

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0126.7
(542 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
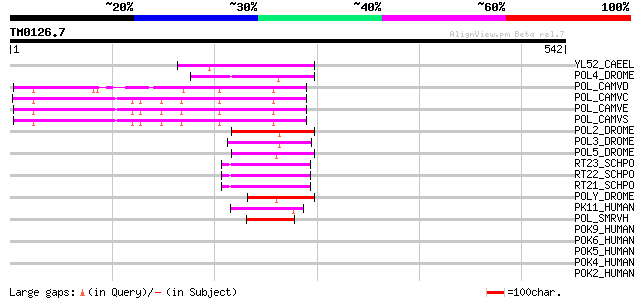
Score E
Sequences producing significant alignments: (bits) Value
YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III 70 1e-11
POL4_DROME (P10394) Retrovirus-related Pol polyprotein from tran... 69 3e-11
POL_CAMVD (P03556) Enzymatic polyprotein [Contains: Aspartic pro... 60 1e-08
POL_CAMVC (P03555) Enzymatic polyprotein [Contains: Aspartic pro... 59 2e-08
POL_CAMVE (Q02964) Enzymatic polyprotein [Contains: Aspartic pro... 59 3e-08
POL_CAMVS (P03554) Enzymatic polyprotein [Contains: Aspartic pro... 57 9e-08
POL2_DROME (P20825) Retrovirus-related Pol polyprotein from tran... 57 1e-07
POL3_DROME (P04323) Retrovirus-related Pol polyprotein from tran... 56 2e-07
POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from tran... 55 5e-07
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 48 6e-05
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 48 6e-05
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 48 6e-05
POLY_DROME (P10401) Retrovirus-related Pol polyprotein from tran... 45 5e-04
PK11_HUMAN (P87893) HERV-K_22q11.21 provirus ancestral Pol prote... 45 6e-04
POL_SMRVH (P03364) Pol polyprotein [Contains: Reverse transcript... 44 8e-04
POK9_HUMAN (P63134) HERV-K_5q13.3 provirus ancestral Pol protein... 44 0.001
POK6_HUMAN (P63133) HERV-K_8p23.1 provirus ancestral Pol protein... 44 0.001
POK5_HUMAN (P63132) HERV-K_19p13.11 provirus ancestral Pol prote... 44 0.001
POK4_HUMAN (P63128) HERV-K_6q14.1 provirus ancestral Pol protein... 44 0.001
POK2_HUMAN (Q9BXR3) HERV-K_7p22.1 provirus ancestral Pol protein... 44 0.001
>YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III
Length = 2186
Score = 70.5 bits (171), Expect = 1e-11
Identities = 49/144 (34%), Positives = 71/144 (49%), Gaps = 11/144 (7%)
Query: 165 AEVQEELVAVPEQLQGLMGDYSELFSTIW-----------DLPPSRVQDHAIHLQEGAAI 213
AE E + E L+ GD +++ I +L + + I L+EGA
Sbjct: 883 AETYERFTTICEHLKRENGDDRKIWDVIEQFQDVFAISDDELGRNSGTECVIELKEGAEP 942
Query: 214 PNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFH 273
+P P K EI K++++ML+ + R S S +SS V+LVKKKDGS R CIDYR +
Sbjct: 943 IRQKPRPIPLALKPEIRKMIQKMLNQKVIRESKSPWSSPVVLVKKKDGSIRMCIDYRKVN 1002
Query: 274 KITIPNKFPILIIDELLYKLGEAK 297
K+ N P+ I+ L L K
Sbjct: 1003 KVVKNNAHPLPNIEATLQSLAGKK 1026
>POL4_DROME (P10394) Retrovirus-related Pol polyprotein from
transposon 412 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1237
Score = 68.9 bits (167), Expect = 3e-11
Identities = 40/128 (31%), Positives = 72/128 (56%), Gaps = 8/128 (6%)
Query: 177 QLQGLMGDYSELFSTIWD-LPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKE 235
QL+ + +Y ++F+ + + + + + L++ + + YR PH Q EI+ V++
Sbjct: 278 QLENICSEYIDIFALESEPITVNNLYKQQLRLKDDEPVYT-KNYRSPHSQVEEIQAQVQK 336
Query: 236 MLDVGIFRPSISQFSSLVILVKKKDG------SWRFCIDYRAFHKITIPNKFPILIIDEL 289
++ I PS+SQ++S ++LV KK WR IDYR +K + +KFP+ ID++
Sbjct: 337 LIKDKIVEPSVSQYNSPLLLVPKKSSPNSDKKKWRLVIDYRQINKKLLADKFPLPRIDDI 396
Query: 290 LYKLGEAK 297
L +LG AK
Sbjct: 397 LDQLGRAK 404
>POL_CAMVD (P03556) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 674
Score = 60.1 bits (144), Expect = 1e-08
Identities = 71/328 (21%), Positives = 136/328 (40%), Gaps = 61/328 (18%)
Query: 4 KSIAEFTSRRSLKVWGEL-----GQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYMV 58
+ + T+ S+ + G L ++ + VD G S S+ ++ H+ E P MV
Sbjct: 15 EQVMNITNPNSIYIKGRLYFKGYKKIELHCFVDTGASLCIASKFVIPEEHWINAERPIMV 74
Query: 59 ELEDGHKVECQGKCRDVQLSIQ----HIPI--EQEFFLFNLGGLDVVLGL*WFASLGHIR 112
++ DG + CRD+ L I HIP +QE G+D ++G
Sbjct: 75 KIADGSSITINKVCRDIDLIIAGEIFHIPTVYQQE------SGIDFIIG----------- 117
Query: 113 ANFEELSLRF-HVGEKVLLRGELGLIRKNASLSSLVKALQQQAEGFWVEFQHLAEVQ--- 168
NF +L F ++V+ + ++ L +A++ EGF + ++ Q
Sbjct: 118 NNFCQLYEPFIQFTDRVIFTKDRTY---PVHIAKLTRAVRVGTEGFLESMKKRSKTQQPE 174
Query: 169 -------------------EELVAVPEQLQGLMGDYSELFSTIWDLPPSRVQDH---AIH 206
EE + + +Q + + E + L P++ + +I
Sbjct: 175 PVNISTNKIAILSEGRRLSEEKLFITQQRMQKIEELLEKVCSENPLDPNKTKQWMKASIK 234
Query: 207 LQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILV----KKKDGS 262
L + + ++P + + E +K +KE+LD+ + +PS S + LV +K+ G
Sbjct: 235 LSDPSKAIKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGK 294
Query: 263 WRFCIDYRAFHKITIPNKFPILIIDELL 290
R ++Y+A +K T+ + + DELL
Sbjct: 295 KRMVVNYKAMNKATVGDAYNPPNKDELL 322
>POL_CAMVC (P03555) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 59.3 bits (142), Expect = 2e-08
Identities = 69/317 (21%), Positives = 134/317 (41%), Gaps = 30/317 (9%)
Query: 3 LKSIAEFTSRRSLKVWGEL-----GQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYM 57
++ + T+ S+ + G L ++ + VD G S S+ ++ H+ E P M
Sbjct: 12 IEQVMNVTNPNSIYIKGRLYFKGYKKIELHCFVDTGASLCIASKFVIPEEHWVNAERPIM 71
Query: 58 VELEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEE 117
V++ DG + C+D+ L I + G+D ++G F L F +
Sbjct: 72 VKIADGSSITISKVCKDIDLIIAGEIFKIPTVYQQESGIDFIIGNN-FCQLYEPFIQFTD 130
Query: 118 L-------SLRFHVGE-----KVLLRGELGLIRKNASLSSL--VKALQQQAEGFWVEFQH 163
S H+ + +V + G L ++K + V + E E
Sbjct: 131 RVIFTKNKSYPVHITKLTRAVRVGIEGFLESMKKRSKTQQPEPVNISTNKIENPLEEIAI 190
Query: 164 LAE---VQEELVAVPEQLQGLMGDYSELFSTIWDLPPSRVQDH---AIHLQEGAAIPNIR 217
L+E + EE + + +Q + + E + L P++ + +I L + + ++
Sbjct: 191 LSEGRRLSEEKLFITQQRMQKIEELLEKVCSENPLDPNKTKQWMKASIKLSDPSKAIKVK 250
Query: 218 PYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILV----KKKDGSWRFCIDYRAFH 273
P + + E +K +KE+LD+ + +PS S + LV +K+ G R ++Y+A +
Sbjct: 251 PMKYSPMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMN 310
Query: 274 KITIPNKFPILIIDELL 290
K TI + + + DELL
Sbjct: 311 KATIGDAYNLPNKDELL 327
>POL_CAMVE (Q02964) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 58.9 bits (141), Expect = 3e-08
Identities = 69/316 (21%), Positives = 132/316 (40%), Gaps = 30/316 (9%)
Query: 4 KSIAEFTSRRSLKVWGEL-----GQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYMV 58
+ + T+ S+ + G L ++ + VD G S S+ ++ H+ E P MV
Sbjct: 13 EQVMNVTNPNSIYIKGRLYFKGYKKIELHCFVDTGASLCIASKFVIPEEHWVNAERPIMV 72
Query: 59 ELEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEEL 118
++ DG + C+D+ L I + G+D ++G F L F +
Sbjct: 73 KIADGSSITISKVCKDIDLIIAREIFKIPTVYQQESGIDFIIGNN-FCQLYEPFIQFTDR 131
Query: 119 -------SLRFHVGE-----KVLLRGELGLIRKNASLSSL--VKALQQQAEGFWVEFQHL 164
S H+ + +V G L ++K + V + E E L
Sbjct: 132 VIFTKNKSYPVHIAKLTRAVRVGTEGFLESMKKRSKTQQPEPVNISTNKIENPLKEIAIL 191
Query: 165 AE---VQEELVAVPEQLQGLMGDYSELFSTIWDLPPSRVQDH---AIHLQEGAAIPNIRP 218
+E + EE + + +Q + + E + L P++ + +I L + + ++P
Sbjct: 192 SEGRRLSEEKLFITQQRMQKIEELLEKVCSENPLDPNKTKQWMKASIKLSDPSKAIKVKP 251
Query: 219 YRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILV----KKKDGSWRFCIDYRAFHK 274
+ + E +K +KE+LD+ + +PS S + LV +K+ G R ++Y+A +K
Sbjct: 252 MKYSPMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNK 311
Query: 275 ITIPNKFPILIIDELL 290
TI + + + DELL
Sbjct: 312 ATIGDAYNLPNKDELL 327
>POL_CAMVS (P03554) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 57.4 bits (137), Expect = 9e-08
Identities = 68/316 (21%), Positives = 131/316 (40%), Gaps = 30/316 (9%)
Query: 4 KSIAEFTSRRSLKVWGEL-----GQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYMV 58
+ + T+ S+ + G L ++ + VD G S S+ ++ H+ E P MV
Sbjct: 13 EQVMNVTNPNSIYIKGRLYFKGYKKIELHCFVDTGASLCIASKFVIPEEHWVNAERPIMV 72
Query: 59 ELEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEEL 118
++ DG + C+D+ L I G+D ++G F L F +
Sbjct: 73 KIADGSSITISKVCKDIDLIIAGEIFRIPTVYQQESGIDFIIGNN-FCQLYEPFIQFTDR 131
Query: 119 -------SLRFHVGE-----KVLLRGELGLIRKNASLSSL--VKALQQQAEGFWVEFQHL 164
S H+ + +V G L ++K + V + E E L
Sbjct: 132 VIFTKNKSYPVHIAKLTRAVRVGTEGFLESMKKRSKTQQPEPVNISTNKIENPLEEIAIL 191
Query: 165 AE---VQEELVAVPEQLQGLMGDYSELFSTIWDLPPSRVQDH---AIHLQEGAAIPNIRP 218
+E + EE + + +Q + + E + L P++ + +I L + + ++P
Sbjct: 192 SEGRRLSEEKLFITQQRMQKIEELLEKVCSENPLDPNKTKQWMKASIKLSDPSKAIKVKP 251
Query: 219 YRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILV----KKKDGSWRFCIDYRAFHK 274
+ + E +K +KE+LD+ + +PS S + LV +K+ G R ++Y+A +K
Sbjct: 252 MKYSPMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNK 311
Query: 275 ITIPNKFPILIIDELL 290
T+ + + + DELL
Sbjct: 312 ATVGDAYNLPNKDELL 327
>POL2_DROME (P20825) Retrovirus-related Pol polyprotein from
transposon 297 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1059
Score = 57.0 bits (136), Expect = 1e-07
Identities = 32/86 (37%), Positives = 52/86 (60%), Gaps = 5/86 (5%)
Query: 217 RPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGS-----WRFCIDYRA 271
+ Y + E+E V+EML+ G+ R S S ++S +V KK + +R IDYR
Sbjct: 210 KQYPLAQTHEIEVENQVQEMLNQGLIRESNSPYNSPTWVVPKKPDASGANKYRVVIDYRK 269
Query: 272 FHKITIPNKFPILIIDELLYKLGEAK 297
++ITIP+++PI +DE+L KLG+ +
Sbjct: 270 LNEITIPDRYPIPNMDEILGKLGKCQ 295
Score = 33.5 bits (75), Expect = 1.4
Identities = 19/53 (35%), Positives = 29/53 (53%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFII 541
G VL Q P++F S ++D S E+EL+A+V A + + L G F+I
Sbjct: 520 GAVLSQNGHPISFISRTLNDHELNYSAIEKELLAIVWATKTFRHYLLGRQFLI 572
>POL3_DROME (P04323) Retrovirus-related Pol polyprotein from
transposon 17.6 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1058
Score = 56.2 bits (134), Expect = 2e-07
Identities = 33/87 (37%), Positives = 53/87 (59%), Gaps = 5/87 (5%)
Query: 213 IPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILV-KKKDGS----WRFCI 267
+P Y P + E+E +++ML+ GI R S S ++S + +V KK+D S +R I
Sbjct: 207 LPLYSKYSYPQAYEQEVESQIQDMLNQGIIRTSNSPYNSPIWVVPKKQDASGKQKFRIVI 266
Query: 268 DYRAFHKITIPNKFPILIIDELLYKLG 294
DYR ++IT+ ++ PI +DE+L KLG
Sbjct: 267 DYRKLNEITVGDRHPIPNMDEILGKLG 293
Score = 32.3 bits (72), Expect = 3.2
Identities = 19/53 (35%), Positives = 29/53 (53%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFII 541
G VL Q PL++ S +++ S E+EL+A+V A + + L G HF I
Sbjct: 521 GAVLSQDGHPLSYISRTLNEHEINYSTIEKELLAIVWATKTFRHYLLGRHFEI 573
>POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from
transposon opus [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1003
Score = 55.1 bits (131), Expect = 5e-07
Identities = 29/86 (33%), Positives = 49/86 (56%), Gaps = 5/86 (5%)
Query: 217 RPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKK-----DGSWRFCIDYRA 271
+ Y P + E+E+ + E+L GI RPS S ++S + +V KK + +R +D++
Sbjct: 127 KSYPYPVNMRGEVERQIDELLQDGIIRPSNSPYNSPIWIVPKKPKPNGEKQYRMVVDFKR 186
Query: 272 FHKITIPNKFPILIIDELLYKLGEAK 297
+ +TIP+ +PI I+ L LG AK
Sbjct: 187 LNTVTIPDTYPIPDINATLASLGNAK 212
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 48.1 bits (113), Expect = 6e-05
Identities = 28/86 (32%), Positives = 47/86 (54%), Gaps = 1/86 (1%)
Query: 208 QEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCI 267
QE +P IR Y P + + + + L GI R S + + V+ V KK+G+ R +
Sbjct: 408 QENYRLP-IRNYPLPPGKMQAMNDEINQGLKSGIIRESKAINACPVMFVPKKEGTLRMVV 466
Query: 268 DYRAFHKITIPNKFPILIIDELLYKL 293
DY+ +K PN +P+ +I++LL K+
Sbjct: 467 DYKPLNKYVKPNIYPLPLIEQLLAKI 492
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 48.1 bits (113), Expect = 6e-05
Identities = 28/86 (32%), Positives = 47/86 (54%), Gaps = 1/86 (1%)
Query: 208 QEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCI 267
QE +P IR Y P + + + + L GI R S + + V+ V KK+G+ R +
Sbjct: 408 QENYRLP-IRNYPLPPGKMQAMNDEINQGLKSGIIRESKAINACPVMFVPKKEGTLRMVV 466
Query: 268 DYRAFHKITIPNKFPILIIDELLYKL 293
DY+ +K PN +P+ +I++LL K+
Sbjct: 467 DYKPLNKYVKPNIYPLPLIEQLLAKI 492
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 48.1 bits (113), Expect = 6e-05
Identities = 28/86 (32%), Positives = 47/86 (54%), Gaps = 1/86 (1%)
Query: 208 QEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCI 267
QE +P IR Y P + + + + L GI R S + + V+ V KK+G+ R +
Sbjct: 408 QENYRLP-IRNYPLPPGKMQAMNDEINQGLKSGIIRESKAINACPVMFVPKKEGTLRMVV 466
Query: 268 DYRAFHKITIPNKFPILIIDELLYKL 293
DY+ +K PN +P+ +I++LL K+
Sbjct: 467 DYKPLNKYVKPNIYPLPLIEQLLAKI 492
>POLY_DROME (P10401) Retrovirus-related Pol polyprotein from
transposon gypsy [Contains: Reverse transcriptase (EC
2.7.7.49); Endonuclease]
Length = 1035
Score = 45.1 bits (105), Expect = 5e-04
Identities = 27/71 (38%), Positives = 43/71 (60%), Gaps = 6/71 (8%)
Query: 233 VKEMLDVGIFRPSISQFSSLVILVKKK------DGSWRFCIDYRAFHKITIPNKFPILII 286
VK++L GI RPS S ++S +V KK + + R ID+R ++ TIP+++P+ I
Sbjct: 201 VKQLLKDGIIRPSRSPYNSPTWVVDKKGTDAFGNPNKRLVIDFRKLNEKTIPDRYPMPSI 260
Query: 287 DELLYKLGEAK 297
+L LG+AK
Sbjct: 261 PMILANLGKAK 271
Score = 31.2 bits (69), Expect = 7.2
Identities = 19/52 (36%), Positives = 27/52 (51%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFI 540
G VL Q RP+ S + Q + EREL+A+V A+ + L+GS I
Sbjct: 507 GAVLSQEGRPITMISRTLKQPEQNYATNERELLAIVWALGKLQNFLYGSREI 558
>PK11_HUMAN (P87893) HERV-K_22q11.21 provirus ancestral Pol protein
(HERV-K101 Pol protein) [Includes: Reverse transcriptase
(RT) (EC 2.7.7.49)]
Length = 254
Score = 44.7 bits (104), Expect = 6e-04
Identities = 24/87 (27%), Positives = 40/87 (45%), Gaps = 15/87 (17%)
Query: 216 IRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKI 275
+ ++ P + + L E L+ G PS S ++S V +++KK G WR D RA + +
Sbjct: 36 VNQWQLPKQKLEALHLLANEQLEKGHIEPSFSPWNSPVFVIQKKSGKWRMLTDLRAVNAV 95
Query: 276 ---------------TIPNKFPILIID 287
IP +P++IID
Sbjct: 96 IQPMGPLQPGLPSPAMIPKDWPLIIID 122
>POL_SMRVH (P03364) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)]
Length = 888
Score = 44.3 bits (103), Expect = 8e-04
Identities = 19/47 (40%), Positives = 30/47 (63%)
Query: 232 LVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIP 278
LV+E L G P+ S +++ + ++KKK GSWR D RA +K+ +P
Sbjct: 47 LVQEQLAAGHIEPTNSPWNTPIFIIKKKSGSWRLLQDLRAVNKVMVP 93
>POK9_HUMAN (P63134) HERV-K_5q13.3 provirus ancestral Pol protein
(HERV-K104 Pol protein) [Includes: Reverse transcriptase
(RT) (EC 2.7.7.49)]
Length = 483
Score = 43.9 bits (102), Expect = 0.001
Identities = 25/87 (28%), Positives = 39/87 (44%), Gaps = 15/87 (17%)
Query: 216 IRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKI 275
+ + P + + L E L+ G PS S ++S V +++KK G WR D RA + I
Sbjct: 36 VNQWPLPKQKLEALHLLANEQLEKGHIEPSFSPWNSPVFVIQKKSGKWRMLTDLRAVNAI 95
Query: 276 ---------------TIPNKFPILIID 287
IP +P++IID
Sbjct: 96 IQPMGPLQSRLPSPAMIPKDWPLIIID 122
>POK6_HUMAN (P63133) HERV-K_8p23.1 provirus ancestral Pol protein
(HERV-K115 Pol protein) [Includes: Reverse transcriptase
(RT) (EC 2.7.7.49); Ribonuclease H (EC 3.1.26.4) (RNase
H); Integrase (IN)]
Length = 956
Score = 43.5 bits (101), Expect = 0.001
Identities = 24/87 (27%), Positives = 39/87 (44%), Gaps = 15/87 (17%)
Query: 216 IRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKI 275
+ + P + + L E L+ G PS S ++S V +++KK G WR D RA + +
Sbjct: 36 VNQWPLPKQKLEALHLLANEQLEKGHIEPSFSPWNSPVFVIQKKSGKWRMLTDLRAVNAV 95
Query: 276 ---------------TIPNKFPILIID 287
IP +P++IID
Sbjct: 96 IQPMGPLQPGLPSPAMIPKDWPLIIID 122
>POK5_HUMAN (P63132) HERV-K_19p13.11 provirus ancestral Pol protein
(HERV-K113 Pol protein) [Includes: Reverse transcriptase
(RT) (EC 2.7.7.49); Ribonuclease H (EC 3.1.26.4) (RNase
H); Integrase (IN)]
Length = 956
Score = 43.5 bits (101), Expect = 0.001
Identities = 24/87 (27%), Positives = 39/87 (44%), Gaps = 15/87 (17%)
Query: 216 IRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKI 275
+ + P + + L E L+ G PS S ++S V +++KK G WR D RA + +
Sbjct: 36 VNQWPLPKQKLEALHLLANEQLEKGHIEPSFSPWNSPVFVIQKKSGKWRMLTDLRAVNAV 95
Query: 276 ---------------TIPNKFPILIID 287
IP +P++IID
Sbjct: 96 IQPMGPLQPGLPSPAMIPKDWPLIIID 122
>POK4_HUMAN (P63128) HERV-K_6q14.1 provirus ancestral Pol protein
(HERV-K109 Pol protein) (HERV-K(C6) Pol protein)
Length = 194
Score = 43.5 bits (101), Expect = 0.001
Identities = 24/87 (27%), Positives = 39/87 (44%), Gaps = 15/87 (17%)
Query: 216 IRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKI 275
+ + P + + L E L+ G PS S ++S V +++KK G WR D RA + +
Sbjct: 36 VNQWPLPKQKLEALHLLANEQLEKGHIEPSFSPWNSPVFVIQKKSGKWRMLTDLRAVNAV 95
Query: 276 ---------------TIPNKFPILIID 287
IP +P++IID
Sbjct: 96 IQPMGPLQPGLPSPAMIPKDWPLIIID 122
>POK2_HUMAN (Q9BXR3) HERV-K_7p22.1 provirus ancestral Pol protein
(HERV-K(HML-2.HOM) Pol protein) (HERV-K108 Pol protein)
(HERV-K(C7) Pol protein) [Includes: Reverse
transcriptase (RT) (EC 2.7.7.49); Ribonuclease H (EC
3.1.26.4) (RNase H); Integrase (IN)]
Length = 956
Score = 43.5 bits (101), Expect = 0.001
Identities = 24/87 (27%), Positives = 39/87 (44%), Gaps = 15/87 (17%)
Query: 216 IRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKI 275
+ + P + + L E L+ G PS S ++S V +++KK G WR D RA + +
Sbjct: 36 VNQWPLPKQKLEALHLLANEQLEKGHIEPSFSPWNSPVFVIQKKSGKWRMLTDLRAVNAV 95
Query: 276 ---------------TIPNKFPILIID 287
IP +P++IID
Sbjct: 96 IQPMGPLQPGLPSPAMIPKDWPLIIID 122
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.349 0.158 0.543
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 54,652,694
Number of Sequences: 164201
Number of extensions: 2088905
Number of successful extensions: 8670
Number of sequences better than 10.0: 85
Number of HSP's better than 10.0 without gapping: 51
Number of HSP's successfully gapped in prelim test: 34
Number of HSP's that attempted gapping in prelim test: 8575
Number of HSP's gapped (non-prelim): 118
length of query: 542
length of database: 59,974,054
effective HSP length: 115
effective length of query: 427
effective length of database: 41,090,939
effective search space: 17545830953
effective search space used: 17545830953
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0126.7


