

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0134.14
(277 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
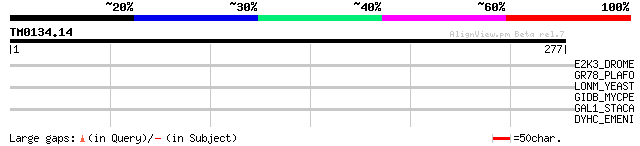
Score E
Sequences producing significant alignments: (bits) Value
E2K3_DROME (Q9NIV1) Eukaryotic translation initiation factor 2-a... 33 0.99
GR78_PLAFO (Q05866) 78 kDa glucose-regulated protein homolog pre... 31 3.8
LONM_YEAST (P36775) Lon protease homolog, mitochondrial precurso... 30 4.9
GIDB_MYCPE (Q8EUW9) Methyltransferase gidB (EC 2.1.-.-) (Glucose... 30 6.4
GAL1_STACA (Q9RGS1) Galactokinase (EC 2.7.1.6) (Galactose kinase) 30 8.4
DYHC_EMENI (P45444) Dynein heavy chain, cytosolic (DYHC) 30 8.4
>E2K3_DROME (Q9NIV1) Eukaryotic translation initiation factor 2-alpha
kinase precursor (EC 2.7.1.-) (PRKR-like endoplasmic
reticulum kinase) (PERK) (PEK) (DmPEK)
Length = 1162
Score = 32.7 bits (73), Expect = 0.99
Identities = 23/63 (36%), Positives = 31/63 (48%), Gaps = 3/63 (4%)
Query: 168 SKLESLAKHFQFFNNHVDERYMCKRFLNGLRADIKDSVRPLGIIRFQALVEKATEVELMK 227
S L S A+H Q H+ YM L G D K + LG+I F+ V +TE+E +K
Sbjct: 1019 SGLPSCARHTQQVGTHL---YMSPEQLLGQHYDYKVDIYSLGLIFFELHVYFSTEMERIK 1075
Query: 228 NRR 230
R
Sbjct: 1076 TLR 1078
>GR78_PLAFO (Q05866) 78 kDa glucose-regulated protein homolog
precursor (GRP 78)
Length = 655
Score = 30.8 bits (68), Expect = 3.8
Identities = 26/110 (23%), Positives = 49/110 (43%), Gaps = 8/110 (7%)
Query: 80 IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKGTRGIMRANHEEVN*NSFRTAF--L 137
IQ++ K E+ + V +T + + D + + RA EE+N + FR +
Sbjct: 283 IQKLRKEVEIAK--RNLSVVHSTQIEIEDIVEGHNFSETLTRAKFEELNDDLFRETLEPV 340
Query: 138 EKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESLAKHFQFFNNHVDER 187
+K + ++ + + L GS +P K++ + K F+FFN R
Sbjct: 341 KKVLDDAKYEKSKIDEIVLVGGSTRIP----KIQQIIKEFEFFNGKEPNR 386
>LONM_YEAST (P36775) Lon protease homolog, mitochondrial precursor
(EC 3.4.21.-)
Length = 1133
Score = 30.4 bits (67), Expect = 4.9
Identities = 21/63 (33%), Positives = 29/63 (45%), Gaps = 2/63 (3%)
Query: 200 DIKDSVRPLGIIRFQALVEKATEVELMKNRRMNGAGTGGPMRSSSQDNLGKGRFQMKKPY 259
D KD+ + R +V K E E KN R + +GG SSS+ + G G + K P
Sbjct: 116 DEKDTDNDVEPTRDDEIVNKDQEGEASKNSR--SSASGGGQSSSSRSDSGDGSSKQKPPK 173
Query: 260 QRP 262
P
Sbjct: 174 DVP 176
>GIDB_MYCPE (Q8EUW9) Methyltransferase gidB (EC 2.1.-.-) (Glucose
inhibited division protein B)
Length = 237
Score = 30.0 bits (66), Expect = 6.4
Identities = 29/102 (28%), Positives = 44/102 (42%), Gaps = 14/102 (13%)
Query: 147 DEREAQFLTLGQGSMTVPEYASKL----------ESLAKHFQFFNNHVDERYMCKRFLNG 196
D + L +G GS VP K+ ES K F NN VD+ + F++
Sbjct: 69 DNKNINLLDIGSGS-GVPGIFLKILFPHINLYIVESNTKKVNFLNNLVDKLELENVFISN 127
Query: 197 LRAD--IKDSVRPLGIIRFQALVEKATEVEL-MKNRRMNGAG 235
R + IKD + +I +A+ E +EL ++NG G
Sbjct: 128 QRCEDYIKDKIEFFDLITCRAVAELRILLELSFPGLKINGIG 169
>GAL1_STACA (Q9RGS1) Galactokinase (EC 2.7.1.6) (Galactose kinase)
Length = 388
Score = 29.6 bits (65), Expect = 8.4
Identities = 18/47 (38%), Positives = 22/47 (46%), Gaps = 1/47 (2%)
Query: 192 RFLNGLRADIKDSVRPLGIIRFQALVEKATEVELMKNRRMNGAGTGG 238
R LN A +K+ GI L E A +VE + RM GAG G
Sbjct: 297 RLLNASHASLKNDYEVTGI-ELDTLAETAQQVEGVLGARMTGAGFAG 342
>DYHC_EMENI (P45444) Dynein heavy chain, cytosolic (DYHC)
Length = 4344
Score = 29.6 bits (65), Expect = 8.4
Identities = 27/106 (25%), Positives = 46/106 (42%), Gaps = 5/106 (4%)
Query: 19 VQAMTVQTNDNAHRRAT--EDTRELHRLQREVALDQNRGLNDFRRQDPPKFTGGTDPDNA 76
+Q M + R+A E L + ++EVAL ++ L+D R +P N
Sbjct: 3235 LQRMVADQREAEQRKAVSLEVQAALEKQEKEVALRKDVVLHDLARAEPAVLEAQKSVSNI 3294
Query: 77 DL*IQEIEKIFEVLQTAEGAKVSL-ATYLLLGDAEYWWKGTRGIMR 121
Q + ++ + G +++L A LLG WK +GI+R
Sbjct: 3295 KR--QHLTEVRSMGNPPAGVRLALEAVCTLLGHKVDSWKTIQGIVR 3338
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.138 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,858,334
Number of Sequences: 164201
Number of extensions: 1273150
Number of successful extensions: 3269
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 3269
Number of HSP's gapped (non-prelim): 8
length of query: 277
length of database: 59,974,054
effective HSP length: 109
effective length of query: 168
effective length of database: 42,076,145
effective search space: 7068792360
effective search space used: 7068792360
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0134.14


