

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0132.6
(162 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
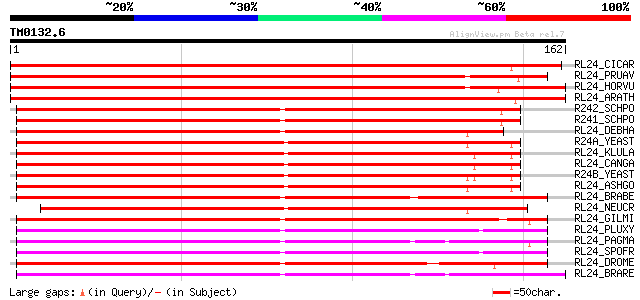
Score E
Sequences producing significant alignments: (bits) Value
RL24_CICAR (O65743) 60S ribosomal protein L24 292 2e-79
RL24_PRUAV (Q9FUL4) 60S ribosomal protein L24 279 2e-75
RL24_HORVU (P50888) 60S ribosomal protein L24 274 7e-74
RL24_ARATH (P38666) 60S ribosomal protein L24 268 5e-72
R242_SCHPO (O74884) 60S ribosomal protein L24-B 135 4e-32
R241_SCHPO (Q92354) 60S ribosomal protein L24-A 134 9e-32
RL24_DEBHA (Q6BNC2) 60S ribosomal protein L24 133 2e-31
R24A_YEAST (P04449) 60S ribosomal protein L24-A (L30A) (RP29) (Y... 131 8e-31
RL24_KLULA (P38665) 60S ribosomal protein L24 (Ribosomal protein... 130 1e-30
RL24_CANGA (Q6FXY9) 60S ribosomal protein L24 130 1e-30
R24B_YEAST (P24000) 60S ribosomal protein L24-B (L30B) (RP29) (Y... 130 1e-30
RL24_ASHGO (Q752U6) 60S ribosomal protein L24 129 2e-30
RL24_BRABE (Q8ISQ3) 60S ribosomal protein L24 125 3e-29
RL24_NEUCR (Q7SDU2) 60S ribosomal protein L24 124 9e-29
RL24_GILMI (Q9DFQ7) 60S ribosomal protein L24 124 1e-28
RL24_PLUXY (Q6F444) 60S ribosomal protein L24 123 2e-28
RL24_PAGMA (Q6Y263) 60S ribosomal protein L24 123 2e-28
RL24_SPOFR (Q962T5) 60S ribosomal protein L24 122 3e-28
RL24_DROME (Q9VJY6) 60S ribosomal protein L24 122 3e-28
RL24_BRARE (Q8JGR4) 60S ribosomal protein L24 122 3e-28
>RL24_CICAR (O65743) 60S ribosomal protein L24
Length = 164
Score = 292 bits (748), Expect = 2e-79
Identities = 146/163 (89%), Positives = 154/163 (93%), Gaps = 2/163 (1%)
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSGAKIYPGRGIRFIR DSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ
Sbjct: 1 MVLKTELCRFSGAKIYPGRGIRFIRGDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKD AQEA +K+RR+TKKPYSRSIVGATLEVIQK+RTEKPEVRDAAREAALREIKER+K
Sbjct: 61 HKKDAAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRTEKPEVRDAAREAALREIKERIK 120
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKGG--KVAAPKGPKLGGGGGK 161
KTKDEK+AKKAEV +KAQKSQGKG K A PKGPK+GGGGGK
Sbjct: 121 KTKDEKKAKKAEVASKAQKSQGKGNVQKGALPKGPKMGGGGGK 163
>RL24_PRUAV (Q9FUL4) 60S ribosomal protein L24
Length = 186
Score = 279 bits (713), Expect = 2e-75
Identities = 142/160 (88%), Positives = 150/160 (93%), Gaps = 4/160 (2%)
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSGAKIYPG+GIRFIRSDSQVFLF NSKCKRYFHNRLKPSKLTWTAM+RKQ
Sbjct: 1 MVLKTELCRFSGAKIYPGKGIRFIRSDSQVFLFANSKCKRYFHNRLKPSKLTWTAMYRKQ 60
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKDIAQEA +K+RR+TKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKER+K
Sbjct: 61 HKKDIAQEAVKKRRRTTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERIK 120
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKGGKV---AAPKGPKLGG 157
KTKDEK+AKKAEV TK+QKSQGKG A PKGPKLGG
Sbjct: 121 KTKDEKKAKKAEV-TKSQKSQGKGSIAKGGAQPKGPKLGG 159
>RL24_HORVU (P50888) 60S ribosomal protein L24
Length = 162
Score = 274 bits (700), Expect = 7e-74
Identities = 139/163 (85%), Positives = 149/163 (91%), Gaps = 2/163 (1%)
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSG KIYPG+GIRFIRSDSQVFLF NSKCKRYFHNRLKP+KL WTAM+RKQ
Sbjct: 1 MVLKTELCRFSGQKIYPGKGIRFIRSDSQVFLFANSKCKRYFHNRLKPAKLCWTAMYRKQ 60
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKDI EAA+K+RR+TKKPYSRSIVGATLEVIQK+R EKPEVRDAAREAALREIKER+K
Sbjct: 61 HKKDIHAEAAKKRRRTTKKPYSRSIVGATLEVIQKKRAEKPEVRDAAREAALREIKERIK 120
Query: 121 KTKDEKRAKKAEVQTKAQKSQ-GKGGKVAAPKGPKLGGGGGKR 162
KTKDEK+AKKAEV TK+QKSQ GKG KGPKLGGGGGKR
Sbjct: 121 KTKDEKKAKKAEV-TKSQKSQGGKGAVQKGSKGPKLGGGGGKR 162
>RL24_ARATH (P38666) 60S ribosomal protein L24
Length = 163
Score = 268 bits (684), Expect = 5e-72
Identities = 133/163 (81%), Positives = 146/163 (88%), Gaps = 1/163 (0%)
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSG KIYPGRGIRFIRSDSQVFLF+NSKCKRYFHN+LKPSKL WTAM+RKQ
Sbjct: 1 MVLKTELCRFSGQKIYPGRGIRFIRSDSQVFLFLNSKCKRYFHNKLKPSKLAWTAMYRKQ 60
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKD AQEA +++RR+TKKPYSRSIVGATLEVIQK+R EKPEVRDAAREAALREIKER+K
Sbjct: 61 HKKDAAQEAVKRRRRATKKPYSRSIVGATLEVIQKKRAEKPEVRDAAREAALREIKERIK 120
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKGGK-VAAPKGPKLGGGGGKR 162
KTKDEK+AKK E +K QK + K AA KGPK+GGGGGKR
Sbjct: 121 KTKDEKKAKKVEFASKQQKVKANFPKAAAASKGPKVGGGGGKR 163
>R242_SCHPO (O74884) 60S ribosomal protein L24-B
Length = 149
Score = 135 bits (340), Expect = 4e-32
Identities = 68/148 (45%), Positives = 102/148 (67%), Gaps = 2/148 (1%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K E+C FSG+K+YPG G F+R D++VF FVN K + F R P +L+WT ++R+ HK
Sbjct: 1 MKVEVCSFSGSKVYPGAGRLFVRGDNKVFRFVNKKSESLFLQRKNPRRLSWTVLYRRMHK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K I++E A+K+ R T K + R IVGA L+VI+++R ++PEVR AAR AAL++ K++ +
Sbjct: 61 KGISEEHAKKRTRRTVK-HQRGIVGANLDVIKEKRNQRPEVRAAARAAALKQRKDKRAAS 119
Query: 123 KDEKRAKKAEVQTKAQKSQG-KGGKVAA 149
+ EK+A KA+ + + Q K KVAA
Sbjct: 120 ESEKKAIKAKSAASSARGQAIKNAKVAA 147
>R241_SCHPO (Q92354) 60S ribosomal protein L24-A
Length = 149
Score = 134 bits (337), Expect = 9e-32
Identities = 67/148 (45%), Positives = 101/148 (67%), Gaps = 2/148 (1%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K E+C FSG+K+YPG G F+R D++VF FVN K + F R P +L+WT ++R+ HK
Sbjct: 1 MKVEVCSFSGSKVYPGAGRLFVRGDNKVFRFVNKKSESLFLQRKNPRRLSWTVLYRRMHK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K I++E A+K+ R T K + R IVGA L+VI+++R ++PEVR AAR AAL++ K++ +
Sbjct: 61 KGISEEHAKKRTRRTVK-HQRGIVGANLDVIKEKRNQRPEVRAAARAAALKQRKDKKAAS 119
Query: 123 KDEKRAKKAEVQTKAQKSQG-KGGKVAA 149
+ EK+A KA+ + + Q K K AA
Sbjct: 120 ESEKKAIKAKSAASSARGQAIKNAKAAA 147
>RL24_DEBHA (Q6BNC2) 60S ribosomal protein L24
Length = 156
Score = 133 bits (335), Expect = 2e-31
Identities = 68/147 (46%), Positives = 99/147 (67%), Gaps = 6/147 (4%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K EL FSG KIYPGRG F+R DS++F F +SK FH R P +++WT ++R+QHK
Sbjct: 1 MKVELDSFSGTKIYPGRGTLFVRGDSKIFRFQSSKSASLFHQRKNPRRISWTVLYRRQHK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K I++E+A+++ R T K R+IVGA+LE+I++RR++KP R AAR+ L + KE K
Sbjct: 61 KGISEESAKRRTRKTVK-NQRAIVGASLELIKERRSQKPSDRKAARDVKLAKDKEAKKAD 119
Query: 123 KDEKRAKKAE-----VQTKAQKSQGKG 144
K ++A+KA+ Q+K K Q KG
Sbjct: 120 KAARKAEKAKSAAAGAQSKVSKQQSKG 146
>R24A_YEAST (P04449) 60S ribosomal protein L24-A (L30A) (RP29)
(YL21)
Length = 155
Score = 131 bits (329), Expect = 8e-31
Identities = 73/153 (47%), Positives = 97/153 (62%), Gaps = 7/153 (4%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K E+ FSGAKIYPGRG F+R DS++F F NSK F R P ++ WT +FRK HK
Sbjct: 1 MKVEIDSFSGAKIYPGRGTLFVRGDSKIFRFQNSKSASLFKQRKNPRRIAWTVLFRKHHK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K I +E A+K+ R T K R I GA+L++I++RR+ KPEVR A RE L+ KE+ K
Sbjct: 61 KGITEEVAKKRSRKTVKA-QRPITGASLDLIKERRSLKPEVRKANREEKLKANKEKKKAE 119
Query: 123 KDEKRAKKAE----VQTKAQKSQGKGG--KVAA 149
K ++A+KA+ +K K Q KG KVAA
Sbjct: 120 KAARKAEKAKSAGTQSSKFSKQQAKGAFQKVAA 152
>RL24_KLULA (P38665) 60S ribosomal protein L24 (Ribosomal protein
L30)
Length = 155
Score = 130 bits (327), Expect = 1e-30
Identities = 69/153 (45%), Positives = 100/153 (65%), Gaps = 7/153 (4%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K E+ FSGAKIYPGRG F+R DS++F F +SK FH R P ++ WT ++R+ HK
Sbjct: 1 MKVEIDSFSGAKIYPGRGTLFVRGDSKIFRFQSSKSASLFHQRKNPRRIAWTVLYRRHHK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K I +E ARK+ R + K R++VGA+LE+I++RR+ KPEVR A R+ + KE+ K
Sbjct: 61 KGITEEVARKRTRKSVKA-QRAVVGASLELIKERRSLKPEVRKAQRDEKKKADKEKKKAD 119
Query: 123 KDEKRAKKAEVQ----TKAQKSQGKGG--KVAA 149
K ++++KA++ +K K Q KG KVAA
Sbjct: 120 KAARKSEKAKLAAAQGSKVSKQQAKGAFQKVAA 152
>RL24_CANGA (Q6FXY9) 60S ribosomal protein L24
Length = 155
Score = 130 bits (327), Expect = 1e-30
Identities = 71/153 (46%), Positives = 99/153 (64%), Gaps = 7/153 (4%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K E+ FSGAKIYPGRG F+R DS++F F NSK F R P ++ WT ++R+ HK
Sbjct: 1 MKVEIDSFSGAKIYPGRGTLFVRGDSKIFRFQNSKSASLFKQRKNPRRVAWTVLYRRHHK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K I +E A+K+ R T K R+IVGA+LE+I++RR+ KPEVR A R+ + KE+ K
Sbjct: 61 KGITEEVAKKRSRKTVKA-QRAIVGASLELIKERRSLKPEVRKAKRDDKAKADKEKKKAD 119
Query: 123 KDEKRAKKAEVQ----TKAQKSQGKGG--KVAA 149
K ++A+KA++ +K K Q KG KVAA
Sbjct: 120 KAARKAEKAKLAAAQGSKVSKQQAKGAFQKVAA 152
>R24B_YEAST (P24000) 60S ribosomal protein L24-B (L30B) (RP29)
(YL21)
Length = 155
Score = 130 bits (327), Expect = 1e-30
Identities = 74/153 (48%), Positives = 99/153 (64%), Gaps = 7/153 (4%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K E+ FSGAKIYPGRG F+R DS++F F NSK F R P ++ WT +FRK HK
Sbjct: 1 MKVEVDSFSGAKIYPGRGTLFVRGDSKIFRFQNSKSASLFKQRKNPRRIAWTVLFRKHHK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K I +E A+K+ R T K R I GA+L++I++RR+ KPEVR A RE L+ KE+ +
Sbjct: 61 KGITEEVAKKRSRKTVKA-QRPITGASLDLIKERRSLKPEVRKANREEKLKANKEKKRAE 119
Query: 123 KDEKRAKKAE---VQ-TKAQKSQGKGG--KVAA 149
K ++A+KA+ VQ +K K Q KG KVAA
Sbjct: 120 KAARKAEKAKSAGVQGSKVSKQQAKGAFQKVAA 152
>RL24_ASHGO (Q752U6) 60S ribosomal protein L24
Length = 155
Score = 129 bits (325), Expect = 2e-30
Identities = 70/153 (45%), Positives = 98/153 (63%), Gaps = 7/153 (4%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K E+ FSGAKIYPGRG F+R DS+VF F +SK FH R P ++ WT ++R+ HK
Sbjct: 1 MKVEIDSFSGAKIYPGRGTLFVRGDSKVFRFHSSKSASLFHQRKNPRRIAWTVLYRRHHK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K I +E A+K+ R T K R++VGA+LE+I++RR+ KPE+R A R+ + KE+ K
Sbjct: 61 KGITEEVAKKRSRKTVKA-QRAVVGASLELIKERRSLKPEIRKAKRDEKAKADKEKKKAD 119
Query: 123 KDEKRAKKAE----VQTKAQKSQGKGG--KVAA 149
K ++A KA+ +K K Q KG KVAA
Sbjct: 120 KAARKADKAKSAATQASKISKQQAKGAFQKVAA 152
>RL24_BRABE (Q8ISQ3) 60S ribosomal protein L24
Length = 154
Score = 125 bits (315), Expect = 3e-29
Identities = 63/155 (40%), Positives = 96/155 (61%), Gaps = 3/155 (1%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K ELC +SG KIYPG G R+ R+D +V+ F+N+KC+ F + P ++TWT ++R++HK
Sbjct: 1 MKIELCLYSGYKIYPGHGKRYARTDGRVYTFLNAKCEASFLMKRNPREITWTVLYRRKHK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K +E +K+ R T K + R+I GA+L I +R +KPEVR A RE A++ KE KK
Sbjct: 61 KGQQEEIQKKRSRRTAK-FQRAITGASLTEIMAKRNQKPEVRKAQREQAVKAAKE--KKK 117
Query: 123 KDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGG 157
D+ +KA + +A + K K P++GG
Sbjct: 118 ADQVGKRKAPAKARATAPKTKAPKTVKAAAPRVGG 152
>RL24_NEUCR (Q7SDU2) 60S ribosomal protein L24
Length = 156
Score = 124 bits (311), Expect = 9e-29
Identities = 64/144 (44%), Positives = 98/144 (67%), Gaps = 3/144 (2%)
Query: 10 FSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHKKDIAQEA 69
FSG +IYPG+G ++R DS++F F N K + F R P ++ WT ++R+QHKK I++E
Sbjct: 8 FSGQRIYPGKGKLYVRGDSKIFRFQNGKSESLFLQRKNPRRIAWTVLYRRQHKKGISEEV 67
Query: 70 ARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKTKDEKRAK 129
A+K+ R T K R+IVGA+LEVI++RR+ +PE R AAR AA++E K + ++T+ K+A+
Sbjct: 68 AKKRSRRTVKA-QRAIVGASLEVIKERRSMRPEARSAARLAAIKESKAKKQETQAAKKAE 126
Query: 130 KAE--VQTKAQKSQGKGGKVAAPK 151
KA+ KA+ + +G K A K
Sbjct: 127 KAKNAANPKARVTSKQGAKGAPVK 150
>RL24_GILMI (Q9DFQ7) 60S ribosomal protein L24
Length = 157
Score = 124 bits (310), Expect = 1e-28
Identities = 65/158 (41%), Positives = 96/158 (60%), Gaps = 6/158 (3%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K ELC FSG KIYPG G R+ R D +VF F+N+KC+ F + P ++ WT ++R++HK
Sbjct: 1 MKVELCSFSGYKIYPGHGRRYARIDGKVFQFLNAKCESAFLAKRNPRQINWTVLYRRKHK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K ++E +K+ R K + R+I GA+L I +R +KPEVR A RE A+R KE K
Sbjct: 61 KGQSEEVTKKRTRRAVK-FQRAITGASLAEIMAKRNQKPEVRKAQREQAIRAAKESKKAK 119
Query: 123 KDEKRAKKAEVQTKAQKSQGKGGKVAAP---KGPKLGG 157
+ K+ A +T A+ +Q K+A P P++GG
Sbjct: 120 QATKKPAAASAKTSAKTAQKP--KIAKPMKISAPRVGG 155
>RL24_PLUXY (Q6F444) 60S ribosomal protein L24
Length = 155
Score = 123 bits (308), Expect = 2e-28
Identities = 65/155 (41%), Positives = 94/155 (59%), Gaps = 2/155 (1%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K LC +SG KIYPG G ++ D + F F+NSKC+ R P K+TWT ++R++ K
Sbjct: 1 MKIGLCAYSGYKIYPGHGKTMVKVDGKNFTFLNSKCEAAHLMRRNPRKVTWTVLYRRKFK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K +E A+K+ R T+K Y R+IVGA+L I +R KPEVR A RE A+++ KE+ K T
Sbjct: 61 KGQEEEQAKKRTRRTQK-YQRAIVGASLSDIMAKRNMKPEVRKAQREQAIKQAKEQKKST 119
Query: 123 KDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGG 157
K K+ + KA + K KV+ P++GG
Sbjct: 120 KASKKTTAPPTKGKA-APKAKAAKVSQKAAPRVGG 153
>RL24_PAGMA (Q6Y263) 60S ribosomal protein L24
Length = 157
Score = 123 bits (308), Expect = 2e-28
Identities = 67/158 (42%), Positives = 95/158 (59%), Gaps = 6/158 (3%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K ELC FSG KIYPG G R+ R D +VF F+N+KC+ F + P ++ WT ++R++HK
Sbjct: 1 MKVELCSFSGYKIYPGHGRRYARIDGKVFQFLNAKCESAFLAKRNPRQINWTVLYRRKHK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K ++E A+K+ R K + R+I GA+L I +R +KPEVR A RE A+R KE KK
Sbjct: 61 KGQSEEVAKKRTRRAVK-FQRAITGASLAEIMAKRNQKPEVRKAQREQAIRAAKE-AKKA 118
Query: 123 KDEKRAKKAEVQTKAQKSQGKGGKVAAP---KGPKLGG 157
K + K A KA + K+A P P++GG
Sbjct: 119 KQAAK-KPAAPSAKASTKTAQKPKIAKPMKISAPRVGG 155
>RL24_SPOFR (Q962T5) 60S ribosomal protein L24
Length = 155
Score = 122 bits (306), Expect = 3e-28
Identities = 64/155 (41%), Positives = 93/155 (59%), Gaps = 2/155 (1%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K LC +SG KIYPG G ++ D + F F+NSKC+ R P K+TWT ++R++ K
Sbjct: 1 MKIGLCAYSGYKIYPGHGKTMVKVDGKTFTFLNSKCEAAHLMRRNPRKVTWTVLYRRKFK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K +E +K+ R T+K Y R+IVGA+L I +R KPEVR A R+ A++ KE+ K T
Sbjct: 61 KGQEEEQTKKRTRRTQK-YQRAIVGASLSDIMAKRNMKPEVRKAQRDQAIKAAKEQKKST 119
Query: 123 KDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGG 157
K K+A + KA + K KV+ P++GG
Sbjct: 120 KAAKKASAPAPKAKA-APKAKAAKVSQKSAPRVGG 153
>RL24_DROME (Q9VJY6) 60S ribosomal protein L24
Length = 155
Score = 122 bits (306), Expect = 3e-28
Identities = 65/157 (41%), Positives = 98/157 (62%), Gaps = 6/157 (3%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K LC FSG KIYPG G ++ D + F F++ KC+R + + P K+TWT ++R++H+
Sbjct: 1 MKIGLCAFSGYKIYPGHGKTMVKIDGKSFTFLDKKCERSYLMKRNPRKVTWTVLYRRKHR 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K I +EA++K+ R T+K + R+IVGA+L I +R KPEVR A R+ A++ KE+ +
Sbjct: 61 KGIEEEASKKRTRRTQK-FQRAIVGASLAEILAKRNMKPEVRKAQRDQAIKVAKEQKRAV 119
Query: 123 KDEKRAKKAEVQTKAQKS--QGKGGKVAAPKGPKLGG 157
K AKKA A+KS + K KV P++GG
Sbjct: 120 ---KAAKKAAAPAPAKKSAPKQKAAKVTQKAAPRVGG 153
>RL24_BRARE (Q8JGR4) 60S ribosomal protein L24
Length = 157
Score = 122 bits (306), Expect = 3e-28
Identities = 66/160 (41%), Positives = 96/160 (59%), Gaps = 3/160 (1%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K ELC FSG KIYPG G R+ R D +VF F+N+KC+ F ++ P ++ WT ++R++HK
Sbjct: 1 MKVELCSFSGYKIYPGHGRRYARVDGKVFQFLNAKCESAFLSKRNPRQINWTVLYRRKHK 60
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K ++E ++K+ R K + R+I GA+L I +R +KPEVR A RE A+R KE KK
Sbjct: 61 KGQSEEVSKKRTRRAVK-FQRAITGASLAEILAKRNQKPEVRKAQREQAIRAAKE-AKKA 118
Query: 123 KDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGGGGGKR 162
K + K+ +KA + K+A P GGKR
Sbjct: 119 KQATK-KQTTQSSKAPAKSAQKQKIAKPMKVSAPRVGGKR 157
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.132 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,657,743
Number of Sequences: 164201
Number of extensions: 646495
Number of successful extensions: 5585
Number of sequences better than 10.0: 468
Number of HSP's better than 10.0 without gapping: 191
Number of HSP's successfully gapped in prelim test: 282
Number of HSP's that attempted gapping in prelim test: 4711
Number of HSP's gapped (non-prelim): 960
length of query: 162
length of database: 59,974,054
effective HSP length: 102
effective length of query: 60
effective length of database: 43,225,552
effective search space: 2593533120
effective search space used: 2593533120
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0132.6


