

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0122.7
(125 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
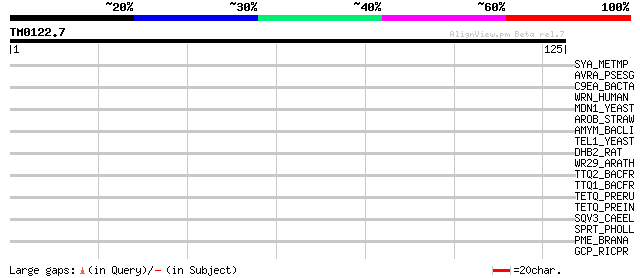
Score E
Sequences producing significant alignments: (bits) Value
SYA_METMP (P61710) Alanyl-tRNA synthetase (EC 6.1.1.7) (Alanine-... 29 1.6
AVRA_PSESG (P11437) Avirulence A protein 29 1.6
C9EA_BACTA (Q9ZNL9) Pesticidial crystal protein cry9Ea (Insectic... 28 2.8
WRN_HUMAN (Q14191) Werner syndrome helicase 28 3.6
MDN1_YEAST (Q12019) Midasin (MIDAS-containing protein) 28 3.6
AROB_STRAW (Q827R8) 3-dehydroquinate synthase (EC 4.2.3.4) 28 3.6
AMYM_BACLI (Q04977) Maltogenic alpha-amylase (EC 3.2.1.133) (Glu... 28 3.6
TEL1_YEAST (P38110) Telomer length regulation protein TEL1 28 4.7
DHB2_RAT (Q62730) Estradiol 17-beta-dehydrogenase 2 (EC 1.1.1.62... 28 4.7
WR29_ARATH (Q9SUS1) Probable WRKY transcription factor 29 (WRKY ... 27 6.2
TTQ2_BACFR (Q08425) Tetracycline resistance protein tetQ (TetA(Q)2) 27 6.2
TTQ1_BACFR (P70882) Tetracycline resistance protein tetQ (Tet(Q)... 27 6.2
TETQ_PRERU (Q52360) Tetracycline resistance protein tetQ (Tet(Q)) 27 6.2
TETQ_PREIN (O05197) Tetracycline resistance protein tetQ (Tet(Q)) 27 6.2
SQV3_CAEEL (P34548) Probable galactosyltransferase sqv-3 (EC 2.4... 27 8.0
SPRT_PHOLL (Q7N118) Protein sprT 27 8.0
PME_BRANA (P41510) Probable pectinesterase precursor (EC 3.1.1.1... 27 8.0
GCP_RICPR (Q9ZEA8) Probable O-sialoglycoprotein endopeptidase (E... 27 8.0
>SYA_METMP (P61710) Alanyl-tRNA synthetase (EC 6.1.1.7)
(Alanine--tRNA ligase) (AlaRS)
Length = 892
Score = 29.3 bits (64), Expect = 1.6
Identities = 13/38 (34%), Positives = 24/38 (62%)
Query: 42 DQVRSTLERFYMNCKEELYLLQIDAKKLGDGLVYEMVD 79
DQ+ T++RF+ KE+ ++ KK+G+ + YE+ D
Sbjct: 760 DQLPKTVKRFFEEWKEQKKTIEELQKKVGELVKYELAD 797
>AVRA_PSESG (P11437) Avirulence A protein
Length = 907
Score = 29.3 bits (64), Expect = 1.6
Identities = 16/66 (24%), Positives = 32/66 (48%)
Query: 47 TLERFYMNCKEELYLLQIDAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAE 106
+++ F N + L++++ + GL Y+ VD F H Y L++D V A+
Sbjct: 414 SMDFFRRNGPAVVMALRMNSLRKNQGLPYKEVDRCEVFTHSYAEIHGAISLTIDGVDPAD 473
Query: 107 KLSLSD 112
K+ + +
Sbjct: 474 KVEVKN 479
>C9EA_BACTA (Q9ZNL9) Pesticidial crystal protein cry9Ea (Insecticidal
delta-endotoxin CryIXE(a)) (Crystaline entomocidal
protoxin) (130 kDa crystal protein)
Length = 1150
Score = 28.5 bits (62), Expect = 2.8
Identities = 20/54 (37%), Positives = 30/54 (55%), Gaps = 4/54 (7%)
Query: 30 DKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGDGLVYEMV-DGSN 82
D + F+ LS D S R NCK Y+L++ AKK+G+G Y + DG++
Sbjct: 1039 DGNMHFLVLSHWDAQVSQQFRVQPNCK---YVLRVTAKKVGNGDGYVTIQDGAH 1089
>WRN_HUMAN (Q14191) Werner syndrome helicase
Length = 1432
Score = 28.1 bits (61), Expect = 3.6
Identities = 20/75 (26%), Positives = 38/75 (50%), Gaps = 4/75 (5%)
Query: 30 DKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGDGLVYEMVDGSNSFPHFYG 89
D + FI L+ + T++RF +N +EE+ L D K + E++D + PH +
Sbjct: 216 DAYAGFIIYRNLEILDDTVQRFAINKEEEILL--SDMNKQLTSISEEVMDLAKHLPHAF- 272
Query: 90 PSRSFSPLSLDVVTK 104
S+ +P + ++ K
Sbjct: 273 -SKLENPRRVSILLK 286
>MDN1_YEAST (Q12019) Midasin (MIDAS-containing protein)
Length = 4910
Score = 28.1 bits (61), Expect = 3.6
Identities = 17/30 (56%), Positives = 19/30 (62%), Gaps = 5/30 (16%)
Query: 58 ELYLLQIDAKKLGDGLVYEMVDGSNSFPHF 87
ELYL +AKKL D +VDGSN PHF
Sbjct: 935 ELYL---EAKKLSDNNT--IVDGSNQKPHF 959
>AROB_STRAW (Q827R8) 3-dehydroquinate synthase (EC 4.2.3.4)
Length = 363
Score = 28.1 bits (61), Expect = 3.6
Identities = 17/54 (31%), Positives = 28/54 (51%)
Query: 27 GNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGDGLVYEMVDG 80
G LD ++A H + L+ V L Y + L +++D K GD L + ++DG
Sbjct: 286 GRLDDATADRHRTVLESVGLPLHYRYDQWPKLLETMKVDKKSRGDLLRFIVLDG 339
>AMYM_BACLI (Q04977) Maltogenic alpha-amylase (EC 3.2.1.133) (Glucan
1,4-alpha-maltohydrolase)
Length = 578
Score = 28.1 bits (61), Expect = 3.6
Identities = 16/48 (33%), Positives = 27/48 (55%), Gaps = 8/48 (16%)
Query: 9 ISTAKEWEELQANGST--------FGGNLDKSSAFIHLSKLDQVRSTL 48
I T++EW+++ NG T F +L K +A IHL + + RS++
Sbjct: 245 IGTSQEWQDVVKNGETSRYKDWFIFILSLLKKAAMIHLRLVPRCRSSI 292
>TEL1_YEAST (P38110) Telomer length regulation protein TEL1
Length = 2787
Score = 27.7 bits (60), Expect = 4.7
Identities = 25/95 (26%), Positives = 35/95 (36%), Gaps = 18/95 (18%)
Query: 35 FIHLSKLDQVRSTLERFYMNCKEELYLLQIDAKK------------LGDGLVYEMVDGSN 82
F L L ++RST+E+ + YL + + LG LV
Sbjct: 1219 FTDLGSLTELRSTVEKLFPTSYLSPYLFENSSVSMRYQYPLHIPLALGATLV------QT 1272
Query: 83 SFPHFYGPSRSFSPLSLDVVTKAEKLSLSDGRFTC 117
F H + F L L V+T EK S G+ C
Sbjct: 1273 QFAHEKNNTHEFKLLFLSVITDLEKTSTYIGKLRC 1307
>DHB2_RAT (Q62730) Estradiol 17-beta-dehydrogenase 2 (EC 1.1.1.62)
(17-beta-HSD 2) (17-beta-hydroxysteroid dehydrogenase 2)
Length = 381
Score = 27.7 bits (60), Expect = 4.7
Identities = 15/47 (31%), Positives = 20/47 (41%)
Query: 22 GSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQIDAKK 68
G +LDK + LD+ E NC E L +LQ+D K
Sbjct: 96 GHALAKHLDKLGFTVFAGVLDKEGPGAEELRKNCSERLSVLQMDVTK 142
>WR29_ARATH (Q9SUS1) Probable WRKY transcription factor 29 (WRKY
DNA-binding protein 29)
Length = 304
Score = 27.3 bits (59), Expect = 6.2
Identities = 10/26 (38%), Positives = 18/26 (68%)
Query: 64 IDAKKLGDGLVYEMVDGSNSFPHFYG 89
I+ ++L +GL +++ GS +FP F G
Sbjct: 260 IEGRRLSNGLPSDLMSGSGTFPSFTG 285
>TTQ2_BACFR (Q08425) Tetracycline resistance protein tetQ (TetA(Q)2)
Length = 641
Score = 27.3 bits (59), Expect = 6.2
Identities = 19/62 (30%), Positives = 29/62 (46%), Gaps = 2/62 (3%)
Query: 16 EELQANGSTFGGNLDKSS--AFIHLSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGDGL 73
E +QA L K I ++K+D+ LER Y++ K L + + + DGL
Sbjct: 103 EGIQAQTKLLFNTLQKLQIPTIIFINKIDRDGVNLERLYLDIKTNLSQDVLFMQTVVDGL 162
Query: 74 VY 75
VY
Sbjct: 163 VY 164
>TTQ1_BACFR (P70882) Tetracycline resistance protein tetQ (Tet(Q))
(TetA(Q)3)
Length = 641
Score = 27.3 bits (59), Expect = 6.2
Identities = 19/62 (30%), Positives = 28/62 (44%), Gaps = 2/62 (3%)
Query: 16 EELQANGSTFGGNLDKSS--AFIHLSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGDGL 73
E +QA L K I ++K+D+ LER YM+ K L + + + DG
Sbjct: 103 EGIQAQTKLLFSTLQKLQIPTIIFINKIDRAGVNLERLYMDIKTNLSQDVLFMQTVVDGS 162
Query: 74 VY 75
VY
Sbjct: 163 VY 164
>TETQ_PRERU (Q52360) Tetracycline resistance protein tetQ (Tet(Q))
Length = 641
Score = 27.3 bits (59), Expect = 6.2
Identities = 19/62 (30%), Positives = 28/62 (44%), Gaps = 2/62 (3%)
Query: 16 EELQANGSTFGGNLDKSS--AFIHLSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGDGL 73
E +QA L K I ++K+D+ LER YM+ K L + + + DG
Sbjct: 103 EGIQAQTKLLFSTLQKLQIPTIIFINKIDRAGVNLERLYMDIKTNLSQDVLFMQTVVDGS 162
Query: 74 VY 75
VY
Sbjct: 163 VY 164
>TETQ_PREIN (O05197) Tetracycline resistance protein tetQ (Tet(Q))
Length = 654
Score = 27.3 bits (59), Expect = 6.2
Identities = 19/62 (30%), Positives = 29/62 (46%), Gaps = 2/62 (3%)
Query: 16 EELQANGSTFGGNLDKSS--AFIHLSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGDGL 73
E +QA L K I ++K+D+ LER Y++ K L + + + DGL
Sbjct: 103 EGIQAQTKLLFNTLQKLQIPTIIFINKIDRDGVNLERLYLDIKTNLSQDVLFMQTVVDGL 162
Query: 74 VY 75
VY
Sbjct: 163 VY 164
>SQV3_CAEEL (P34548) Probable galactosyltransferase sqv-3 (EC
2.4.1.-) (Squashed vulva protein 3)
Length = 289
Score = 26.9 bits (58), Expect = 8.0
Identities = 14/36 (38%), Positives = 19/36 (51%)
Query: 57 EELYLLQIDAKKLGDGLVYEMVDGSNSFPHFYGPSR 92
+E YL ID+K + D SN+F H +GP R
Sbjct: 189 DEFYLRIIDSKLNLTRVSGLSTDSSNTFRHIHGPKR 224
>SPRT_PHOLL (Q7N118) Protein sprT
Length = 194
Score = 26.9 bits (58), Expect = 8.0
Identities = 18/63 (28%), Positives = 28/63 (43%), Gaps = 13/63 (20%)
Query: 37 HLSKLDQVRSTLERFYMNCKEELYLLQIDAK-----------KLGDGLVYEMVDGSNSFP 85
H K+D VRS +Y NC++ L+ K K G+ L+ + +D N P
Sbjct: 115 HKFKIDSVRSQTFTYYCNCQQHELTLRRHNKILRGESHYICQKCGEKLIAKSIDSLN--P 172
Query: 86 HFY 88
+ Y
Sbjct: 173 NLY 175
>PME_BRANA (P41510) Probable pectinesterase precursor (EC 3.1.1.11)
(Pectin methylesterase) (PE)
Length = 584
Score = 26.9 bits (58), Expect = 8.0
Identities = 19/89 (21%), Positives = 45/89 (50%), Gaps = 9/89 (10%)
Query: 16 EELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLE---RFYMNCKEELYLLQIDAKKLGDG 72
E+L+ G +L +S SK+DQ++ L + +C +++ ++ K +G+G
Sbjct: 124 EDLETIVEEMGEDLQQSG-----SKMDQLKQWLTGVFNYQTDCIDDIEESEL-RKVMGEG 177
Query: 73 LVYEMVDGSNSFPHFYGPSRSFSPLSLDV 101
+ + + SN+ F+ + + S +++ V
Sbjct: 178 IAHSKILSSNAIDIFHALTTAMSQMNVKV 206
>GCP_RICPR (Q9ZEA8) Probable O-sialoglycoprotein endopeptidase (EC
3.4.24.57) (Glycoprotease)
Length = 387
Score = 26.9 bits (58), Expect = 8.0
Identities = 19/59 (32%), Positives = 28/59 (47%), Gaps = 3/59 (5%)
Query: 23 STFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGDGLVYEMVDGS 81
+ FGG + + +A HLS LDQ L+ EL + A G GL+ ++ GS
Sbjct: 38 AVFGGVVPEIAARSHLSNLDQ---ALKNVLKKSNTELTEISAIAATSGPGLIGGVIVGS 93
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.136 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,504,581
Number of Sequences: 164201
Number of extensions: 547297
Number of successful extensions: 1168
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 1162
Number of HSP's gapped (non-prelim): 18
length of query: 125
length of database: 59,974,054
effective HSP length: 101
effective length of query: 24
effective length of database: 43,389,753
effective search space: 1041354072
effective search space used: 1041354072
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0122.7


