94 TPGARSQSAHPEVTCFRCGKNGHYANACL-------GQGPRCFNCNQIGHLAVNCKTFQA 146
T + +C CGK GHYA C + CF C + GH++ C
Sbjct: 4 TEDVKRPRTESSTSCRNCGKEGHYARECPEADSKGDERSTTCFRCGEEGHMSREC----- 58
Query: 147 GPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC-------SATGPRCFNCHRVGHL 199
P++ + G ++C+RCGE GH + C +A G C+ C + GHL
Sbjct: 59 -PNEAR---------SGAAGAMTCFRCGEAGHMSRDCPNSAKPGAAKGFECYKCGQEGHL 108
Query: 200 AVVCKKPKVEPSVNTARGKHPAARGK 225
+ C S +RG + RG+
Sbjct: 109 SRDCPS-----SQGGSRGGYGQKRGR 129
Score = 58.9 bits (141), Expect = 2e-08
Identities = 40/153 (26%), Positives = 58/153 (37%), Gaps = 45/153 (29%)
Query: 95 PGARSQSAHPEVTCFRCGKNGHYANAC--------------------------------- 121
P A S+ TCFRCG+ GH + C
Sbjct: 32 PEADSKGDERSTTCFRCGEEGHMSRECPNEARSGAAGAMTCFRCGEAGHMSRDCPNSAKP 91
Query: 122 -LGQGPRCFNCNQIGHLAVNCKTFQAGPSD--NKMKGKFPAIARGTEKQVSCYRCGEIGH 178
+G C+ C Q GHL+ +C + Q G + +G+ A G +CY+CG+ GH
Sbjct: 92 GAAKGFECYKCGQEGHLSRDCPSSQGGSRGGYGQKRGRSGA-QGGYSGDRTCYKCGDAGH 150
Query: 179 FASRC-------SATGPR-CFNCHRVGHLAVVC 203
+ C S G R C+ C GH++ C
Sbjct: 151 ISRDCPNGQGGYSGAGDRTCYKCGDAGHISRDC 183
Score = 53.1 bits (126), Expect = 8e-07
Identities = 31/127 (24%), Positives = 51/127 (39%), Gaps = 35/127 (27%)
Query: 96 GARSQSAHPEVTCFRCGKNGHYANAC-LGQG-------PRCFNCNQIGHLAVNCKTFQAG 147
G S + TC++CG GH + C GQG +C+ C + GH++ C + +
Sbjct: 158 GQGGYSGAGDRTCYKCGDAGHISRDCPNGQGGYSGAGDRKCYKCGESGHMSRECPSAGST 217
Query: 148 PSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATG-----------PRCFNCHRV 196
S ++ +CY+CG+ GH + C G C+ C
Sbjct: 218 GSGDR----------------ACYKCGKPGHISRECPEAGGSYGGSRGGGDRTCYKCGEA 261
Query: 197 GHLAVVC 203
GH++ C
Sbjct: 262 GHISRDC 268
Score = 51.2 bits (121), Expect = 3e-06
Identities = 27/85 (31%), Positives = 45/85 (52%), Gaps = 17/85 (20%)
Query: 108 CFRCGKNGHYANAC-----LGQGPR-CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIA 161
C++CG++GH + C G G R C+ C + GH++ C +AG S +G
Sbjct: 198 CYKCGESGHMSRECPSAGSTGSGDRACYKCGKPGHISRECP--EAGGSYGGSRG------ 249
Query: 162 RGTEKQVSCYRCGEIGHFASRCSAT 186
G ++ +CY+CGE GH + C ++
Sbjct: 250 -GGDR--TCYKCGEAGHISRDCPSS 271
>GLH4_CAEEL (O76743) ATP-dependent RNA helicase glh-4 (EC 3.6.1.-)
(Germline helicase-4)
Length = 1156
Score = 55.1 bits (131), Expect = 2e-07
Identities = 37/108 (34%), Positives = 46/108 (42%), Gaps = 27/108 (25%)
Query: 108 CFRCGKNGHYANAC-LGQGPR--CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGT 164
C CG+ GH + C + PR C NC Q+GH A +C P + RG
Sbjct: 572 CHNCGEEGHISKECDKPKVPRFPCRNCEQLGHFASDCDQ--------------PRVPRGP 617
Query: 165 EKQVSCYRCGEIGHFASRCSAT----GPRCFNCHRVGHLAVVCKKPKV 208
C CG GHFA C GP C NC + GH A C+ +V
Sbjct: 618 -----CRNCGIEGHFAVDCDQPKVPRGP-CRNCGQEGHFAKDCQNERV 659
Score = 51.2 bits (121), Expect = 3e-06
Identities = 33/91 (36%), Positives = 41/91 (44%), Gaps = 25/91 (27%)
Query: 123 GQGPR-CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFAS 181
G+ PR C NC + GH++ C D +FP C C ++GHFAS
Sbjct: 566 GERPRGCHNCGEEGHISKEC--------DKPKVPRFP-----------CRNCEQLGHFAS 606
Query: 182 RCSAT----GPRCFNCHRVGHLAVVCKKPKV 208
C GP C NC GH AV C +PKV
Sbjct: 607 DCDQPRVPRGP-CRNCGIEGHFAVDCDQPKV 636
Score = 45.1 bits (105), Expect = 2e-04
Identities = 25/64 (39%), Positives = 35/64 (54%), Gaps = 8/64 (12%)
Query: 148 PSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATG-PR--CFNCHRVGHLAVVCK 204
PSDN+ +G + G E+ C+ CGE GH + C PR C NC ++GH A C
Sbjct: 555 PSDNQ-RGNWD----GGERPRGCHNCGEEGHISKECDKPKVPRFPCRNCEQLGHFASDCD 609
Query: 205 KPKV 208
+P+V
Sbjct: 610 QPRV 613
Score = 41.6 bits (96), Expect = 0.003
Identities = 27/84 (32%), Positives = 36/84 (42%), Gaps = 21/84 (25%)
Query: 104 PEVTCFRCGKNGHYANAC----LGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPA 159
P C CG GH+A C + +GP C NC Q GH A +C+ + +M+ P
Sbjct: 614 PRGPCRNCGIEGHFAVDCDQPKVPRGP-CRNCGQEGHFAKDCQNERV-----RMEPTEP- 666
Query: 160 IARGTEKQVSCYRCGEIGHFASRC 183
C RC E GH+ C
Sbjct: 667 ----------CRRCAEEGHWGYEC 680
Score = 32.7 bits (73), Expect = 1.2
Identities = 15/46 (32%), Positives = 19/46 (40%), Gaps = 6/46 (13%)
Query: 104 PEVTCFRCGKNGHYANACLGQGPR------CFNCNQIGHLAVNCKT 143
P C CG+ GH+A C + R C C + GH C T
Sbjct: 637 PRGPCRNCGQEGHFAKDCQNERVRMEPTEPCRRCAEEGHWGYECPT 682
>GRP2_NICSY (P27484) Glycine-rich protein 2
Length = 214
Score = 50.8 bits (120), Expect = 4e-06
Identities = 26/76 (34%), Positives = 32/76 (41%), Gaps = 5/76 (6%)
Query: 112 GKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCY 171
G G Y G G CF C + GH A +C G + G G CY
Sbjct: 143 GGGGGYGGGGSGGGSGCFKCGESGHFARDCSQSGGGGGGGRFGGGGGGGGGG-----GCY 197
Query: 172 RCGEIGHFASRCSATG 187
+CGE GHFA C++ G
Sbjct: 198 KCGEDGHFARECTSGG 213
>GLH2_CAEEL (Q966L9) ATP-dependent RNA helicase glh-2 (EC 3.6.1.-)
(Germline helicase-2)
Length = 974
Score = 50.8 bits (120), Expect = 4e-06
Identities = 33/119 (27%), Positives = 45/119 (37%), Gaps = 20/119 (16%)
Query: 108 CFRCGKNGHYANAC----LGQGPR-CFNCNQIGHLAVNCKT--------------FQAGP 148
CF C + GH +N C + PR C+NC Q GH + +C F G
Sbjct: 373 CFNCQQPGHRSNDCPEPKKEREPRVCYNCQQPGHNSRDCPEERKPREGRNGFTSGFGGGN 432
Query: 149 SDNKMKGKFPAIARGTEK-QVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKP 206
G E+ + C+ C GH ++ C CFNC GH + C P
Sbjct: 433 DGGFGGGNAEGFGNNEERGPMKCFNCKGEGHRSAECPEPPRGCFNCGEQGHRSNECPNP 491
Score = 37.0 bits (84), Expect = 0.063
Identities = 36/134 (26%), Positives = 46/134 (33%), Gaps = 32/134 (23%)
Query: 108 CFRCGKNGHYANAC----LGQGPR-CFNCNQIGHLAVNC----------KTFQAGPSD-N 151
CF C + GH +N C + PR C+NC Q GH + +C F G S
Sbjct: 259 CFNCQQPGHRSNDCPEPKKEREPRVCYNCQQPGHNSRDCPEERKPREGRNGFTGGSSGFG 318
Query: 152 KMKGKFPAIARGTEKQVSCYRCGEIGH----FASRCSATGP------------RCFNCHR 195
G G GE G F + G CFNC +
Sbjct: 319 GGNGGGTGFDSGLTNGFGSGNNGESGFGSGGFGGNSNGFGSGGGGQDRGERNNNCFNCQQ 378
Query: 196 VGHLAVVCKKPKVE 209
GH + C +PK E
Sbjct: 379 PGHRSNDCPEPKKE 392
Score = 30.4 bits (67), Expect = 5.9
Identities = 16/51 (31%), Positives = 27/51 (52%), Gaps = 8/51 (15%)
Query: 165 EKQVSCYRCGEIGHFASRC----SATGPR-CFNCHRVGHLAVVC---KKPK 207
E+ +C+ C + GH ++ C PR C+NC + GH + C +KP+
Sbjct: 254 ERNNNCFNCQQPGHRSNDCPEPKKEREPRVCYNCQQPGHNSRDCPEERKPR 304
>YL92_SCHPO (Q9HFF2) Hypothetical protein C683.02c in chromosome I
Length = 218
Score = 48.5 bits (114), Expect = 2e-05
Identities = 27/80 (33%), Positives = 35/80 (43%), Gaps = 19/80 (23%)
Query: 108 CFRCGKNGHYANACLGQGP----RCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARG 163
CFRCG H NAC +GP +CF C++ GHL+ C + KG +P
Sbjct: 102 CFRCGSKEHSLNACSKKGPLKFAKCFICHENGHLSGQC--------EQNPKGLYP----- 148
Query: 164 TEKQVSCYRCGEIGHFASRC 183
K C C + H A C
Sbjct: 149 --KGGCCKFCSSVHHLAKDC 166
Score = 45.8 bits (107), Expect = 1e-04
Identities = 25/82 (30%), Positives = 34/82 (40%), Gaps = 23/82 (28%)
Query: 128 CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATG 187
CF C Q GH+ +C + S C+RCG H + CS G
Sbjct: 79 CFACRQQGHIVQDCPEAKDNVS-------------------ICFRCGSKEHSLNACSKKG 119
Query: 188 P----RCFNCHRVGHLAVVCKK 205
P +CF CH GHL+ C++
Sbjct: 120 PLKFAKCFICHENGHLSGQCEQ 141
Score = 35.4 bits (80), Expect = 0.18
Identities = 27/106 (25%), Positives = 32/106 (29%), Gaps = 28/106 (26%)
Query: 108 CFRCGKNGHYANACLGQGPR---CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGT 164
CF C + GH C CF C H C +G
Sbjct: 79 CFACRQQGHIVQDCPEAKDNVSICFRCGSKEHSLNACS------------------KKGP 120
Query: 165 EKQVSCYRCGEIGHFASRCSAT-------GPRCFNCHRVGHLAVVC 203
K C+ C E GH + +C G C C V HLA C
Sbjct: 121 LKFAKCFICHENGHLSGQCEQNPKGLYPKGGCCKFCSSVHHLAKDC 166
>GLH1_CAEEL (P34689) ATP-dependent RNA helicase glh-1 (EC 3.6.1.-)
(Germline helicase-1)
Length = 763
Score = 47.8 bits (112), Expect = 4e-05
Identities = 34/121 (28%), Positives = 48/121 (39%), Gaps = 22/121 (18%)
Query: 108 CFRCGKNGHYANAC----LGQGPR-CFNCNQIGHLAVNCKTFQ-------------AGPS 149
CF C + GH ++ C + PR C+NC Q GH + C + AG
Sbjct: 160 CFNCQQPGHRSSDCPEPRKEREPRVCYNCQQPGHTSRECTEERKPREGRTGGFGGGAGFG 219
Query: 150 DNKMKGKFPA---IARGTEK-QVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKK 205
+N F G E+ + C+ C GH ++ C CFNC GH + C
Sbjct: 220 NNGGNDGFGGDGGFGGGEERGPMKCFNCKGEGHRSAECPEPPRGCFNCGEQGHRSNECPN 279
Query: 206 P 206
P
Sbjct: 280 P 280
Score = 34.3 bits (77), Expect = 0.41
Identities = 18/53 (33%), Positives = 28/53 (51%), Gaps = 8/53 (15%)
Query: 163 GTEKQVSCYRCGEIGHFASRC----SATGPR-CFNCHRVGHLAVVC---KKPK 207
G E+ +C+ C + GH +S C PR C+NC + GH + C +KP+
Sbjct: 153 GGERNNNCFNCQQPGHRSSDCPEPRKEREPRVCYNCQQPGHTSRECTEERKPR 205
>GAG_FIVT2 (P31821) Gag polyprotein [Contains: Core protein p15;
Major core protein p24; Nucleic acid binding protein
p10]
Length = 449
Score = 47.8 bits (112), Expect = 4e-05
Identities = 24/60 (40%), Positives = 30/60 (50%), Gaps = 1/60 (1%)
Query: 100 QSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPA 159
Q+ P + CF C K GH A C + RC NC + GHLA NC S N+ G+ A
Sbjct: 369 QTKGPRLVCFNCKKPGHLARQCK-EAKRCNNCGKPGHLAANCWQGGRKTSGNEKVGRAAA 427
Score = 38.1 bits (87), Expect = 0.028
Identities = 24/79 (30%), Positives = 31/79 (38%), Gaps = 25/79 (31%)
Query: 124 QGPR--CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFAS 181
+GPR CFNC + GHLA CK + C CG+ GH A+
Sbjct: 371 KGPRLVCFNCKKPGHLARQCKEAKR-----------------------CNNCGKPGHLAA 407
Query: 182 RCSATGPRCFNCHRVGHLA 200
C G + +VG A
Sbjct: 408 NCWQGGRKTSGNEKVGRAA 426
Score = 37.4 bits (85), Expect = 0.048
Identities = 20/57 (35%), Positives = 28/57 (49%), Gaps = 1/57 (1%)
Query: 170 CYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVEPSVNTARGKHPAARGKV 226
C+ C + GH A +C RC NC + GHLA C + + S N G+ A +V
Sbjct: 377 CFNCKKPGHLARQCKEA-KRCNNCGKPGHLAANCWQGGRKTSGNEKVGRAAAPVNQV 432
Score = 31.6 bits (70), Expect = 2.6
Identities = 14/23 (60%), Positives = 16/23 (68%), Gaps = 2/23 (8%)
Query: 187 GPR--CFNCHRVGHLAVVCKKPK 207
GPR CFNC + GHLA CK+ K
Sbjct: 372 GPRLVCFNCKKPGHLARQCKEAK 394
>GR2B_ARATH (Q38896) Glycine-rich protein 2b (AtGRP2b)
Length = 201
Score = 46.6 bits (109), Expect = 8e-05
Identities = 24/64 (37%), Positives = 32/64 (49%), Gaps = 2/64 (3%)
Query: 128 CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ--VSCYRCGEIGHFASRCSA 185
CF C + GH+A C G S G++ + G +SCY CGE GHFA C++
Sbjct: 138 CFKCGEPGHMARECSQGGGGYSGGGGGGRYGSGGGGGGGGGGLSCYSCGESGHFARDCTS 197
Query: 186 TGPR 189
G R
Sbjct: 198 GGAR 201
Score = 31.2 bits (69), Expect = 3.4
Identities = 17/59 (28%), Positives = 21/59 (34%), Gaps = 24/59 (40%)
Query: 169 SCYRCGEIGHFASRCS------------------------ATGPRCFNCHRVGHLAVVC 203
SC++CGE GH A CS G C++C GH A C
Sbjct: 137 SCFKCGEPGHMARECSQGGGGYSGGGGGGRYGSGGGGGGGGGGLSCYSCGESGHFARDC 195
>GAG_SIVAI (Q02843) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 513
Score = 46.6 bits (109), Expect = 8e-05
Identities = 22/54 (40%), Positives = 29/54 (52%), Gaps = 5/54 (9%)
Query: 96 GARSQSAHPEVTCFRCGKNGHYANACLGQGPR---CFNCNQIGHLAVNCKTFQA 146
G + + + CF CGK GH C + PR CF C +IGH+A +CK QA
Sbjct: 380 GPQKKGPRGPLKCFNCGKFGHMQREC--KAPRQIKCFKCGKIGHMAKDCKNGQA 431
Score = 43.1 bits (100), Expect = 9e-04
Identities = 16/44 (36%), Positives = 24/44 (54%), Gaps = 1/44 (2%)
Query: 162 RGTEKQVSCYRCGEIGHFASRCSATGP-RCFNCHRVGHLAVVCK 204
+G + C+ CG+ GH C A +CF C ++GH+A CK
Sbjct: 384 KGPRGPLKCFNCGKFGHMQRECKAPRQIKCFKCGKIGHMAKDCK 427
Score = 42.0 bits (97), Expect = 0.002
Identities = 19/61 (31%), Positives = 28/61 (45%), Gaps = 22/61 (36%)
Query: 124 QGP-RCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASR 182
+GP +CFNC + GH+ CK +Q+ C++CG+IGH A
Sbjct: 387 RGPLKCFNCGKFGHMQRECKA---------------------PRQIKCFKCGKIGHMAKD 425
Query: 183 C 183
C
Sbjct: 426 C 426
>GK12_HUMAN (P63130) HERV-K_1q22 provirus ancestral Gag polyprotein
(Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III)
Gag protein) [Contains: Matrix protein; Capsid protein;
Nucleocapsid protein]
Length = 665
Score = 45.4 bits (106), Expect = 2e-04
Identities = 24/59 (40%), Positives = 29/59 (48%), Gaps = 6/59 (10%)
Query: 125 GPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC 183
G +C+NC QIGHL NC P NK A G E C RC + H+AS+C
Sbjct: 542 GGKCYNCGQIGHLKKNC------PVLNKQNITIQATTTGREPPDLCPRCKKGKHWASQC 594
>GK11_HUMAN (P63145) HERV-K_22q11.21 provirus ancestral Gag
polyprotein (Gag polyprotein) (HERV-K101 Gag protein)
[Contains: Matrix protein; Capsid protein; Nucleocapsid
protein]
Length = 665
Score = 45.4 bits (106), Expect = 2e-04
Identities = 24/59 (40%), Positives = 29/59 (48%), Gaps = 6/59 (10%)
Query: 125 GPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC 183
G +C+NC QIGHL NC P NK A G E C RC + H+AS+C
Sbjct: 542 GGKCYNCGQIGHLKKNC------PVLNKQNITIQATTTGREPPDLCPRCKKGKHWASQC 594
>GK10_HUMAN (P87889) HERV-K_5q33.3 provirus ancestral Gag
polyprotein (Gag polyprotein) (HERV-K10 Gag protein)
(HERV-K107 Gag protein) [Contains: Matrix protein;
Capsid protein; Nucleocapsid protein]
Length = 665
Score = 45.4 bits (106), Expect = 2e-04
Identities = 24/59 (40%), Positives = 29/59 (48%), Gaps = 6/59 (10%)
Query: 125 GPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC 183
G +C+NC QIGHL NC P NK A G E C RC + H+AS+C
Sbjct: 542 GGKCYNCGQIGHLKKNC------PVLNKQNITIQATTTGREPPDLCPRCKKGKHWASQC 594
>GAK6_HUMAN (P62685) HERV-K_8p23.1 provirus ancestral Gag
polyprotein (Gag polyprotein) (HERV-K115 Gag protein)
[Contains: Matrix protein; Capsid protein; Nucleocapsid
protein]
Length = 646
Score = 45.4 bits (106), Expect = 2e-04
Identities = 24/59 (40%), Positives = 29/59 (48%), Gaps = 6/59 (10%)
Query: 125 GPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC 183
G +C+NC QIGHL NC P NK A G E C RC + H+AS+C
Sbjct: 542 GGKCYNCGQIGHLKKNC------PVLNKQNITIQATTTGREPPDLCPRCKKGKHWASQC 594
>GAK5_HUMAN (P62684) HERV-K_19p13.11 provirus ancestral Gag
polyprotein (Gag polyprotein) (HERV-K113 Gag protein)
[Contains: Matrix protein; Capsid protein; Nucleocapsid
protein]
Length = 665
Score = 45.4 bits (106), Expect = 2e-04
Identities = 24/59 (40%), Positives = 29/59 (48%), Gaps = 6/59 (10%)
Query: 125 GPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC 183
G +C+NC QIGHL NC P NK A G E C RC + H+AS+C
Sbjct: 542 GGKCYNCGQIGHLKKNC------PVLNKQNITIQATTTGREPPDLCPRCKKGKHWASQC 594
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.134 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,836,526
Number of Sequences: 164201
Number of extensions: 1629216
Number of successful extensions: 4850
Number of sequences better than 10.0: 125
Number of HSP's better than 10.0 without gapping: 67
Number of HSP's successfully gapped in prelim test: 59
Number of HSP's that attempted gapping in prelim test: 4071
Number of HSP's gapped (non-prelim): 437
length of query: 311
length of database: 59,974,054
effective HSP length: 110
effective length of query: 201
effective length of database: 41,911,944
effective search space: 8424300744
effective search space used: 8424300744
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0134.4


