

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0133.3
(700 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
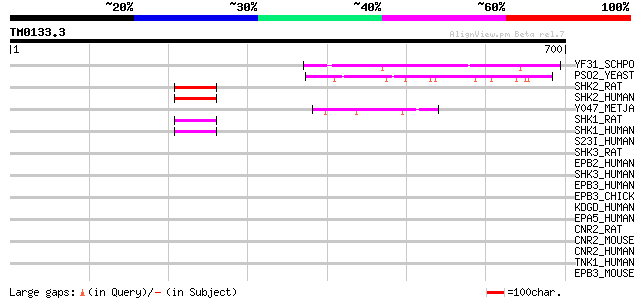
Score E
Sequences producing significant alignments: (bits) Value
YF31_SCHPO (Q10264) Hypothetical protein C22A12.01c in chromosome I 155 4e-37
PSO2_YEAST (P30620) DNA cross-link repair protein PSO2/SNM1 140 1e-32
SHK2_RAT (Q9QX74) SH3 and multiple ankyrin repeat domains protei... 49 5e-05
SHK2_HUMAN (Q9UPX8) SH3 and multiple ankyrin repeat domains prot... 49 5e-05
Y047_METJA (Q60355) Hypothetical protein MJ0047 47 1e-04
SHK1_RAT (Q9WV48) SH3 and multiple ankyrin repeat domains protei... 45 8e-04
SHK1_HUMAN (Q9Y566) SH3 and multiple ankyrin repeat domains prot... 45 8e-04
S23I_HUMAN (Q9Y6Y8) SEC23 interacting protein (p125) (MSTP053) 44 0.001
SHK3_RAT (Q9JLU4) SH3 and multiple ankyrin repeat domains 3 (Sha... 44 0.002
EPB2_HUMAN (P29323) Ephrin type-B receptor 2 precursor (EC 2.7.1... 43 0.003
SHK3_HUMAN (Q9BYB0) SH3 and multiple ankyrin repeat domains prot... 42 0.004
EPB3_HUMAN (P54753) Ephrin type-B receptor 3 precursor (EC 2.7.1... 42 0.004
EPB3_CHICK (Q07498) Ephrin type-B receptor 3 (EC 2.7.1.112) (Tyr... 42 0.004
KDGD_HUMAN (Q16760) Diacylglycerol kinase, delta (EC 2.7.1.107) ... 42 0.006
EPA5_HUMAN (P54756) Ephrin type-A receptor 5 precursor (EC 2.7.1... 42 0.007
CNR2_RAT (Q9Z1T4) Connector enhancer of kinase suppressor of ras... 42 0.007
CNR2_MOUSE (Q80YA9) Connector enhancer of kinase suppressor of r... 42 0.007
CNR2_HUMAN (Q8WXI2) Connector enhancer of kinase suppressor of r... 42 0.007
TNK1_HUMAN (O95271) Tankyrase 1 (EC 2.4.2.30) (TANK1) (Tankyrase... 41 0.009
EPB3_MOUSE (P54754) Ephrin type-B receptor 3 precursor (EC 2.7.1... 41 0.009
>YF31_SCHPO (Q10264) Hypothetical protein C22A12.01c in chromosome I
Length = 560
Score = 155 bits (391), Expect = 4e-37
Identities = 113/351 (32%), Positives = 172/351 (48%), Gaps = 35/351 (9%)
Query: 371 PFRVDAFKYLRGD-CSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNI 429
PF VDAF Y D +FL+HFH D L T + G + E L+ +
Sbjct: 194 PFAVDAFAYGAIDGVEAYFLSHFHSDHYGGL------TPKWKHGPIYCSEVTGNLLINVM 247
Query: 430 GIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILF---QPPNGKAVLHTGDFRFSD 486
+ + L LNQ I GI V LDANHCPGS + +F Q + VLH GDFR S
Sbjct: 248 HVDEQYVKRLKLNQPYNIMGITVYVLDANHCPGSAMFVFETLQSNQTRRVLHCGDFRASK 307
Query: 487 ELAINPVLRMCPVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQ-AEAFNPGTLFLIGS 544
+ ++PVLR +H + LDTTY NP+Y FP Q V+Q D AI ++ + L ++ +
Sbjct: 308 DHVMHPVLREKTIHKVYLDTTYLNPKYTFPPQADVVQACADKAISIKKSTDSRLLVVVST 367
Query: 545 YTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTL 604
Y+IGKE++ + +A+SL ++YV K+ ++K LE ++ + T + ++++H+ M +
Sbjct: 368 YSIGKEKVAVAIAKSLSSRIYVVPRKMHIIKQLE-NQDLIDLLTDDPTQASVHMVTMMGI 426
Query: 605 ASLKRLKHVSSQYASRFSLIVAFSPTGWTFGKGKKKSP---------------------G 643
L ++ QY S F I+ + TGWTF + ++
Sbjct: 427 HPNSLLDYL-EQYNSSFDKIIGYKVTGWTFQPLENRAQLSSSLDSIISRPPKFVEYDLRA 485
Query: 644 RRWQNGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESANAM 694
R + + PYSEH SF +L F ++ +IIP+VN S M
Sbjct: 486 IRGSTDKVAAFVAPYSEHSSFYDLTMFCLSMNIGHIIPTVNVGSQRSREKM 536
>PSO2_YEAST (P30620) DNA cross-link repair protein PSO2/SNM1
Length = 661
Score = 140 bits (352), Expect = 1e-32
Identities = 124/408 (30%), Positives = 183/408 (44%), Gaps = 100/408 (24%)
Query: 374 VDAFKYLRGDC-SHWFLTHFHLDRTRLLDLNIFDTD---LRQMGYLIKFECFARLVNMNI 429
VD F Y + S +FL+HFH D L + + D +++ Y K A LVN+
Sbjct: 230 VDGFNYKASETISQYFLSHFHSDHYIGLKKSWNNPDENPIKKTLYCSKIT--AILVNLKF 287
Query: 430 GIPYDKLHILPLNQKVEIAG-IDVTCLDANHCPGSIIILFQP--PNG-----KAVLHTGD 481
IP D++ ILP+N++ I I V LDANHCPG+II+LFQ N + +LHTGD
Sbjct: 288 KIPMDEIQILPMNKRFWITDTISVVTLDANHCPGAIIMLFQEFLANSYDKPIRQILHTGD 347
Query: 482 FRFSDELAINPVLRMC------PVHTLILDTTYCNPQYDFPKQEAVIQFVIDAI------ 529
FR S+ I + + + + LDTTY Y+FP Q +V + V D
Sbjct: 348 FR-SNAKMIETIQKWLAETANETIDQVYLDTTYMTMGYNFPSQHSVCETVADFTLRLIKH 406
Query: 530 ---------QAEAFN-----------PGTLFLIGSYTIGKERLFLEVARSLRRKVYV--N 567
Q F+ LFL+G+YTIGKE+L +++ L+ K++V N
Sbjct: 407 GKNKTFGDSQRNLFHFQRKKTLTTHRYRVLFLVGTYTIGKEKLAIKICEFLKTKLFVMPN 466
Query: 568 AAKLRLLKCL--EFEEEDMQW----FTLNEHESNIHVAPMWTLAS-------LKRLKHVS 614
+ K ++ + E ++ W T N HES++H+ P+ L S LK LK +
Sbjct: 467 SVKFSMMLTVLQNNENQNDMWDESLLTSNLHESSVHLVPIRVLKSQETIEAYLKSLKELE 526
Query: 615 SQYASRFSLIVAFSPTGWTFGKG----KKKSPGRRWQNG--------------------- 649
+ Y +V F PTGW+ G KK +G
Sbjct: 527 TDYVKDIEDVVGFIPTGWSHNFGLKYQKKNDDDENEMSGNTEYCLELMKNDRDNDDENGF 586
Query: 650 ---TIIR----------YEVPYSEHCSFTELKDFVKHISPANIIPSVN 684
+I+R + VPYSEH SF +L F + + +IP+VN
Sbjct: 587 EISSILRQYKKYNKFQVFNVPYSEHSSFNDLVKFGCKLKCSEVIPTVN 634
>SHK2_RAT (Q9QX74) SH3 and multiple ankyrin repeat domains protein 2
(Shank2) (Proline-rich synapse associated protein 1)
(ProSAP1) (Cortactin-binding protein 1) (CortBP1)
(GKAP/SAPAP interacting protein) (SPANK-3)
Length = 1474
Score = 48.9 bits (115), Expect = 5e-05
Identities = 23/53 (43%), Positives = 33/53 (61%)
Query: 208 VVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL 260
V DWL L LG++++ F+ E+D L L +EDL+ +GVT +G R I AL
Sbjct: 1416 VADWLESLNLGEHKETFMDNEIDGSHLPNLQKEDLIDLGVTRVGHRMNIERAL 1468
>SHK2_HUMAN (Q9UPX8) SH3 and multiple ankyrin repeat domains protein 2
(Shank2)
Length = 1253
Score = 48.9 bits (115), Expect = 5e-05
Identities = 23/53 (43%), Positives = 33/53 (61%)
Query: 208 VVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL 260
V DWL L LG++++ F+ E+D L L +EDL+ +GVT +G R I AL
Sbjct: 1195 VADWLESLNLGEHKEAFMDNEIDGSHLPNLQKEDLIDLGVTRVGHRMNIERAL 1247
>Y047_METJA (Q60355) Hypothetical protein MJ0047
Length = 428
Score = 47.4 bits (111), Expect = 1e-04
Identities = 45/180 (25%), Positives = 72/180 (40%), Gaps = 23/180 (12%)
Query: 383 DCSHWFLTHFHLDRT--------RLLDLNIFDTDLRQMGYLIKFECFARLVNM-NIGIPY 433
D F++H HLD + R +D+ + T+L + + + ++ N IPY
Sbjct: 49 DVDKVFISHAHLDHSGALPVLFHRKMDVPVITTELSKKLIKVLLKDMVKIAETENKKIPY 108
Query: 434 DK-------LHILPLN--QKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRF 484
+ H +PLN K A H PGS IL N K +L+TGD +
Sbjct: 109 NNHDVKEAIRHTIPLNYNDKKYYKDFSYELFSAGHIPGSASILLNYQNNKTILYTGDVKL 168
Query: 485 SDELAINPV---LRMCPVHTLILDTTYCNPQYDFPKQEAVIQFVIDAIQAEAFNPGTLFL 541
D + LI+++TY N + P ++AV I+ I+ F G +
Sbjct: 169 RDTRLTKGADLSYTKDDIDILIIESTYGNSIH--PDRKAVELSFIEKIKEILFRGGVALI 226
>SHK1_RAT (Q9WV48) SH3 and multiple ankyrin repeat domains protein 1
(Shank1) (GKAP/SAPAP interacting protein) (SPANK-1)
(Synamon) (Somatostatin receptor interacting protein)
(SSTR interacting protein) (SSTRIP)
Length = 2167
Score = 44.7 bits (104), Expect = 8e-04
Identities = 23/53 (43%), Positives = 31/53 (58%)
Query: 208 VVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL 260
V DWL LGL ++ F+ E+D L LT+ED + +GVT +G R I AL
Sbjct: 2109 VADWLEWLGLSEHRAQFLDHEIDGSHLPALTKEDYVDLGVTRVGHRMNIDRAL 2161
>SHK1_HUMAN (Q9Y566) SH3 and multiple ankyrin repeat domains protein 1
(Shank1) (Somatostatin receptor interacting protein)
(SSTR interacting protein) (SSTRIP)
Length = 2161
Score = 44.7 bits (104), Expect = 8e-04
Identities = 23/53 (43%), Positives = 31/53 (58%)
Query: 208 VVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL 260
V DWL LGL ++ F+ E+D L LT+ED + +GVT +G R I AL
Sbjct: 2103 VADWLEWLGLAEHRAQFLDHEIDGSHLPALTKEDYVDLGVTRVGHRMNIDRAL 2155
>S23I_HUMAN (Q9Y6Y8) SEC23 interacting protein (p125) (MSTP053)
Length = 1000
Score = 43.9 bits (102), Expect = 0.001
Identities = 29/107 (27%), Positives = 57/107 (53%), Gaps = 12/107 (11%)
Query: 204 KVSRVVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL--- 260
+V + + L L L +Y F +E++D ++L T +DL MG+ LGPRKKI + +
Sbjct: 645 EVLTLQETLEALSLSEYFSTFEKEKIDMESLLMCTVDDLKEMGI-PLGPRKKIANFVEHK 703
Query: 261 -------CEFRKGIAASNEKPEDALAEPRRNRNQNVKMQPHRPERKV 300
+K +AA++ K ++ A+ ++ ++ + + P+RK+
Sbjct: 704 AAKLKKAASEKKAVAATSTKGQEQSAQKTKDM-ASLPSESNEPKRKL 749
>SHK3_RAT (Q9JLU4) SH3 and multiple ankyrin repeat domains 3 (Shank3)
(Proline-rich synapse associated protein 2) (ProSAP2)
(SPANK-2)
Length = 1815
Score = 43.5 bits (101), Expect = 0.002
Identities = 22/53 (41%), Positives = 31/53 (57%)
Query: 208 VVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL 260
V DWL + LG++ D F E++ L LT+ED + +GVT +G R I AL
Sbjct: 1757 VGDWLESIHLGEHRDRFEDHEIEGAHLPALTKEDFVELGVTRVGHRMNIERAL 1809
>EPB2_HUMAN (P29323) Ephrin type-B receptor 2 precursor (EC 2.7.1.112)
(Tyrosine-protein kinase receptor EPH-3) (DRT) (Receptor
protein-tyrosine kinase HEK5) (ERK)
Length = 1055
Score = 42.7 bits (99), Expect = 0.003
Identities = 29/109 (26%), Positives = 49/109 (44%), Gaps = 5/109 (4%)
Query: 208 VVDWLRDLGLGKYEDVFVREE-VDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCEFRKG 266
V +WL + +G+Y++ F +D + + ED+L +GVT G +KKI++++ R
Sbjct: 918 VDEWLEAIKMGQYKESFANAGFTSFDVVSQMMMEDILRLGVTLAGHQKKILNSIQVMRAQ 977
Query: 267 IAASNEKPEDALAEPRRNRNQNVKMQPHRPERKV----DGTSKPAANKK 311
+ LA R + + QP +K DG K KK
Sbjct: 978 MNQIQSVEGQPLARRPRATGRTKRCQPRDVTKKTCNSNDGKKKGMGKKK 1026
>SHK3_HUMAN (Q9BYB0) SH3 and multiple ankyrin repeat domains protein
3 (Shank3) (Proline-rich synapse-associated protein 2)
(ProSAP2) (Fragment)
Length = 797
Score = 42.4 bits (98), Expect = 0.004
Identities = 21/53 (39%), Positives = 31/53 (57%)
Query: 208 VVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL 260
V DWL + LG++ D F E++ L LT++D + +GVT +G R I AL
Sbjct: 739 VGDWLESIHLGEHRDRFEDHEIEGAHLPALTKDDFVELGVTRVGHRMNIERAL 791
>EPB3_HUMAN (P54753) Ephrin type-B receptor 3 precursor (EC
2.7.1.112) (Tyrosine-protein kinase receptor HEK-2)
Length = 998
Score = 42.4 bits (98), Expect = 0.004
Identities = 22/58 (37%), Positives = 36/58 (61%), Gaps = 1/58 (1%)
Query: 208 VVDWLRDLGLGKYEDVFVREE-VDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCEFR 264
V DWL + +G+Y++ FV +D + +T EDLL +GVT G +KKI+ ++ + R
Sbjct: 930 VGDWLDAIKMGRYKESFVSAGFASFDLVAQMTAEDLLRIGVTLAGHQKKILSSIQDMR 987
>EPB3_CHICK (Q07498) Ephrin type-B receptor 3 (EC 2.7.1.112)
(Tyrosine-protein kinase receptor CEK10) (Fragment)
Length = 988
Score = 42.4 bits (98), Expect = 0.004
Identities = 22/58 (37%), Positives = 36/58 (61%), Gaps = 1/58 (1%)
Query: 208 VVDWLRDLGLGKYEDVFVREE-VDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCEFR 264
V DWL + +G+Y++ FV +D + +T EDLL +GVT G +KKI+ ++ + R
Sbjct: 920 VGDWLDAIKMGRYKENFVNAGFASFDLVAQMTAEDLLRIGVTLAGHQKKILSSIQDMR 977
>KDGD_HUMAN (Q16760) Diacylglycerol kinase, delta (EC 2.7.1.107)
(Diglyceride kinase) (DGK-delta) (DAG kinase delta) (130
kDa diacylglycerol kinase) (Fragment)
Length = 1195
Score = 42.0 bits (97), Expect = 0.006
Identities = 21/62 (33%), Positives = 32/62 (50%)
Query: 208 VVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCEFRKGI 267
V WL L L +Y+D+F R ++ L L DL +GVT +G K+I+ + E +
Sbjct: 1131 VAAWLEHLSLCEYKDIFTRHDIRGSELLHLERRDLKDLGVTKVGHMKRILCGIKELSRSA 1190
Query: 268 AA 269
A
Sbjct: 1191 PA 1192
>EPA5_HUMAN (P54756) Ephrin type-A receptor 5 precursor (EC 2.7.1.112)
(Tyrosine-protein kinase receptor EHK-1) (Eph homology
kinase-1) (Receptor protein-tyrosine kinase HEK7)
Length = 1037
Score = 41.6 bits (96), Expect = 0.007
Identities = 25/81 (30%), Positives = 46/81 (55%), Gaps = 5/81 (6%)
Query: 189 DVAVRDDNVDAQFAPKVS----RVVDWLRDLGLGKYEDVFVREEVD-WDTLQWLTEEDLL 243
+ + R N+ A+ +P S V +WL + +G+Y ++F+ D + +T EDL
Sbjct: 947 NASCRVSNLLAEHSPLGSGAYRSVGEWLEAIKMGRYTEIFMENGYSSMDAVAQVTLEDLR 1006
Query: 244 SMGVTALGPRKKIVHALCEFR 264
+GVT +G +KKI+++L E +
Sbjct: 1007 RLGVTLVGHQKKIMNSLQEMK 1027
>CNR2_RAT (Q9Z1T4) Connector enhancer of kinase suppressor of ras 2
(Connector enhancer of KSR2) (CNK2) (Membrane-associated
guanylate kinase-interacting protein) (MAGUIN)
Length = 1032
Score = 41.6 bits (96), Expect = 0.007
Identities = 25/73 (34%), Positives = 41/73 (55%), Gaps = 5/73 (6%)
Query: 206 SRVVDWLRDLG--LGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHA---L 260
S+VVDW++ L L +Y F RE++ D L +T ++L +GV+ +G ++ I+ A L
Sbjct: 14 SQVVDWMKGLDDCLQQYIKNFEREKISGDQLLRITHQELEDLGVSRIGHQELILEAVDLL 73
Query: 261 CEFRKGIAASNEK 273
C G+ N K
Sbjct: 74 CALNYGLETENLK 86
>CNR2_MOUSE (Q80YA9) Connector enhancer of kinase suppressor of ras
2 (Connector enhancer of KSR2) (CNK2)
Length = 1032
Score = 41.6 bits (96), Expect = 0.007
Identities = 25/73 (34%), Positives = 41/73 (55%), Gaps = 5/73 (6%)
Query: 206 SRVVDWLRDLG--LGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHA---L 260
S+VVDW++ L L +Y F RE++ D L +T ++L +GV+ +G ++ I+ A L
Sbjct: 14 SQVVDWMKGLDDCLQQYIKNFEREKISGDQLLRITHQELEDLGVSRIGHQELILEAVDLL 73
Query: 261 CEFRKGIAASNEK 273
C G+ N K
Sbjct: 74 CALNYGLETENLK 86
>CNR2_HUMAN (Q8WXI2) Connector enhancer of kinase suppressor of ras
2 (Connector enhancer of KSR2) (CNK2)
Length = 1034
Score = 41.6 bits (96), Expect = 0.007
Identities = 25/73 (34%), Positives = 41/73 (55%), Gaps = 5/73 (6%)
Query: 206 SRVVDWLRDLG--LGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHA---L 260
S+VVDW++ L L +Y F RE++ D L +T ++L +GV+ +G ++ I+ A L
Sbjct: 14 SQVVDWMKGLDDCLQQYIKNFEREKISGDQLLRITHQELEDLGVSRIGHQELILEAVDLL 73
Query: 261 CEFRKGIAASNEK 273
C G+ N K
Sbjct: 74 CALNYGLETENLK 86
>TNK1_HUMAN (O95271) Tankyrase 1 (EC 2.4.2.30) (TANK1) (Tankyrase I)
(TNKS-1) (TRF1-interacting ankyrin-related ADP-ribose
polymerase)
Length = 1327
Score = 41.2 bits (95), Expect = 0.009
Identities = 28/111 (25%), Positives = 46/111 (41%), Gaps = 1/111 (0%)
Query: 161 CPLCEVDISNLTEEQRHLHTNDCLDRGDDVAVRDDNVDAQFAPKVSRVVDWLRDLGLGKY 220
C I NLT L + GD A + + + A + +L+ LGL
Sbjct: 985 CLSAASSIDNLTGPLAELAVGGASNAGDGAA-GTERKEGEVAGLDMNISQFLKSLGLEHL 1043
Query: 221 EDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCEFRKGIAASN 271
D+F E++ D L + E+L +G+ A G R K++ + G +N
Sbjct: 1044 RDIFETEQITLDVLADMGHEELKEIGINAYGHRHKLIKGVERLLGGQQGTN 1094
>EPB3_MOUSE (P54754) Ephrin type-B receptor 3 precursor (EC
2.7.1.112) (Tyrosine-protein kinase receptor MDK-5)
(Developmental kinase 5) (SEK-4)
Length = 993
Score = 41.2 bits (95), Expect = 0.009
Identities = 22/58 (37%), Positives = 36/58 (61%), Gaps = 1/58 (1%)
Query: 208 VVDWLRDLGLGKYEDVFVREE-VDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCEFR 264
V DWL + +G+Y++ FV +D + +T EDLL +GVT G +KKI+ ++ + R
Sbjct: 925 VGDWLDAIKMGRYKESFVGAGFASFDLVAQMTAEDLLRIGVTLAGHQKKILCSIQDMR 982
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.134 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 84,173,431
Number of Sequences: 164201
Number of extensions: 3739922
Number of successful extensions: 10176
Number of sequences better than 10.0: 100
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 76
Number of HSP's that attempted gapping in prelim test: 10001
Number of HSP's gapped (non-prelim): 170
length of query: 700
length of database: 59,974,054
effective HSP length: 117
effective length of query: 583
effective length of database: 40,762,537
effective search space: 23764559071
effective search space used: 23764559071
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0133.3


