

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0121b.6
(378 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
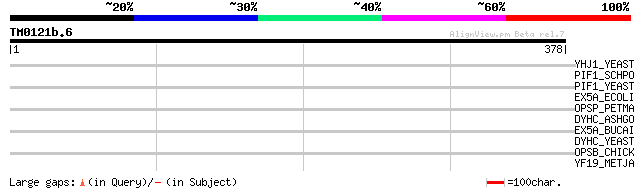
Score E
Sequences producing significant alignments: (bits) Value
YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergeni... 43 0.002
PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1, m... 40 0.010
PIF1_YEAST (P07271) DNA repair and recombination protein PIF1, m... 40 0.013
EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 37 0.082
OPSP_PETMA (O42490) Pineal opsin (P-opsin) (Pineal gland-specifi... 32 2.0
DYHC_ASHGO (Q9C1M7) Dynein heavy chain, cytosolic (DYHC) 32 2.0
EX5A_BUCAI (P57530) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 31 4.5
DYHC_YEAST (P36022) Dynein heavy chain, cytosolic (DYHC) 31 4.5
OPSB_CHICK (P28682) Blue-sensitive opsin (Blue cone photorecepto... 31 5.9
YF19_METJA (Q58914) Hypothetical protein MJ1519 30 7.7
>YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergenic
region
Length = 723
Score = 42.7 bits (99), Expect = 0.002
Identities = 29/77 (37%), Positives = 48/77 (61%), Gaps = 7/77 (9%)
Query: 277 RNTMGETTFIPRMSLTPSN--ADIPFK---FQRRQFPVILCFAMTINKSQGQSSTHVGLY 331
R T+G+ +I + + P DIP + +R Q P++LC+A++I+K+QGQ+ + +
Sbjct: 615 RWTVGKNKYIHEL-MVPERFPIDIPRENVGLERTQIPLMLCWALSIHKAQGQTIQRLKVD 673
Query: 332 LPRPVFTHVQLYVALSR 348
L R +F Q+YVALSR
Sbjct: 674 L-RRIFEAGQVYVALSR 689
>PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1,
mitochondrial precursor
Length = 805
Score = 40.0 bits (92), Expect = 0.010
Identities = 23/49 (46%), Positives = 33/49 (66%), Gaps = 1/49 (2%)
Query: 304 RRQFPVILCFAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
R Q P+IL +A++I+K+QGQ+ V + L R VF Q YVALSR ++
Sbjct: 708 RSQIPLILAYAISIHKAQGQTLDRVKVDLGR-VFEKGQAYVALSRATTQ 755
>PIF1_YEAST (P07271) DNA repair and recombination protein PIF1,
mitochondrial precursor
Length = 857
Score = 39.7 bits (91), Expect = 0.013
Identities = 24/49 (48%), Positives = 33/49 (66%), Gaps = 1/49 (2%)
Query: 304 RRQFPVILCFAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
R Q P++L ++++I+KSQGQ+ V + L R VF Q YVALSR SR
Sbjct: 691 RVQLPLMLAWSLSIHKSQGQTLPKVKVDL-RRVFEKGQAYVALSRAVSR 738
>EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5) (Exodeoxyribonuclease V 67 kDa polypeptide)
Length = 608
Score = 37.0 bits (84), Expect = 0.082
Identities = 36/117 (30%), Positives = 51/117 (42%), Gaps = 22/117 (18%)
Query: 239 GVPIMLIRNIDQTAGLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADI 298
G P+M+ RN D GL NG + I L+R G+ T R+ + +I
Sbjct: 479 GRPVMIARN-DSALGLFNGD------------IGIALDR---GQGT---RVWFAMPDGNI 519
Query: 299 PFKFQRRQFPVILCFAMTINKSQGQSSTHVGLYLP---RPVFTHVQLYVALSRVKSR 352
R +AMT++KSQG H L LP PV T +Y A++R + R
Sbjct: 520 KSVQPSRLPEHETTWAMTVHKSQGSEFDHAALILPSQRTPVVTRELVYTAVTRARRR 576
>OPSP_PETMA (O42490) Pineal opsin (P-opsin) (Pineal gland-specific
opsin)
Length = 444
Score = 32.3 bits (72), Expect = 2.0
Identities = 20/63 (31%), Positives = 32/63 (50%), Gaps = 2/63 (3%)
Query: 128 HSFRFLLIFIEPCKDPLLEIVNFSYPKLLFNLEKNSFFQERAILAPTLESVEEINNFMLA 187
H F+L+F C L ++ FSY KL+ L+K S Q L T + +++ ++
Sbjct: 207 HDHTFILMFFSTCFIFPLAVIFFSYGKLIQKLKKASETQRG--LESTRRAEQQVTRMVVV 264
Query: 188 MIL 190
MIL
Sbjct: 265 MIL 267
>DYHC_ASHGO (Q9C1M7) Dynein heavy chain, cytosolic (DYHC)
Length = 4083
Score = 32.3 bits (72), Expect = 2.0
Identities = 17/74 (22%), Positives = 40/74 (53%), Gaps = 7/74 (9%)
Query: 259 RMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFAMTIN 318
+M++ T+ + ++++ + + ET+F+ RM+ +N+D+P F+ ++ +L
Sbjct: 2795 QMLLRCGTESEKICLIIDESNILETSFLERMNTLLANSDVPGLFEADEYEALL------- 2847
Query: 319 KSQGQSSTHVGLYL 332
GQ + +GL L
Sbjct: 2848 SKIGQRISQLGLLL 2861
>EX5A_BUCAI (P57530) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5)
Length = 602
Score = 31.2 bits (69), Expect = 4.5
Identities = 31/114 (27%), Positives = 52/114 (45%), Gaps = 21/114 (18%)
Query: 238 VGVPIMLIRNIDQTAGLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNAD 297
+G PIM+I N ++ + NG I N +N+N + + +F+ + T +N
Sbjct: 480 IGKPIMIINN-NRALNVSNGNIGITN-----------INKNGILQVSFLKENN-TINN-- 524
Query: 298 IPFKFQRRQFPVILCFAMTINKSQGQSSTHVGLYLPR---PVFTHVQLYVALSR 348
IP K R +A+T++KSQG + L LP + LY ++R
Sbjct: 525 IPVKILRNYKTA---WAITVHKSQGSEFMNTALILPNFNSHILNKDTLYTGITR 575
>DYHC_YEAST (P36022) Dynein heavy chain, cytosolic (DYHC)
Length = 4092
Score = 31.2 bits (69), Expect = 4.5
Identities = 13/39 (33%), Positives = 25/39 (63%)
Query: 273 IVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVIL 311
++++ + + ET F+ RM+ +NADIP FQ ++ +L
Sbjct: 2815 LIIDESNILETAFLERMNTLLANADIPDLFQGEEYDKLL 2853
>OPSB_CHICK (P28682) Blue-sensitive opsin (Blue cone photoreceptor
pigment)
Length = 361
Score = 30.8 bits (68), Expect = 5.9
Identities = 22/75 (29%), Positives = 37/75 (49%), Gaps = 6/75 (8%)
Query: 128 HSFRFLLIFIEPCKDPLLEIVNFSYPKLLFNLEKNSFFQERAILAPTLESVEEINNFMLA 187
H+ ++L C L I+ FSY +LL L + QE++ A T ++ E+ ++
Sbjct: 206 HNESYVLFLFTFCFGVPLAIIVFSYGRLLITLRAVARQQEQS--ATTQKADREVTKMVVV 263
Query: 188 MILGEEIEYLNCDTP 202
M+LG +L C P
Sbjct: 264 MVLG----FLVCWAP 274
>YF19_METJA (Q58914) Hypothetical protein MJ1519
Length = 1175
Score = 30.4 bits (67), Expect = 7.7
Identities = 17/43 (39%), Positives = 26/43 (59%), Gaps = 3/43 (6%)
Query: 313 FAMTINKSQGQSSTHVGLYLPRPVFTHV---QLYVALSRVKSR 352
+A+TI+KSQG +V L +P+ + V LY A++R K R
Sbjct: 961 YAITIHKSQGSGFENVILIIPKGLNKFVSKEMLYTAITRAKKR 1003
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.343 0.151 0.491
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,731,139
Number of Sequences: 164201
Number of extensions: 1414111
Number of successful extensions: 5323
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 5315
Number of HSP's gapped (non-prelim): 12
length of query: 378
length of database: 59,974,054
effective HSP length: 112
effective length of query: 266
effective length of database: 41,583,542
effective search space: 11061222172
effective search space used: 11061222172
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0121b.6


